GENETICS AND PLANT BREEDING
Overall heritability in popcorn estimated by meta-analysis
Overall heritability in popcorn estimated by meta-analysis
Acta Scientiarum. Agronomy, vol. 43, e53721, 2021
Editora da Universidade Estadual de Maringá - EDUEM
Received: 18 May 2020
Accepted: 03 September 2020
ABSTRACT: The objective of this study was to conduct a meta-analysis and test its efficiency in summarizing the heterogeneous data of heritability estimates for the traits of grain yield (GY) and popping expansion (PE), and to provide reliable estimates of selection gains in popcorn. Therefore, 97 heritability estimates (ĥ2) for popcorn GY and PE in the broad and narrow sense were used. The main procedures underlying the estimation of the combined heritability (ĥ2+ ) using the technique of meta-analysis consisted of i) an exploratory analysis of the set of heritability estimates to detect outliers using a box-plot chart, ii) the verification of the required statistical assumptions, iii) testing the involved heritability estimates for homogeneity, and iv) the calculation of the estimates of combined heritability. The meta-analysis facilitated the synthesis of the information pertaining to heritability in popcorn. The combined heritability estimates (ĥ2+ ) in the broad sense for GY and PE were 0.5208 ± 0.0229 and 0.6356 ± 0.0209, respectively, and in the narrow sense were 0.3290 ± 0.0292 and 0.3083 ± 0.0298, respectively.
Keywords: meta-analysis, data compilation, summary data.
Introduction
The concept of heritability (h2), introduced to distinguish genetic and non-genetic differences among plants, is fundamental in popcorn breeding programs to guide decisions related to the selection method applied, traits selected in the early and advanced stages of the program, and the estimation of genetic gains. According to Fehr (1993), high heritability indicates that selection in the early generations of selfing is effective, whereas low heritability indicates that selection should only be applied in the more advanced generations as an increase in homozygosis increases narrow-sense heritability.
Heritability reflects the inheritable proportion of phenotypic variation, that is, it quantifies the reliability of the phenotypic value as a guide to the genetic value. Only the phenotypic value of a plant is measurable, but it is the genetic value that will influence the next generation. Thus, it is important to know how much of the phenotypic variation is attributable to genotypic variation, which is measured by heritability (Falconer & Mackay, 1996; Ramalho, Abreu, Santos, & Nunes, 2012). Two types of heritability can be estimated: heritability in the broad sense (ĥ2a ), defined as the ratio of genotypic to phenotypic variance, and in the narrow sense (ĥ2r ), defined as the ratio of additive genetic to phenotypic variance (Allard, 1971).
Popping expansion (PE) is an oligogenic trait, with heritability estimates varying from 70 to 90% (Pereira & Amaral Júnior, 2001; Rangel, Amaral Júnior, Gonçalves, Freitas Júnior, & Candido, 2011), increasing the chances of obtaining gains by selection (Alexander, 1988). According to the multiple inheritance studies (Dofing, D’Croz-Mason, & Thomas-Compton, 1991; Pereira & Amaral Júnior, 2001; Rangel et al., 2011; Moterle et al., 2012; Coan et al., 2019), the main component of genetic variation of PE is the additive genetic effect. On the other hand, the heritabilities of yield traits are generally lower than those estimated for PE; therefore, the selection of inbred families based on yield is more difficult. Additionally, for PE, as well as for grain yield (GY), substantial variation is observed in the heritability estimates, be it in the broad or narrow sense.
In an analysis of S1 and S2 families of the popcorn population, Vilarinho, Viana, Santos, and Câmara (2003) found broad-sense heritabilities of 60 and 32% for PE, and 33.58 and 27.83% for GY, respectively. In another study, in a test of S2 families of the Beija-flor popcorn population, Santos, Viana, Vilarinho, and Câmara (2004) obtained broad-sense heritability estimates of 72% for PE and 56.92% for GY. For PE of genotypes with different inbreeding levels, Arnhold, Mora, and Deitos (2006) found estimates from 35.93 to 90% among families, from 59.13 to 88.79% within families, and from 16.92 to 81.73 % at the plant level, whereas for GY, the estimates varied from 33.48 to 90.18% among families, from 22.47 to 55.99% within families, and from 21.93 to 48.82% at the plant level. In experiments at three sites, with topcross hybrids derived from S3 lines, Arnhold, Mora, Silva, Good-God, and Rodovalho (2009) obtained heritability estimates between 67 and 85% for PE and between 48 and 62% for GY. In another study, with S3 families, Lima et al. (2020) obtained heritability estimates of 76% for PE and 40% for GY.
Usually, heritability is estimated from an analysis of variance. The occurrence of errors associated with heritability estimates and other components of genetic variance is normal (Pešek & Baker, 1969). There is a large variation in heritability estimates of the same trait that can be attributed to sampling, population differences, and environmental differences (Vencovsky, 1987). For certain traits, these variations in results hamper decision making. A possible way of overcoming this problem is to compute a mean based on the combination of estimates of pre-existing results (Koots, Gibson, Smith, & Wilton, 1994), establishing a synthesis.
Combined estimates can be obtained by a meta-analysis (Giannotti, Packer, & Mercadante, 2005), which consists of a statistical procedure that provides a quantitative and summarized overview of different but related studies (Glass, 1976; Gurevitch, Koricheva, Nakagawa, & Stewart, 2018). The statistical methods used in the meta-analysis ensure that the combined estimate is accurate, mainly because of the increase in the number of observations, and consequently, the statistical power and possibility of detecting variability among the studies (Fagard, Staessen, & Thijs, 1996), which is not possible by simply averaging the published results.
The objective of this study was to apply the methodology of meta-analysis to the problem of testing the efficiency of summarizing information from heritability estimates (ĥ2) in the broad and narrow sense, for GY and PE, to provide reliable estimates of heritability.
Material and methods
Two types of meta-analysis models can be applied to combine information: the fixed-effect and the random-effect models (Sousa & Ribeiro, 2009). For the fixed-effect model, the studies are assumed to be homogeneous, that is, the observed variability among the results of the studies underlying the meta-analysis is ascribed only to the internal sample variability of each study (Whitehead, 2002). On the other hand, the random-effect model assumes that the studies are dissimilar, with variability within and between studies.
The data used in this study included 97 heritability estimates of S1 to S4 inbred popcorn families, in the broad and narrow sense, for GY and PE. These data were compiled and summarized from independent but related studies. These consisted of scientific articles published in national and international journals, annals of congresses, theses, and dissertations, as well as in the databases SCIELO, SCOPUS, and ISI, using the search terms: ‘heritability’, ‘popcorn’, ‘popping expansion’, ‘diallel’, ‘top cross’, and ‘genetic parameters’.
To ensure the validity of the procedures used in the meta-analysis, the assumptions of the normality and independence of the heritability estimates must be met (Hedges & Olkin, 1985). The assumption of independence was partly satisfied because the estimates were extracted from different studies. However, the normality assumption was tested using the Shapiro-Wilk test (Shapiro & Wilk, 1965).
The main procedures involved in calculating the estimates of combined heritability (ĥ2+ ), in the broad and narrow sense, for PE and GY in popcorn by meta-analysis were: i) exploratory analysis of the datasets (compiled heritability estimates), ii) verification of the required statistical assumptions, iii) homogeneity test of the heritability estimates involved, and iv) calculation of combined heritability estimates.
The set of heritability estimates was subjected to exploratory analysis to detect outliers using a box-plot chart. This chart was used to evaluate the empirical data distribution, such as the position, dispersion, symmetry, and discrepancy of the data. In summary, the objective was to determine whether the datasets were comparable with each other (Bussab & Morettin, 2003).
To obtain the combined estimate and test for homogeneity, the variances (s2i ) associated with the heritability estimates (ĥ2i ) must be known. However, because the vast majority of studies did not contain this information it had to be calculated by the method described by Falconer and Mackay (1996): , where N is the number of estimates involved in the meta-analysis.
The homogeneity test is important for determining the best model for a particular meta-analysis. Thus, based on ĥ2i and s2i , we tested the null hypothesis, that is, the statement that the studies that make up the meta-analysis are homogeneous, Ho: , against the hypothesis that at least one heritability estimate differs from the others, utilizing Cochran's Q test (Cochran, 1937), given by: , where wi is the weight of study i, given by Wi = 1/s2i ; ĥ2i is the i-th heritability for a given trait; ĥ2+ is the combined estimate of the heritabilities, given by, , and k is the number of values sampled for that trait (Hedges & Olkin, 1985). A significant result of the statistical test implies that the variation in the estimates among the studies is greater than that expected by chance; thus, the hypothesis that the estimates do not differ from each other is rejected (Wang & Bushman, 1999). If the null hypothesis is not rejected, the studies are considered homogeneous (p ≥ 0.01).
Thus, in the case of homogeneity, we adopted the fixed-effect model, with the structure: and, in case of heterogeneity, the random-effect model was used, structured as: , where (i is the random effect of each study i; and ei is the random error of study i. In the random-effects model, it is assumed that random errors have a normal distribution, with mean 0 and known variance , the same assumption as that of the fixed-effect model. The random effects have a normal distribution with a mean of 0 and unknown variance . In this model, the point estimate for ĥ2+ similarly consists of the weighted mean between the effect measures of each study with the difference in the estimate of (2( influencing the weighting (Deeks, Higgins, & Altman, 2019). Because εi and ei are independent, we have . The parameter (2( represents the variability between the studies and it should be estimated when using the random-effect model. Subsequently, the combined standard deviation associated with ĥ2+ was obtained by
.
Statistical analyses were performed with the Rcmdr plug-in and metafor package of R software (R Core Team, 2019).
Results and discussion
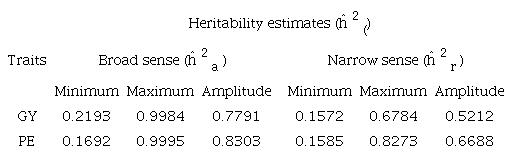
The minimum and maximum values and amplitude of variation of the heritability estimates (ĥ2() in the broad and narrow sense for PE and GY (Table 1) were 3.5 and 3.3 times higher than the minimum broad and narrow-sense values for GY, respectively. However, for PE, the amplitudes were 4.9 and 4.2 times higher than the minimum value in the broad and narrow sense, respectively. These high amplitudes confirmed the discordant results indicated by the literature, consequently justifying the application of the technique of meta-analysis.
For the exploratory analysis of the set of heritability (ĥ2() estimates of the traits under study, a box-plot chart was used (Figure 1) to show the distribution and summary of the main statistics of the data, as well as those performed by Senhorinho et al. (2019). For the estimates of broad-sense heritability (ĥ2a ), values below 0.2667 were considered discrepant for PE. For narrow-sense heritability estimates (ĥ2r ), values above 0.6784 for GY and 0.8273 for PE were also considered discrepant.
Source: first author.
No exclusion criterion was adopted for the discrepant values because, in the studies from which they were originally extracted, the authors explained the reasons why they obtained low or high heritability values for the traits studied (Giannotti et al., 2005). As conditions for the validity of the statistical methods used in the meta-analysis, the assumptions of the normality and independence of the heritability estimates must be satisfied. The assumption of independence was partly satisfied because the estimates were extracted from different studies. The normality assumption, on the other hand, was tested by the Shapiro-Wilk test, and all traits evaluated in the broad and narrow sense had a normal probability distribution (Table 2).
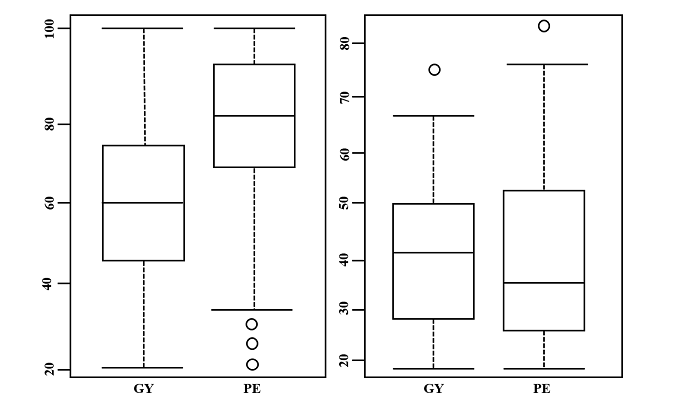
Figure 1
Box-plots of heritability estimates (%) in the broad sense (ĥ2a ) and narrow sense (ĥ2r ), for the traits of yield (GY) and popping expansion (PE) in popcorn
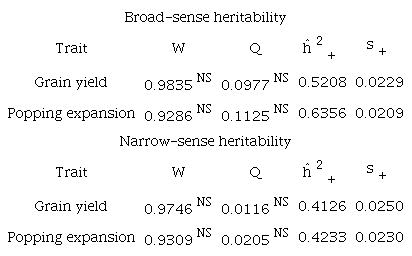

NS was non-significant by the Shapiro-Wilk test and Q test at a 1% probability.
Once the required assumptions were met, the homogeneity between the heritability estimates was tested using the Q statistic. This homogeneity test is fundamental for decisions regarding the choice of either a fixed or random-effect model. The hypothesis of homogeneity among heritability estimates was accepted for all studied traits, and consequently, the fixed-effect model was adopted (Table 2), based on: .
For the combined estimates of heritabilities (ĥ2+ ) for the evaluated traits and the combined standard deviations associated with these combined estimates (Table 2), we found broad-sense heritability values of 0.5208 for GY and 0.6356 for PE. These values were different from the results obtained if a simple arithmetic mean was calculated, having values of 0.5925 and 0.7182, respectively. It is worth mentioning that for PE, the value obtained is below, but relatively close to the lowest value (70%) of the parameter established by Pereira and Amaral Júnior (2001) and Lima et al. (2020), who determined that heritability estimates should be within a range between 70 and 90%. Therefore, the PE estimate by the meta-analysis was sufficiently high to ensure satisfactory gains with selection.
For narrow-sense heritability, ĥ2+ values of 0.4126 for GY and 0.4233 for PE were obtained. These values also differed from those of the arithmetic mean, in which the estimates were 0.3677 and 0.3766, respectively. Heritability in the narrow sense considers only the effect of additive genes, unlike heritability in the broad sense, which considers the effect of additive and non-additive genes. Consequently, heritability in a narrow sense will always tend to be less. These were far different from the extreme values found in the literature, corresponding to less than half of the amplitude obtained for the traits PE and GY by meta-analysis. The magnitude of heritability indicates whether the trait of interest is inheritable. Thus, high-performing parents tend to produce high-performing progenies.
In scientific research, the number of similar articles published in different areas of knowledge, resulting from similar research objectives, is increasing, thereby generating interest in the estimation of the synthesis of their results (Fagard et al., 1996; Wang et al., 2013), as in this study. In this context, the application of meta-analysis has contributed to obtaining an overall measure of the traits under study, which is even more relevant when considering the extreme lack of studies involving this technique, especially in the areas of breeding and crop production.
Conclusion
The method of meta-analysis is recommended to synthesize information about heritability in popcorn, and is indispensable for the appropriate analysis of the dataset to be summarized, such that reliable estimates can be obtained to achieve satisfactory gains with selection. The combined heritability estimates (ĥ2+ ) in the broad sense for GY and PE were 0.5208 ± 0.0229 and 0.6356 ± 0.0209, respectively, and in the narrow sense were 0.3290 ± 0.0292 and 0.3083 ± 0.0298, respectively.
References
Alexander, D. E. (1988). Breeding special industrial and nutritional types. In G. P. Sprague, & J. W. Dudley (Eds.), Corn and corn improvement (3rd ed., p. 869-880, Agronomy Monograph, 18). DOI: 10.2134/agronmonogr18.3ed.c14
Allard, R. W. (1971). Princípios do melhoramento genético das plantas. São Paulo, SP: Edgard Blüchner.
Arnhold, E., Mora, F., & Deitos, A. (2006). Correlaciones genéticas en famílias S4 de maíz (Zea mays). Ciencia e Investigación Agraria, 33(1), 125-131. DOI: 10.7764/rcia.v33i2.335
Arnhold, E., Mora, F., Silva, R. G., Good-God, P. I. V., & Rodovalho, M. A. (2009). Evaluation of top-cross popcorn hybrids using mixed linear model methodology. Chilean Journal of Agricultural Research, 69, 46-53. DOI: 10.4067/S0718-58392009000100006
Bussab, W. O., & Morettin, P. A. (2003). Estatística básica (5. ed.). São Paulo, SP: Saraiva.
Coan, M. M. D., Pinto, R. J. B., Kuki, M. C., Amaral Junior, A. T., Figueiredo, A. S. T., Scapim, C. A., & Warburton, M. L. (2019). Inheritance study for popping expansion in popcorn vs. flint corn genotypes. Agronomy Journal, 3(5), 2174-2183. DOI: 10.2134/agronj2019.04.0295
Cochran, W. G. (1937). Problems arising in the analysis of a series of similar experiments. Journal of the Royal Statistical Society, 4(1), 102-118. DOI: 10.2307/2984123
Deeks, J. J., Higgins, J. P., & Altman, D. G. (2019). Analysing data and undertaking meta‐analyses. Cochrane Handbook for Systematic Reviews of Interventions, 241-284. DOI: 10.1002/9781119536604.ch10
Dofing, S. M., D’Croz-Mason, N., & Thomas-Compton, M. A. (1991). Inheritance of expansion volume and yield in two popcorn x dent corn crosses. Crop Science, 31(3), 715-718. DOI: 10.2135/cropsci1991.0011183x003100030035x
Fagard, R. H., Staessen, J. A., & Thijs, L. (1996). Advantages and disadvantages of the meta-analysis approach. Journal of Hypertension, 14(2), 9-12. DOI: 10.1097/00004872-199609002-00004
Falconer, D. S., & Mackay, T. F. C. (1996). Introduction to quantitative genetics. Edinburgh, UK: Addison Wesley Longman.
Fehr, W. R. (1993). Principles of cultivar development (v. 1). New York, NY: MacMillan Publishing Company.
Giannotti, J. G., Packer, I. U., & Mercadante, M. E. Z. (2005). Meta-análise das estimativas de herdabilidade para características de crescimento em bovinos de corte. Revista Brasileira de Zootecnia, 34, 1173-1180. DOI: 10.1590/s1516-35982005000400011
Glass, G. V. (1976). Primary, secondary, and meta-analysis of research. Educational Researcher, 5(10), 3-8. DOI: 10.3102/0013189x005010003
Gurevitch, J., Koricheva, J., Nakagawa, S., & Stewart, G. (2018). Meta-analysis and the science of research synthesis. Nature, 555(7695), 175-182. DOI: 10.1038/nature25753
Hedges, L.V., & Olkin, I. (1985). Statistical methods for meta-analysis. London, UK: Academic Press.
Koots, K. R., Gibson, J. P., Smith, C., & Wilton, J. W. (1994). Analysis of published genetic parameter estimates for beef production traits. 1. Heritability. Animal Breeding abstracts, 62(5), 309-338.
Lima, V. J., Viana, A. P., Amaral Júnior, A. T., Kamphorst, S. H., Leite, J. T., Santos, P. H. A. D., ... Santos, T. O. (2020). Exploring the use of testers to maximize selection accuracy of partially inbred S3 popcorn progênies. Revista Brasileira de Ciências Agrárias, 15(2), e6557. DOI: 10.5039/agraria.v15i2a6557
Moterle, L. M., Braccini, A. L., Scapim, C. A., Pinto, R. J. B., Gonçalves, L. S. A., Rodrigues, R., & Amaral Júnior, A. T. (2012). Combining ability of popcorn lines for seed quality and agronomic traits. Euphytica, 185, 337-347. DOI: 10.1007/s10681-011-0458-2
Pereira, M. G., & Amaral Júnior, A. T. (2001). Estimation of genetic components in popcorn based on nested design. Crop Breeding and Applied Biotechnology, 71(1), 3-10. DOI: 10.13082/1984-7033.v01n01a01
Pešek, J., & Baker, R. J. (1969). Desired improvement in relation to selection indices. Canadian Journal of Plant Science, 49, 803-804. DOI: 10.4141/cjps69-137
Ramalho, M. A. P., Abreu, A. F. B., Santos, J. B., & Nunes, J. A. R. (2012). Aplicações da genética quantitativa no melhoramento de plantas autógamas. Lavras, MG: UFLA.
Rangel, R. M., Amaral Júnior, A. T., Gonçalves, L. S. Z., Freitas Júnior, S. P., & Candido, L. S. (2011). Análise biométrica de ganhos por seleção em população de milho pipoca de quinto ciclo de seleção recorrente. Revista Ciência Agronômica, 42(2), 473-481. DOI: 10.1590/s1806-66902011000200029
R Core Team (2019). R: A language and environment for statistical computing. Vienna, AT: R Foundation for Statistical Computing.
Santos, J. F., Viana, J. M. S., Vilarinho, A. A., & Câmara, T. M. M. (2004). Efficiency of S2 progeny selection strategies in popcorn. Crop Breeding and Applied Biotechnology, 4(2), 183-191.
Senhorinho, H. J. C., Coan, M. M. D., Marino, T. P., Kuki, M. C., Pinto, R. J. B., Scapim, C. A., & Holland, J. B. (2019). Genomic‐wide association study of popping expansion in tropical popcorn and field corn germplasm. Crop Science, 59, 2007-2019. DOI: 10.2135/cropsci2019.02.0101
Shapiro, S. S., & Wilk, M. B. (1965). An analysis of variance test for normality (complete samples). Biometrika, 52(3), 591-611. DOI: 10.2307/2333709
Sousa, M. R., & Ribeiro, A. L. P. (2009). Revisión sistemática y meta-análisis de estúdios de diagnóstico y pronóstico: uma guia. Arquivos Brasileiros de Cardiologia, 92, 241-251. DOI: 10.1590/S0066-782X2009000300013
Vencovsky, R. (1987). Herança quantitativa. In E. Paterniani (Coord.), Melhoramento e produção de milho no Brasil (p. 122-201). Piracicaba, SP: Fundação Cargill.
Vilarinho, A. A., Viana, J. M. S., Santos, J. F., & Câmara, T. M. M. (2003). Eficiência da seleção de progênies S1 e S2 de milho-pipoca, visando à produção de linhagens. Bragantia, 6, 9-17. DOI: 10.1590/S0006-87052003000100002
Wang, M. C., & Bushman, B. J. (1999). Integration results: through meta-analytic review using SAS software. Cary, NC: SAS Institute.
Wang, Y., Huang, Z., Deng, D., Ding, H., Zhang, R., Wang, S., … Xu, X. (2013). Meta-analysis combined with syntenic metaQTL mining dissects candidate loci for maize yield. Molecular Breeding, 31, 601-614. DOI: 10.1007/s11032-012-9818-4
Whitehead, A. (2002). Meta-analysis of controlled clinical trial. Chichester, UK: John Wiley & Sons.
Author notes
* Author for correspondence. E-mail: renan_uhdre@hotmail.com
