PLANT BREEDING
Selection of Theobroma grandiflorum clones adapted to agroforestry systems using an additive index
Selection of Theobroma grandiflorum clones adapted to agroforestry systems using an additive index
Acta Scientiarum. Agronomy, vol. 45, e57519, 2023
Editora da Universidade Estadual de Maringá - EDUEM
Received: 22 January 2021
Accepted: 13 May 2021
ABSTRACT: In fruit tree breeding, selection indices are used to identify the genotypes that combine desirable commercial and non-commercial characteristics. As Theobroma grandiflorum is generally cultivated in agroforestry systems (AFS), there is a need to develop cultivars that are adapted to such environments. In this study, the objective was to select the most promising genotypes for their future use in AFS based on the additive index, a pioneering method for this crop. The trial was carried out for 12 years in an agroforestry system in the municipality of Tomé-Açu, Pará State, Brazil. The 16 evaluated clones were completely randomised with a variable number of repetitions. The average number of fruits produced as well as the morpho-agronomic characteristics of the fruits were analysed. Mixed linear models were used to estimate the components of variance and predict the genotypic values. The genetic correlation between the variables was estimated, and the selection of genotypes was based on the additive index, with a positive orientation of all variables except the thickness of the fruit shells and the weight of the fruits. Clones 42, 44, 46, 47, 57, 61, and 64 performed well for all the analysed variables, resulting in a selection gain of 7.3% and low incidence rates of witches’ broom disease. These genotypes can be made available to producers in the form of clones for use in AFS and can further be included in future hybridisations in T. grandiflorum breeding.
Keywords: Selection index, fruit tree breeding, genetic parameters, Moniliophthora perniciosa.
Introduction
The cultivation of native fruit trees in the Brazilian Amazon, particularly in agroforestry systems (AFS), provides an alternative land for cultivations, especially for areas that have already been deforested. This practice offers environmental benefits, such as forest restoration, as well as socio-economic benefits by generating income for producers, keeping them from migrating to other lands and, thus avoiding the deforestation of preserved areas (Homma, 2014). Native fruit trees cultivation has gained increasing prominence nationally and internationally due to the vast species diversity, which translates to a multiplicity of biochemical compounds with health benefits (Lima et al., 2019; Faria et al., 2020).
Among these species, the cupuassu tree (Theobroma grandiflorum Willd. (ex. Spreng) Schumm.) has emerged as a promising material. It is an arboreal species belonging to the Malvaceae family and the Byttnerioideae subfamily. Its fruit, cupuassu, offers two products that are currently exploited by agroindustries: pulp, which is widely used to produce juices, jellies, sweets, and other foods (Pereira, Abreu, & Rodrigues, 2018); and almond, which provides the raw material for several products in the pharmaceutical and cosmetic industries owing to its emollient and antioxidant properties (Costa et al., 2020). Almonds are also used to make "cupulate" or cupuassu chocolate (Genovese & Lannes, 2009). The shell of the fruit can be used for human consumption, to manufacture compost, and as animal feed (Salgado, Rodrigues, Donado-Pestana, Dias, & Morzelle, 2011).
The state of Pará, Brazil, is the largest T. grandiflorum producer in the world, with approximately 27,000 tons of fruit produced in 2018 (Pará, 2020). T. grandiflorum crops can be found in all states in the north of Brazil, as well as in some states of the Northeast, in addition to other Latin-American countries. However, the volume of fruits currently produced is below the genetic potential of the crop, which makes it difficult for the T. grandiflorum production chain to evolve. This can be attributed to two factors: the lack of varieties with productive values that are adapted to the different edaphoclimatic conditions throughout Brazil, and the fungus Moniliophthora perniciosa (Stahel) Aime & Phillips-Mora, which is the etiological agent of witches’ broom disease that affects both cupuassu and cacao (Theobroma cacao L.). Outbreaks of the disease have been responsible for losses that reach up to 90% of production (Meinhardt et al., 2008), in some cases leading farmers to abandon orchards.
As such, there is an urgent need to guide T. grandiflorum genetic improvement programs, focusing on the development of cultivars adapted to AFS, preferably in small-scale farming environments, which make up the majority of T. grandiflorum producers. Furthermore, there is a need for cultivars with desirable characteristics, such as high rates of productivity, resistance to M. perniciosa, and commercially viable fruits, i.e., with high yields of pulp and almonds (Alves, Chaves, Gama, Pedroza Neto, & Santos, 2020).
To optimise the selection of genotypes based on more than one characteristic, methodologies involving selection indexes can be employed. Such methods weigh the different characteristics considering their relative importance and select genotypes based on a ranking of scores (Céron-Rojas & Crossa, 2018). The technique of assigning weights to each character differentiates the methodologies.
In this study, we used the additive index, which considers the coefficient of genotypic variation as the weight attributed to a variable. The main advantage of this method is that it is based on mixed modelling, allowing it to be used with unbalanced data, which is a common situation with perennial fruit trees with a long breeding cycle. This index has been used with other fruit trees, such as Jatropha curcas L. (Alves et al., 2019) and Carica papaya L. (Pereira, Abreu, & Rodrigues, 2019). However, there is no precedent for the use of selection indexes for T. grandiflorum. As such, this is a pioneering study that will inform future applications of T. grandiflorum.
Thus, the objective of this study was to use an additive index to select genotypes that are best adapted to environments with interspecific competition (i.e., AFS), with special emphasis on the productive characteristics, fruit quality, and resistance to M. perniciosa.
Material and methods
Test implementation
The test was conducted in an experimental area of Embrapa Amazonia Oriental, in the municipality of Tomé-Açu (02°26’08’’ S; 48°09’08’’ W; 145 m), northeastern Pará State, 120 km from the state capital, Belém, Brazil. The soil is well-structured, with a medium texture and low natural fertility, and is classified as yellow latosol or oxisol. According to the Köppen classification, the climate is categorised as Ami and is mesothermic and humid (Bolfe & Batistella, 2011). The relative air humidity was approximately 85%, with an average annual precipitation of 2,300 mm and an average temperature of 26°C during the years of evaluation, according to data provided by the weather station installed at Embrapa Amazonia Oriental Headquarters.
The preparation of the area and seedlings, as well as the implementation and management of the crop, followed the guidelines recommended for T. grandiflorum crop production (Souza, 2007; Alves, 2012). Sixteen selection candidates were evaluated (Table 1) from a range of genotypes with different geographical origins to exploit the vast heterogeneity among T. grandiflorum populations throughout the Amazon region (Alves, Sebbenn, Artero, Clement, & Figueira, 2007).

The experiment was installed in an AFS composed of banana (Musa sp.) (5 × 5 m), cupuassu tree (5 × 5 m), and taperebazeiro (Spondias mombin L.) (30 × 30 m). All species were planted simultaneously in March of 2007. Along the two borders of the experimental area, seedlings of abiu (Pouteria caimito (Ruiz and Pavon) Radlk.) were planted to serve as a windbreak, at a spacing of 5 m between plants, with each plant placed at the head of a row of T. grandiflorum. The other two lateral sides were bordered by a secondary forest at an intermediate stage of succession.
The experiment was conducted over 12 years and followed a completely randomised design. This experimental design was necessary considering the negative effects of self-incompatibility and, in some cases, inter-incompatibility of the species (Venturieri, 2011). Thus, individuals of four different clones surrounded an individual of a given clone. This pattern was repeated throughout the experimental area to maximise pollination efficiency and fruiting. The number of repetitions varied from 15 to 30 per clone, with one plant per plot (single-tree plot).
Data acquisition
Evaluations were conducted at the plant level over nine consecutive harvests (2010-2011 to 2018-2019). In the cupuassu tree, the harvest opening coincides with the beginning of the rainy season and extends over the entire period of about six months (Venturieri, 2011). Therefore, each harvest was divided into four evaluations with 45-day intervals between them. The data collected included the average number of fruits produced per harvest (NF) and the rate of plants affected by witches’ broom disease (WB), based on the identification of symptomatic individuals within the same genotype. The fruit characteristics were also assessed. Five fruits/plant/harvest were sampled for each clone, and the fruit length (FL) (mm), fruit diameter (FD) (mm), shell thickness (ST) (mm), shell weight (ShW) (g), seed weight (SeW) (g), pulp weight (PW) (g), and total fruit weight (FW) (g) were measured.
Statistical procedures
Considering that the data are unbalanced, mixed linear models were used to estimate the components of variance (REML) and the prediction of genotypic values (BLUP). The statistical model used was as follows: y = Xu + Zg + e, where y is the data vector; u is the scale referring to the general average, considered as a fixed effect; g is the vector of genotypic effects, considered as random [g ~ N (0, σg2)]; and e is the residual vector [e ~ N (0, σe2)]. The capital letters X and Z represent the incidence matrices for these effects. The significance of the genotypic effects was tested by deviance analysis, that is, the likelihood ratio test: LRT= (2(LogL-LogLR), where LogL is the logarithm of the maximum point of the residual likelihood function (L) for the complete model, and LogLR is the logarithm of the maximum point of the residual likelihood function (LR ) for the model without genotypic effects, using the chi-square statistic with one degree of freedom and p ≤ 0.01. The averages of the nine harvests were used to perform genetic and statistical analyses.
The following components of variance were estimated for eight evaluated characteristics, except WB: individual phenotypic variance (σy2), genotypic variance (σg2), residual variance (σe2), heritability of total genotypic effects (hg2), and coefficients of genotypic variation (CVgi) and residual variation (CVe). For the latter two, the relative coefficient of variation (CVr) was obtained. The additive index was used to select superior genotypes, as determined by Resende, Silva, and Azevedo (2014) as: , where wt is the weight attributed to variable t, GVit is the genotypic value of individual i for variable t, and σt is the standard error of the genotypic values for variable t. The coefficients of genotypic variation were used as the weight, and the direction of selection was "higher" for all variables except ST and ShW. These analyses were performed using Selegen-REML/BLUP software (Resende, 2016).
Correlations between the variables’ genetic values, except for WB, were estimated to assess their interrelation at the genetic level for the experiment in question. These values were employed for the elaboration of a correlation network, using the statistical software R (R Core Team, 2020), with the aid of the extensions "qgraph" (Epskamp, Vramer, Waldorp, Schmittman, & Borsboom, 2012) and "biotools" (Silva, Malafaia, & Menezes, 2017).
Results
Fruit production began slowly in the first two harvests but progressed considerably from the third when the genotypes expressed their full potential. In the fourth harvest, the production stability phase was reached, although minor fluctuations were observed in the subsequent harvests (Figure 1). As an exception, the sixth harvest stands out, corresponding to the end of 2015 and the beginning of 2016, which suffered from the effects of El Niño 2015 (Jiménez-Muñoz et al., 2016), which caused a prolonged drought that year, affecting the performance of genotypes.
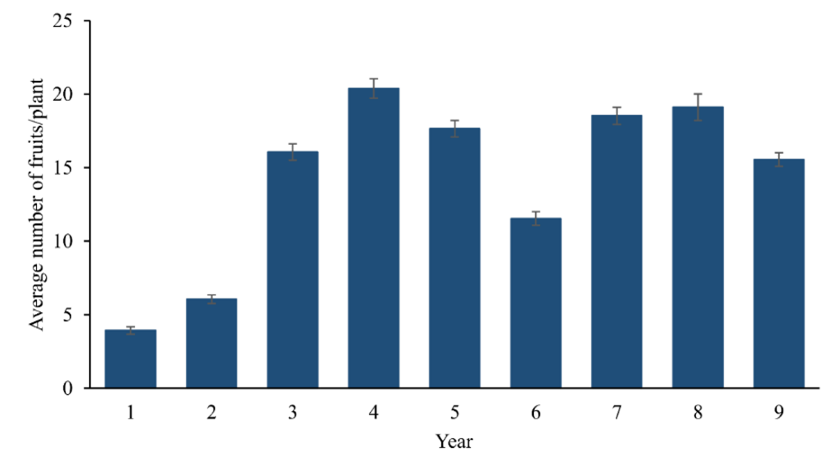
Figure 1
Average and standard error in the number of fruits (NF) produced per plant over nine harvests in a clonal competition trial of T. grandiflorum established in an agroforestry system, in the municipality of Tomé-Açu, State of Pará, Brazil.
In general, we observed a genetic values correlation of median levels between the analysed variables. The greatest correlations were between FW with ShW (0.8) and PW (0.86), demonstrating the influence of these two components on the total fruit weight. Negative correlations were found between NF and ShW (−0,65), FW and NF (−0.67), and FL and ST (−0.51); positive correlations were found between FD and FW (0.78), as well as between ShW (0.77) and PW (0.61). The only non-significant correlation was between the NF and SeW (Figure 2).
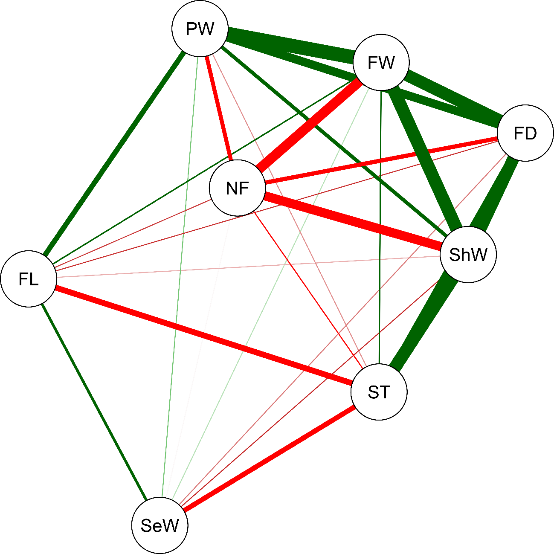
Figure 2
Network of correlations between the genetic values of eight variables1 analyzed for 16 T. grandiflorum clones established in an agroforestry system in the municipality of Tomé-Açu, Pará, Brazil. The green and red lines represent positive and negative significant correlations, respectively. The width of the line represents the intensity of the correlation. 1Average number of fruits per plant (NF), average fruit length (FL) in mm, average fruit diameter (FD) in mm, average shell thickness (ST) in mm, average shell weight per fruit (ShW) in g, average seed weight per fruit (SeW) in g, average pulp weight per fruit (PW) in g, and average fruit weight per plant (FW) in g.
Estimates of the variance components showed the presence of genetic variability among clones for all variables, considering the significance of their genotypic variances, with p ≤ 0.01, according to the deviance analysis. The highest coefficients of both genotypic and residual variation were observed for NF, while for the other characteristics, there was a lower magnitude of these parameters. The total heritability values (hg2) estimated for all variables were considered as median (Table 2).
The predicted genotypic values indicated that different genotypes stood out among the variables. For NF, clones 42 and 44 showed the highest values. On the other hand, for FW, the best results were observed for clones 56 and 174, with the latter also showing the highest value for ShW and ST and the former, a good result for PW. Clones 63 and 61 stood out for FD, while clones 57, 56, and 47 exhibited the highest FL values (Table 3).
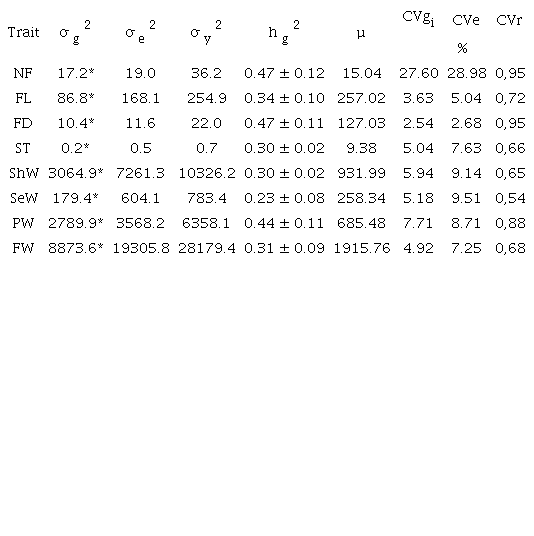
¹σg2: genotypic variance; σe2: residual variance; σy2: individual phenotypic variance; hg2: heritability of total genotypic effects; µ = general average; CVgi = coefficient of genotypic variation; CVe = coefficient of residual variation; CVr = coefficient of relative variation. ²See Figure 2. *Significant at 1%, according to the deviance analysis.
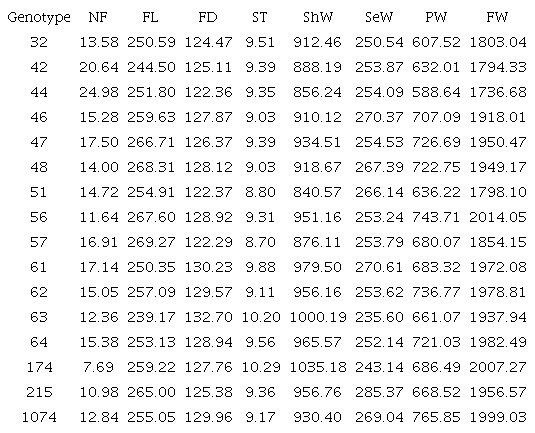
¹See Figure 2.
Using the additive index made possible to rank the best genotypes of the experiment in descending order. The top ten genotypes were 44, 47, 57, 42, 46, 48, 62, 61, 1074, and 64, which together would result in an average gain of 6.53%. However, considering the high incidence of witches’ broom disease (WB) presented by some clones, three (48, 62, and 1074) of the ten top-ranked genotypes were discarded (Table 4).
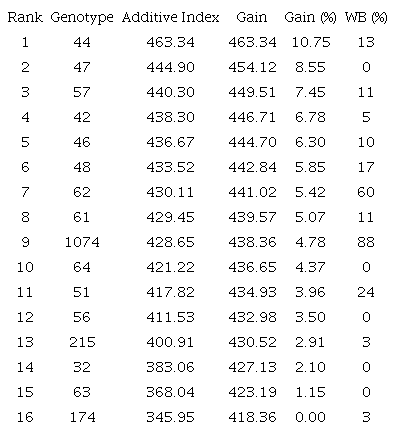
¹See Figure 2.
Thus, genotypes 44, 47, 57, 42, 46, 61, and 64 (43% of the original) were selected for inclusion in the T. grandiflorum breeding program, reflecting an average gain of 7.03%. Of these, the clones 47 and 64 stood out in terms of resistance, as none of the individuals presented signs of witches’ broom disease in over 12 years of field observation.
Discussion
The treatments in this experiment included grafted plants, which showed a tendency to attain stability earlier than the non-grafted seedlings used by Alves et al. (2020), in which the stability was attained after the seventh year. The grafting technique offers advantages, such as uniformity and early production, especially in perennial fruit trees such as T. grandiflorum (Baron, Amaro, Pina, & Ferreira, 2019). Fluctuations in production observed in this experiment during the stable productive phase are common in commercial crops and are attributed to climatic factors, especially long periods of drought after the beginning of fructification, the ecophysiology of the crop, and interactions with pollinating insects (Venturieri, 2011).
In terms of genetic values correlation between the studied characteristics, a relatively high correlation between FD and FW was observed, similar to that reported by Alcoforado, Pedrozo, Mayer, and Lima-Primo (2019), suggesting that this correlation is typical for the species and the existence of gene linkage or epistatic interactions. The greater influence of FD on the total fruit weight (FW) indicates that genotypes that produce fruits with larger diameters should be prioritised over those with greater length. However, other relationships should be considered, such as the positive correlation between FD and ST and the negative correlation between FL and ST.
Fruits with thicker and heavier shells are not suitable for industry because the shell accounts for a greater proportion of the total fruit weight, to the detriment of the percentage of pulp and almonds, which are pillars of the T. grandiflorum production chain. Therefore, FL and FD are equally important in the selection of genotypes, and both are directed to positive in the selection index.
The negative correlation between NF and FW indicates that, depending on the genotype, the partitioning of photoassimilates will prioritise the production of either a greater quantity of small fruits or a smaller number of heavier fruits. This behaviour was also observed for cocoa, regarding the weight of almonds and the number of fruits produced (Doaré, Ribeyre, & Cilas, 2020). Therefore, one of the objectives of breeding programs is to select genotypes that produce a large number of heavy fruits, which leads to greater production of pulp and almonds. For this reason, the selection index prioritised genotypes that performed well for both NF and FW.
The highest values found for CVgi for NF indicate that this characteristic is highly influenced by the heterogeneity of genes that govern it. This influence is comparatively greater than that for the other studied characteristics, which are more homogeneous despite being quantitative. The CVgi is a good indicator of the magnitude of variability available in the population for a given characteristic, even allowing inter-population comparison of this parameter (Silva et al., 2020).
Considering a minimum CVgi value of 10%, as recommended by Resende (2002) for clonal selection, none of the characteristics reached this requirement except for NF. This indicates that individualised selection for these variables is compromised by low levels of variability. However, the use of CVgi as a weight in the selection index enables NF to be considered the main characteristic of commercial importance without discarding the other traits. Furthermore, although the variability is low for these traits, the significance of the deviance analysis shows that it does exist and can be exploited.
The CVr, or the ratio between CVgi and CVe, enables a more direct interpretation of the relationship between genotypic and environmental influences. Values greater or close to unity indicate good conditions for selection and greater gains, leading to better accuracy (Silva, Vidal Neto, & Vale, 2017; Yamamoto et al., 2017). Taking this parameter into consideration, the variables NF (0.95%), FD (0.94%), and PW (0.88%) provided the best results.
Heritability is another parameter of paramount importance that should be considered in the breeding cycle of any species. According to Schmidt, Hartung, Bennewitz, and Piepho (2019), heritability may reflect the precision and usefulness of test results. Taking into account the classification proposed by Resende (2002), the heritability values found herein are median, denoting efficiency in selection and consistency between the predicted gains and the real values.
Compared to the present study, Maia et al. (2011) found slightly lower values for FD, FW, and PW of T. grandiflorum clones, which may be due to the low amplitude of the genetic base of the clones used in their experiment. It should be noted that the genetic structure of the population under study influences the magnitude of heritability. For example, Alves et al. (2020) studied a population with greater variability (25 half-sibling progeny) and found a narrow-sense heritability of 0.65, a value higher than that found here for total heritability.
The variability indicated by the above parameters can be observed in the difference between the genotypic values of the 16 clones, particularly for NF, with a 70% difference between the least productive (174) and the most productive (44) clones. For the other variables, the difference was less pronounced but fluctuated between 25% and 10%. By analysing the behaviour of the genotypes for each characteristic in detail, we can see that direct individual selection would prioritise different genotypes. Therefore, it is pertinent to use a selection index that considers the relative importance of each variable. Setyawan, Taryono, and Mitrowihardjo (2016) emphasized the need to develop selection indexes for cocoa that gather all the variables that characterize a productive genotype. Mustiga et al. (2018) and Jaimez, Vera, Mora, Loor, and Bailey (2020) also concluded that the use of selection indices can be more advantageous than direct selection, especially when characteristics of different natures are considered, such as vegetative development, morpho-agronomic fruit characteristics, productivity, and disease resistance.
In the present experiment, the additive index enabled us to rank the 16 studied genotypes considering all of the eight analysed variables (Table 4), from which we identified the best seven. Alves et al. (2019) demonstrated the efficiency of the additive index for J. curcas L., with an average gain of 27%, and highlighted the possibility of using this method for other perennial species. Pereira et al. (2019) also emphasised the ability of this index to distinguish genotypes that have superior characteristics, as shown through its use with C. papaya L., another fruit tree.
The analysis of the productive characteristics of T. grandiflorum should not be disconnected from the resistance of the genotypes to M. perniciosa. The pathogen is one of the main challenges for the cultivation of T. grandiflorum and T. cacao in Brazil (Mournet et al., 2020). Genetic improvement through the development of resistant cultivars is the most cost-effective and ecologically sustainable option to combat the effects of this fungus. Therefore, we individualised the analysis of this variable due to its biological and economic significance. Although ten clones were initially selected, three showed relatively high rates of infection with the fungus. Therefore, these three were excluded as susceptible individuals in an orchard can increase the pathogen pressure on resistant individuals, culminating in the breakdown of their resistance (Albuquerque et al., 2010). These seven selected clones will be used for future hybridization, along with other resistant sources, to facilitate the pyramidization of these genes to avoid, or at least delay, the weakening of the resistance of the cultivars (Crété, Pires, Barbetti, & Renton, 2020).
Conclusion
We identified seven genotypes (44, 47, 57, 42, 46, 61, and 64) exhibiting significant potential in terms of productivity and low incidence rates of the witches’ broom disease. These genotypes are also beneficial to obtain fruits with desirable morpho-agronomic characteristics and are adapted to the conditions of interspecific competition. As such, we recommend the use of these genotypes as cultivars in agroforestry systems and their inclusion in the T. grandiflorum breeding programs for future hybridization.
References
Albuquerque, P. S. B., Silva, S. D. V. M., Luz, E. D. M. N., Pires, J. L., Vieira, A. M. C., Demétrio, C. G. B., ... Figueira, A. (2010). Novel sources of witches’ broom resistance (causal agent Moniliophthora perniciosa) from natural populations of Theobroma cacao from the Brazilian Amazon. Euphytica, 172(1), 125-138. DOI: https://doi.org/10.1007/s10681-009-0068-4
Alcoforado, A. T., Pedrozo, C. A., Mayer, M. M., & Lima-Primo, H. E. (2019). Repeatability of morpho-agronomic characters of Theobroma grandiflorum fruits. Revista Brasileira de Fruticultura, 41(2), 1-7. DOI: https://doi.org/10.1590/0100-29452019142
Alves, R. M., Sebbenn, A. M., Artero, A. S., Clement, C., & Figueira, A. (2007). High levels of genetic divergence and inbreeding in populations of cupuassu (Theobroma grandiflorum). Tree Genetics & Genomes, 3(4), 289-298. DOI: https://doi.org/10.1007/s11295-006-0066-9
Alves, R. M. (2012). Implantação de um pomar de cupuaçuzeiro com a cultivar BRS Carimbó. Brasília, DF: Embrapa.
Alves, R. M., Chaves, S. F. S., Gama, M. A. P., Pedroza Neto, J. L., & Santos, T. G. D. (2020). Simultaneous selection of cupuassu tree and Brazilian mahogany genotypes in an agroforestry system in Pará state, Brazil. Acta Amazonica, 50(3), 183-191. DOI: https://doi.org/10.1590/1809-4392202000711
Alves, R. S., Teodoro, P. E., Peixoto, L. A., Silva, L. A., Laviola, B. G., Resende, M. D. V., & Bhering, L. L. (2019). Multiple-trait BLUP in longitudinal data analysis on Jatropha curcas breeding for bioenergy. Industrial Crops and Products, 130, 558-561. DOI: https://doi.org/10.1016/j.indcrop.2018.12.019
Baron, D., Amaro, A. C. E., Pina, A., & Ferreira, G. (2019). An overview of grafting re-establishment in woody fruit species. Scientia Horticulturae, 243, 84-91. DOI: https://doi.org/10.1016/j.scienta.2018.08.012
Bolfe, E. D., & Batistella, M. (2011). Análise florística e estrutural de sistemas silviagrícolas em Tomé-Açu, Pará. Pesquisa Agropecuária Brasileira, 46(10), 1139-1147. DOI: https://doi.org/10.1590/S0100-204X2011001000004
Céron-Rojas, J. J., & Crossa, J. (2018). Linear selection indices in modern plant breeding. Cham, SW: Springer Nature. DOI: https://doi.org/10.1007/978-3-319-91223-3
Costa, R. S., Pinheiro, W. B. S., Arruda, M. S. P., Costa, C. E. F., Converti, A., Costa, R. M. R., & Silva Júnior, J. O. C. (2020). Thermoanalytical and phytochemical study of the cupuassu (Theobroma grandiflorum Schum.) seed by-product in different processing stages. Journal of Thermal Analysis and Calorimetry, 147, 275-284. DOI: https://doi.org/10.1007/s10973-020-10347-0
Crété, R., Pires, R. N., Barbetti, M. J., & Renton, M. (2020). Rotating and stacking genes can improve crop resistance durability while potentially selecting highly virulent pathogen strains. Scientific Reports, 10(1), 1-18. DOI: https://doi.org/10.1038/s41598-020-76788-7
Doaré, F., Ribeyre, F., & Cilas, C. (2020). Genetic and environmental links between traits of cocoa beans and pods clarify the phenotyping processes to be implemented. Scientific Reports, 10(1), 1-6. DOI: https://doi.org/10.1038/s41598-020-66969-9
Epskamp, S., Cramer, A. O., Waldorp, L. J., Schmittmann, V. D., & Borsboom, D. (2012). qgraph: Network visualizations of relationships in psychometric data. Journal of Statistical Software, 48(4), 1-18. DOI: https://doi.org/10.18637/jss.v048.i04
Faria, J. V., Valido, I. H., Paz, W. H., Silva, F. M., Souza, A. D., Acho, L. R., ... Bataglion, G. A. (2020). Comparative evaluation of chemical composition and biological activities of tropical fruits consumed in Manaus, central Amazonia, Brazil. Food Research International, 139, 109836. DOI: https://doi.org/10.1016/j.foodres.2020.109836
Genovese, M. I., & Lannes, S. C. D. S. (2009). Comparison of total phenolic content and antiradical capacity of powders and “chocolates" from cocoa and cupuassu. Food Science and Technology, 29(4), 810-814. DOI: https://doi.org/10.1590/S0101-20612009000400017
Homma, A. K. O. (2014). Setenta anos de pesquisa agropecuária na Amazônia: contribuições da Embrapa para fruticultura tropical. In A. K. O. Homma (Ed.), Extrativismo vegetal na Amazônia: história, ecologia, economia e domesticação (p. 377-403). Belém, PA: Embrapa Amazonia Oriental.
Jaimez, R. E., Vera, D. I., Mora, A., Loor, R. G., & Bailey, B. A. (2020). A disease and production index (DPI) for selection of cacao (Theobroma cacao) clones highly productive and tolerant to pod rot diseases. Plant Pathology, 69(4), 698-712. DOI: https://doi.org/10.1111/ppa.13156
Jiménez-Muñoz, J. C., Mattar, C., Barichivich, J., Santamaría-Artigas, A., Takahashi, K., Malhi, Y., ... Van Der Schrier, G. (2016). Record-breaking warming and extreme drought in the Amazon rainforest during the course of El Niño 2015-2016. Scientific Reports, 6(1), 1-7. DOI: https://doi.org/10.1038/srep33130
Lima, A. K. O., Vasconcelos, A. A., Sousa Júnior, J. J. V., Escher, S. K. S., Nakazato, G., & Taube Júnior, P. S. (2019). Green synthesis of silver nanoparticles using Amazon fruits. International Journal of Nanoscience and Nanotechnology, 15(3), 179-188.
Maia, M. C. C., Resende, M. D. V., Oliveira, L. C., Álvares, V. S., Maciel, V. T., & Lima, A. C. (2011). Seleção de clones experimentais de cupuaçu para características agroindustriais via modelos mistos. Revista Agro@mbiente On-line, 5(1), 35-43. DOI: https://doi.org/10.18227/1982-8470ragro.v5i1.388
Mournet, P., Albuquerque, P. S. B., Alves, R. M., Silva-Werneck, J. O., Rivallan, R., Marcellino, L. H., & Clément, D. (2020). A reference high-density genetic map of Theobroma grandiflorum (Willd. ex Spreng) and QTL detection for resistance to witches’ broom disease (Moniliophthora perniciosa). Tree Genetics & Genomes, 16(6), 89. DOI: https://doi.org/10.1007/s11295-020-01479-3
Meinhardt, L. W., Rincones, J., Bailey, B. A., Aime, M. C., Griffith, G. W., Zhang, D., & Pereira, G. A. (2008). Moniliophthora perniciosa, the causal agent of witches’ broom disease of cacao: what's new from this old foe? Molecular Plant Pathology, 9(5), 577-588. DOI: https://doi.org/10.1111/j.1364-3703.2008.00496.x
Mustiga, G. M., Gezan, S. A., Phillips-Mora, W., Arciniegas-Leal, A., Mata-Quirós, A., & Motamayor, J. C. (2018). Phenotypic description of Theobroma cacao L. for yield and vigor traits from 34 hybrid families in Costa Rica based on the genetic basis of the parental population. Frontiers in Plant Science, 9(808), 1-17. DOI: https://doi.org/10.3389/fpls.2018.00808
Pará. (2020). Secretaria de Estado de Desenvolvimento Agropecuário e da Pesca. Indicadores agropecuários. Retrieved on Dec. 08, 2020 from 08, 2020 from http://www.sedap.pa.gov.br/content/cupua%C3%A7u
Pereira, A. L., Abreu, V. K., & Rodrigues, S. (2018). Cupuassu-Theobroma grandiflorum. In S. Rodrigues, E. O. Silva, & E. Brito (Eds.), Exotic fruits (p. 159-162). Amsterdam, NT: Elsevier Inc.
Pereira, M. G., Poltronieri, T. P. S., Pereira, T. N. S., Ramos, H. C. C., Catarina, R. S., Vettorazzi, J. C. F., ... Ferregueit, G. A. (2019). Twenty-two-year papaya breeding program: from breeding strategy establishment to cultivar development. Functional Plant Breeding Journal, 1(2), 9-27. DOI:https://doi.org/10.35418/2526-4117/v1n2a2
R Core Team. (2020). R: A language and environment for statistical computing. Vienna, AT: R Foundation for Statistical Computing. Retrieved on Dec. 08, 2020 from 08, 2020 from https://www.R-project.org/.
Resende, M. D. V. (2002). Genética biométrica e estatística no melhoramento de plantas perenes. Brasília, DF: Embrapa Informação Tecnológica.
Resende, M. D. V., Silva, F. F., & Azevedo, C. F. (2014). Estatística matemática, biométrica e computacional: Modelos mistos, multivariados, categóricos e generalizados (REML/BLUP), inferência bayesiana, regressão aleatória, seleção genômica, QTL, GWAS, estatística espacial e temporal, competição, sobrevivência. Viçosa, MG: Editora UFV.
Resende, M. D. V. (2016). Software Selegen-REML/BLUP: a useful tool for plant breeding. Crop Breeding and Applied Biotechnology, 16(4), 330-339. DOI: https://doi.org/10.1590/1984-70332016v16n4a49
Salgado, J. M., Rodrigues, B. S., Donado-Pestana, C. M., Dias, C. T. S., & Morzelle, M. C. (2011). Cupuassu (Theobroma grandiflorum) peel as potential source of dietary fiber and phytochemicals in whole-bread preparations. Plant Foods for Human Nutrition, 66(4), 384-390. DOI: https://doi.org/10.1007/s11130-011-0254-0
Schmidt, P., Hartung, J., Bennewitz, J., & Piepho, H. P. (2019). Heritability in plant breeding on a genotype-difference basis. Genetics, 212(4), 991-1008. DOI: https://doi.org/10.1534/genetics.119.302134
Setyawan, B., Taryono, T., & Mitrowihardjo, S. (2016). Determination of selection index of cocoa (Theobroma cacao L.) yield traits using regression methods. Pelita Perkebunan, 32(2), 101-108. DOI: https://doi.org/10.22302/iccri.jur.pelitaperkebunan.v32i2.229
Silva, A. R., Malafaia, G., & Menezes, I. P. P. (2017). Biotools: an R function to predict spatial gene diversity via an individual-based approach. Genetics and Molecular Research, 16(2), 1-6. DOI: https://doi.org/10.4238/gmr16029655
Silva, T. P. D., Vidal Neto, F. D. C., & Vale, J. C. (2017). Prediction of genetic gains with selection between and within S2 progenies of papaya using the REML/Blup analysis. Pesquisa Agropecuária Brasileira, 52(12), 1167-1177. DOI: https://doi.org/10.1590/s0100-204x2017001200005
Silva, J. O. C., Bruckner, C. H., Carneiro, P. C. S., Resende, M. D. V., Alves, R. S., & Silva, D. F. P. (2020). Estimates of genetic parameters, genetic variability, and selection in the S1 generation of peach. Ciência Rural, 50(10), 1-7. DOI: https://doi.org/10.1590/0103-8478cr20190976
Souza, A. G. C. (2007). Boas práticas agrícolas da cultura do cupuaçuzeiro. Manaus, AM: Embrapa Amazônia Ocidental.
Venturieri, G. A. (2011). Flowering levels, harvest season and yields of cupuassu (Theobroma grandiflorum). Acta Amazonica, 41(1), 143-152. DOI: https://doi.org/10.1590/S0044-59672011000100017
Yamamoto, E. L. M., Holanda, I. S. A., Morais, P. L. D., Nunes, G. H. S., Antonio, R. P., Lemos, M. S., & Silva, J. R. (2017). Selection of umbu-cajazeira clones using the REML/BLUP. Revista Brasileira de Ciências Agrárias, 12(4), 496-502. DOI: https://doi.org/10.5039/agraria.v12i4a5485
Author notes
* Author for correspondence. E-mail: saulo.chaves@ufv.br
