Biotecnologia
Extra and intracellular laccase titers of xylophagic bacteria isolated from adult termite
Extra and intracellular laccase titers of xylophagic bacteria isolated from adult termite
Acta Scientiarum. Technology, vol. 43, e51805, 2021
Universidade Estadual de Maringá
Recepción: 14 Enero 2020
Aprobación: 30 Mayo 2020
Abstract: Laccase is an important enzyme in terms of its versatile applicability, but its commercial use is limited by factors such as high production cost, low activity and/or stability under given conditions. The objective of this study was to screen xylophagic bacteria isolated from termites for the production of extracellular and intracellular laccases. Six laccase-positive strains were isolated, namely CA, A3, A5, A6, A7 and A8. They were molecularly identified by sequence analysis of 16S rRNA and classified under the genera Bacillus (A7, A8, CA) and Pseudomonas (A3, A5, A6). Laccase was produced by these bacterial isolates by submerged fermentation and was optimized at 37°C, pH 5.5, 6.2 and 7.0, with agitation and 0.5 mM guaiacol (as carbon source). Laccase activity was determined by measuring the oxidation of guaiacol and ABTS (2,2.-azino bis[3-ethylbenzthiazoline-6-sulfonate]). Strain A5 produced extracellular laccase titers ranging from 123 to 168 U ml-1. Guaiacol was identified as a better substrate for the quantification of laccase. In conclusion, bacteria harboring the gut of termites can produce extracellular laccase with activity at medium to moderate acidity.
Keywords: kraft lignin, extracellular laccase, Bacillus. Pseudomonas spp.
Introduction
Laccases are ubiquitous enzymes found commonly in plants, fungi and bacteria. They possess several biological functions including degradation of complex polymers (lignin, humic acid), lignification, and bioremediation, among others (Strong, 2011). They have been extensively exploited for biotechnological applications such as biobleaching, bioremediation of xenobiotics, decolorization of textile dyes industrial effluents, biosensors, food industry, and degradation of plastics (Du et al., 2015; Shraddha, Simran, Mohit, & Ajay, 2011).
Bacterial production of laccase has been reported in several genera. They are produced by gram-negative bacteria such as Pseudomonas, Enterobacter, Delfia, Proteobacterium and Alteromonas (Neifar, Chouchane, Mahjoubi, Jaouani, & Cherif, 2016) and gram-positive bacteria such as Bacillus, Geobacillus, Streptomyces, Rhodococcus, Staphylococcus, Azospirillum and Lisinibacillus (Margot et al., 2013; Muthukumarasamy, Jackson, Raj, & Sevanan, 2015; Narayanan, Murugan, Eva, Devina, & Kalidass, 2015; Sondhi et al., 2015; Wang, Zhao, Lu, Wei, & Li, 2011; Desai, 2017). Bacterial laccases have not yet been exploited on an industrial scale as most of the reported laccases are localized intracellularly, or are spore bound, which makes their production and purification in large quantities extremely difficult. However, analysis of several bacterial genomes have shown that 76 % of putative laccase genes carry the signal sequences, which could enable them to be secreted outside the cell (Ausec, Zakrzewski, Goesmann, Schluter, & Mandic-Mulec, 2011).
Most industrial operations for enzyme production are carried out under conditions of high temperature, pH and high salt concentrations. Bacterial laccases are known to possess several properties that are not characteristic of fungal laccases, such as stability at high temperatures (30-85°C) and pH (3.0-9.0), or tolerance to high concentrations of salts, e.g., > 1 M NaCl (Muthukumarasamy et al., 2015; Sondhi et al., 2015).
In the past decade, bacterial laccases have attracted more attention for their potential in biodegrading environmentally significant phenolic pollutants (Hildén, Hakala, & Lundell, 2009). The discovery of new laccases from different environmental niches and with different substrate specificities, besides action at different pH values, could be important and amenable for industrial applications.
One biological niche for screening of laccase-producing bateria is the gut of termites (family Termitidae), which harbors various xylophagic bacteria capable of digesting the components of wood (cellulose, hemicellulose, lignin). As these termites feed mainly on wood , it is possible that microbes capable of degrading complex organic matter may be found in their gut. These bacteria can be exploited for enzymes that digest lignin, a natural aromatic biopolymer. Bacteria from termites are known to produce laccases (Gonzalo, Colpa, Habib, Fraaije, 2016; Xu et al., 2018).
Bearing this aspect in mind, we isolated novel lignin-degrading bacteria present in xylophagic insects such as termites. Local termites of the genus Heterotermes were screened for bacteria having the ability to oxidize laccase substrates such as ABTS (2,21-azino bis[3-ethylbenzthiazoline-6-sulfonate]) and guaiacol (2-methoxyphenol). Herein, we report six laccase-positive bacterial strains isolated from termites that were molecularly identified by sequence analysis of the 16S rRNA gene, and classified under the genera Bacillus and Pseudomonas.
Material and methods
Termite collection
Adult Heterotermes sp. (order Isoptera; family Termitidae) colonies were collected from decaying wood in the northern region of the State of Paraná in the South of Brazil, and maintained in sealed plastic boxes. Ten adult termites were sterilized in 70% ethanol (10 min.), rinsed in sterile distilled water, air-dried by shaking for several seconds under aseptic conditions. The termites were next placed in a sterile microcentrifuge tube and aseptically macerated in 10 mM sodium phosphate-buffered saline (PBS, pH 6.0) with the aid of a sterile plastic pestle.
Isolation and plate assay screening for laccase-producing bacteria
The disrupted insect debris (homogenate) obtained was serially diluted and plated on Nutrient Agar (NA) medium (HiMedia Laboratories Pvt. Ltd, Mumbai, India) containing 0.2 mM CuSO4.5H2O, 1% (v v-1) Tween 20, and 1% (w/v) Kraft lignin (KL, extracted from Kraft pulped softwood black liquor by acid precipitation). The plates were incubated for 48 h at 37ºC.
Bacteria from individual colony-forming units (CFUs) were randomly selected and transferred to NA-agar medium plates containing 2 mM guaiacol, and in two sets of experiments the pH was adjusted to pH 5.5 and 7.0, and the plates incubated at 37°C for 24h. Laccase-producing bacterial isolates were examined for color changes in the agar (Strong, 2011). Bacteria producing laccases showed brown zones around the growing colonies as a consequence of oxidation of guaiacol.
Identification of bacterial isolates
The bacterial isolates were identified morphologically, and also by molecular identification through sequencing the 16S rDNA gene. The colour, shape, size, nature and pigmentation of the colonies were assessed to identify the strains morphologically. Gram-staining characteristics and endospores were examined microscopically. The strains isolated were determined according to the methods described in Bergey’s Manual of Determinative Bacteriology (Buchanan & Gibbons, 1974).
Preparation of bacterial extracellular and intracellular fractions for assaying laccase activity
The selected bacterial colonies were cultured in NA broth containing 2 mM of guaiacol at pH adjusted to 6.0, and the cultures incubated at 37°C on a rotary shaker (180 rpm) for 24h. The bacterial cultures were then centrifuged (9,000 x g) for 20 min at 4°C, and the supernatant collected and filter sterilized (cellulose ester membrane of 0.22 μm; Millipore; Sigma-Aldrich, St Louis, MO, USA), and was used as the source of the crude extracellular enzyme (CEE). Catalase solution (10 µl; 1.000 U mL-1 -Sigma-Aldrich, USA) was added to 200 µl of CEE solution and incubated for 1 h at 37°C to remove any H2O2 produced by the bacteria that could possibly interfere with the laccase assay procedure.
The bacterial cells recovered as a pellet following centrifugation were washed twice with ultra-pure water and resuspended in 5.0 ml of phosphate buffer (10mM, pH 7). The cells were disrupted by sonication (five cycles each of 120 s) in an ice bath, and the lysed cells centrifuged (9,000 x g for 20 min.). The supernatant was recovered, and is the source of intracellular enzyme (IE). Cell lysis was confirmed by seeding the pellet obtained by centrifugation on NA plates, and examining for growth.
Laccase assay and determination of the optimum pH
The pH versus the laccase activity profile was evaluated by varying the pH of reaction mixtures using two buffer systems (0.1 M sodium phosphate buffer pH 6.2 and 7.0; 0.05 M sodium citrate buffer, pH 5.0). Laccase was assayed against the substrates guaiacol (2 mM) and ABTS (2.5 mM). The assay mixture contained 100 μl of CEE, 100 μl of guaiacol or ABTS solution, and 800 μl of each buffer in a final volume of 1.0 ml, and was incubated at room temperature. The oxidized products were determined spectrophotometrically by monitoring the increase in absorbance at 520 and 420 nm, for the respective laccase substrates.
One unit of a laccase activity is described as the required amount of enzyme to oxidize 1 μmol of guaiacol (ε: 48,000 M-1 cm-1) or ABTS (ε: 36,000 M-1 cm-1) per min under the assay conditions (Leonowicz & Grzywnowicz, 1981).
Total DNA extraction
Bacterial DNA was extracted using the boiling technique. Bacteria exhibiting color changes in the plate-assay screening procedure were inoculated into 3 mL of Brain Heart Infusion (BHI) (HiMedia Laboratories Pvt. Ltd, Mumbai, India), and incubated at 37°C for 18h while stirring. After growth, the samples were centrifuged (11,200 x g for 20 min.), and the supernatant discarded. The cell pellet recovered was added to sterile deionized water (100 μL) and the samples heated at 100°C for 30 min., followed by their immersion in an ice bath for 3 min. The samples were again centrifuged, and 50 μl of the supernatant containing the extracted DNA was collected.
The 16S rDNA analysis
Universal primer F27 (5’AGAGTTTGATCCTGGCTCAG3’) and R1492 (5’GGTTACCTTGTTACGACTT3’) were used in PCR experiments for sequencing of the 16S rDNA gene. The PCR reaction conditions consisted of an initial denaturation step at 94°C for 10 min., and 30 cycles of denaturation at 94°C each for 1 min., annealing at 45°C for 1 min., extension at 72°C for 2 min., and a final extension cycle at 72°C for 10 min. The PCR product was sequenced using a Ludwig Biotecnologia (Brasil) sequencer. The generated gene sequences were compared with sequences available in GenBank by using the BLASTn program (Basic Local Alignment Search Tools)[1] (Altschul, Gish, Miller, Myers, & Lipman, 1990). The sequences were aligned using ClustalW and MEGA 5.0 software was used for construction of phylogenetic tree.
Statistical analysis
Data analysis were subjected to one way ANOVA using PROC ANOVA (SAS version 9.3) and comparisons were made using Tukey’s post-hoc significance test at p value of less than 0.05. All the experiments were carried out in triplicates.
Results and discussion
In the first stage of this study 15 random bacterial colonies producing laccases were isolated from adult termites by plate assay screening. Their characteristics, colonial morphology, borders and texture were examined. Upon Gram staining, 11 isolates were found to be bacilli and 4 were cocci, and they were predominantly gram-positive (Table 1).
These isolates showed growth on two conditions of acidification of the culture medium (pH 5.5 and 7.0). Of these, 6 colonies, denoted as CA, A3, A5, A6, A7 and A8, showed whitish to brownish colored zones around them, indicating that the substrate guaiacol was oxidized. This suggested laccase production. Hence, these colonies were subjected to further screening (Figure 2 A) to evaluate laccase production in submerged liquid medium. NA culture medium supplemented with CuSO4 (0.2 mM), Tween 20 (1%) and guaiacol was found to be more appropriate for screening and assay of laccase activity. Copper acts both as an inducer and as a micronutrient and has the potential to raise the laccase titers considerably (Bakkiyaraj, Aravindan, Arrivukkarasan, & Viruthagiri, 2013). Similar media has been used by other researchers (Shah & Jobanputra, 2017) for screening of laccase-producing bacteria from diverse environments. Guaiacol is also a known inducer of laccases in bacteria (Mongkolthanaruk, Tongbopit, & Bhoonobtong, 2012).

+ and – indicates whether the bacterial strains supported growth at acid and neutral pH.
The morphological characteristics of the bacterial strains (Table 1) was established according to Bergey’s Manual of Systematic Bacteriology (Buchanan & Gibbons, 1974). The isolated bacteria were identified as Bacillus (A7, A8 and CA) and Pseudomonas (A3, A5 and A6) by 16S rDNA sequence submitted in Genban. A phylogenetic tree was constructed using the ClustalW distance tree method as shown in Figure 1. The 16S rDNA gene sequence analysis method was not sufficient to identify the isolated strains at the species level. However, it is known that isolates belonging to the same genus exhibiting less than 97% similarity in the 16S rRNA gene sequence should be considered as members of the different species (Stackebrandt & Goebel, 1994).
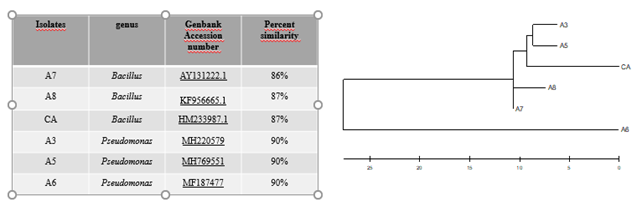
Figure 1. Identification of bacterial isolates and phylogenetic tree Using 16S rRNA gene sequencing BLAST analysis found in this study.
Besides, both the genera have been well-described as laccase producers. Several reports demonstrated the Bacillus (Lončar, Gligorijević, Božić, & Vujčić, 2014; Mishra & Srivastava, 2016) and Pseudomonas (Chauhan & Jha, 2018; Peter & Vandana, 2014) are producers of ligninolytic enzymes.
In this study, we tested the activity of laccase at three different pH values - 5.0, 6.2 and 7.0. We used two substrates for quantification of laccase by spectrophotometry assays (Figure 2 B). All the strains could produce extracellular and intracellular laccases, as measured on fractions CEE and IE, and enzyme activity was expressed as U mL-1.
The results showed that the best substrate for quantification of laccase was guaiacol. With this substrate, no significant difference was observed between enzyme stability at the pH tested, indicating that the bacterial laccase was stable at low to moderate acidity. No difference (p < 0.05) in laccase activity was observed between the different substrates. However, the results were quite promising, with values ranging from a minimum of 123 to a maximum of 168 U mL-1 (A5 isolate at pH 6.2) when guaiacol was used as substrate.
Bacterial laccases have been reported to be either intracellularly localized, or as a component of bacterial spore coat protein in Bacillus spp. (Ausec et al., 2011). In the present study, however, all isolates analyzed were found to produce extracellular laccase, as CEE was used in the experiments. Different techniques like spectrophotometry, voltammetry and fluorometry can be used to assay laccase activity. The spectrophotometric method, however, is more desirable because it allows use of suitable organic substrates that can detect laccase activity. A literature survey has revealed various organic compounds, including guaiacol and ABTS, that have been used for assaying laccase activity (Moshtaghioun et al., 2011).
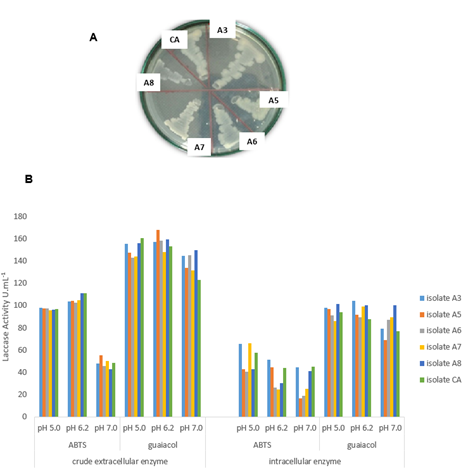
Figure 2. Coloration of the bacterial colonies isolated from termites (whitish to brownish) on nutrient agar medium containing 2 mM of guaiacol (A). Effect of pH on laccase activity of the isolated bacterial strains. Laccase activity was measured as U mL-1 using ABTS and guaiacol as substrates at pH 5.0, 6.2 and 7.0; laccase activity of the crude extracellular and intracellular enzyme (B). All the experiments were carried out in triplicates.
In our study, guaiacol broth was used for laccase production as suggested by Sharma, Goel and Capalash (2007). Although the ABTS method is more sensitive, the results with guaiacol were in accordance with some literature reports (Robles, Lucas, Cienfuegos, & Gálvez, 2000). Our results clearly showed that bacterial laccase activity assayed using guaiacol as substrate was higher than ABTS. This contradicts the results reported by Liers, Ullrich, Pecyna, Schlosser and Hofrichter (2007) for a purified laccase from an ascomycete, and for a laccase from a basidiomycete. However, this in agreement with other studies on B. subtilis laccase (Sheikhi, Ardakani, Enayatizamir, & Rodriguez-Couto, 2012).
The isolates of Bacillus sp. and Pseudomonas sp. from termites showed high laccase activity when assayed on both substrates; however, we found difficulties in comparing our data with the literature findings, since different studies used different units of measurement for laccase activity. It is worth mentioning that the production and activity of laccases is affected by several factors, including medium pH and composition, time and temperature of microbial growth, and carbon and nitrogen ratios (Lorenzo, Moldes, Couto, & Sanromán, 2002).
We analyzed CEE and IE fractions for laccase activity when the isolates were cultured on a medium containing guaiacol as an inducer. The resulting enzyme activities were 123 to 168 U mL-1 for extracellular laccase, which was consistent with the findings of Muthukumarasamy et al. (2015), who reported up to 230 U mL-1 of laccase activity from B. subtilis. With agroindustrial residues as a substrate in the nutrient medium, some authors reported laccase activity of 2.28 U L-1 (ABTS) and 1.6 U L-1 (guaiacol) by B. subtilis (Sheikhi et al., 2012). Strong (2011) stated that the addition of inducer prior to inoculation was more effective in inducing laccase compared to the addition of inducer during the fermentation run at timepoint 0h.
The intracellular laccase activity showed large variations depending upon the isolate and did not show much uniformity in enzyme production The laccase activity measured intracellularly is possibly a measure of the enzyme that has not yet been excreted (de novo synthesis).
In bacteria, most of the laccases studied so far are mainly localized intracellularly (Diamantidis, Effosse, Potier, & Bally, 2000), and show cytochrome C oxidase activity and quinone resistance (Sharma et al,. 2007). Cytochrome C peroxidase might play an important role in the lignin degradative process (Zhu et al., 2017).
Bacterial laccase of the genus Bacillus was first reported by Claus and Filip (1997). Since then, more bacterial laccases that can oxidize ABTS and guaiacol have been discovered (Sheikhi et al., 2012). Rajeswari and Bhuvaneswari (2016) reported high laccase production by Bacillus sp. isolated from the environment. In addition, several Kraft lignin-degrading Bacillus spp. have been isolated (Bandounas, Wierckx, Winde, & Ruijssenaars, 2011; Chandra, Raj, Purohit, & Kapley, 2007).
Besides being extracellular, other properties of enzymes such as stability at different pH and temperature values are required for the enzyme to be exploited industrially. Sondhi et al (2015) reported an extracellular, active, acid-stable to alkali-stable laccase from Bacillus tequilensis. The pH strongly affects the enzymatic reactions and is sensitive to hydrogen ion concentration present in the medium across the cell membrane. The optimum pH for production of laccase by Trametes and Rhizoctonia fungi was found to be 7.0 (Murugesan, Dhamija, Nam, Kim, & Chang, 2007).
In the present study, pH 5.0 and 6.2 were found suitable for CEE laccase produced by Bacillus sp. and Pseudomonas sp., when ABTS and guaiacol were used as substrates. The laccase produced by Bacillus sp. and Pseudomonas sp. was extracellular, as quantified in the CEE fraction. Despite having low concentration, the enzyme showed high laccase activity and enzyme titers. In addition, the extracellular laccase preparation appeared to be stable at moderate to low pH, suggesting that it may be a useful candidate for commercial applications.
Conclusion
In the present study, six Kraft lignin-degrading bacteria were isolated from adult termites on the basis of their laccase activities towards ABTS and guaiacol as substrates. The Bacillus and Pseudomonas genera were predominant, and produced laccase at several pH. A study on bacterial laccases is important from the perspectives of basic science as well as for the development of novel biotechnological applications.
Therefore, keeping in view the importance of laccase in bioremediation of industrial effluents and wastewater treatment, bioprospecting new bacterial strains displaying laccase activity are the need of time.
Acknowledgements
This work was supported by the Universidade Tecnológica Federal do Paraná (UTFPR), Fundação Araucária/Governo do Paraná – Brazil, CNPq-Brazil, and the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES) - Finance Code 001.
References
Altschul, S. F., Gish, W., Miller, W., Myers, E.W., & Lipman, D. J. (1990). Basic local alignment search tool. Journal of Molecular Biology, 215, 403-410. doi: 10.1016/S0022-2836(05)80360-2
Ausec, L., Zakrzewski, M., Goesmann, A., Schluter, A., & Mandic-Mulec, I. (2011). Bioinformatic analysis reveals high diversity of bacterial genes for laccase-like enzymes. PLoS One, 6, e25724. doi: 10.1371/journal.pone.0025724
Bakkiyaraj, S., Aravindan, R., Arrivukkarasan, S., & Viruthagiri, T. (2013). Enhanced laccase production by Tramates hirusta using wheat bran under submerged fermentation. International Journal of ChemTech. Research, 5(3), 1224-1238.
Bandounas, L., Wierckx, N. J. P., Winde, J. H. D., & Ruijssenaars, H. J. (2011). Isolation and characterization of novel bacterial strains exhibiting ligninolytic potential. BMC Biotechnology, 11, 1-11. doi: 10.1186/1472-6750-11-94
Buchanan, R. E., & Gibbons, N. E. (1974). Bergey's manual of determinative bacteriology (8th ed.). Baltimore, MD: Williams and Wilkins Co.
Chandra, R., Raj, A., Purohit, H. J., & Kapley, A. (2007). Characterisation and optimisation of three potential aerobic bacterial strains for kraft lignin degradation from pulp paper waste. Chemosphere, 67, 839-846. doi: 10.1016/j.chemosphere.2006.10.011
Chauhan, P. S., & Jha, B. (2018). Pilot scale production of extracellular thermo‐alkali stable laccase from Pseudomonas sp. S2 using agro waste and its application in organophosphorous pesticides degradation. Journal of Chemical Technology & Biotechnology, 93. 1022-1030. doi: 10.1002/jctb.5454
Claus, H., & Filip, Z. (1997). The evidence of a laccase-like enzyme activity in a Bacillus sphaericus strain. Microbiological Research, 152, 209-216. doi: 10.1016/S0944-5013(97)80014-6
Desai, S. A. (2017). Isolation and characterization of Laccase producing bacteria from contaminated sites. Bioscience Discovery, 8, 567-573.
Diamantidis, G., Effosse, A., Potier, P., & Bally, R. (2000). Purification and characterization of the first bacterial laccase in the rhizospheric bacterium Azospirillum lipoferum. Soil Biology and Biochemistru, 32(7), 919-927. doi: 10.1016/S0038-0717(99)00221-7
Du, W., Sun, C., Liang, J., Han, Y., Yu, J., & Liang, Z. (2015). Improvement of laccase production and its characterization by mutagenesis. Journal of Food Biochemistry, 39, 101-108. doi: 10.1111/jfbc.12111
Gonzalo, G., Colpa, D. I., Habib, M. H. M., Fraaije, M. W. (2016). Bacterial enzymes involved in lignin degradation. Journal of Biotechnology, 236, 110-119. doi: 10.1016/j.jbiotec.2016.08.011
Hildén, K., Hakala, T. K., & Lundell, T. (2009). Thermotolerant and thermostable laccases. Biotechnology Lett, 31, 1117-1128. doi: 10.1007/s10529-009-9998-0.
Leonowicz, A., & Grzywnowicz, K. (1981). Quantitative estimation of laccase forms in some white-rot fungi using syringaldazine as a substrate. Enzyme Microbial Technology, 3, 55-58. doi: 10.1016/0141-0229(81)90036-3
Liers, C., Ullrich, R., Pecyna, M., Schlosser, D., & Hofrichter, M. (2007). Production, purification and partial enzymatic and molecular characterization of a laccase from the wood-rotting ascomycete Xylaria polymorpha. Enzyme and Microbial Technology, 41(6-7), 785-793. doi: 10.1016/j.enzmictec.2007.07.002
Lončar, N., Gligorijević, N., Božić, N., & Vujčić, Z. (2014). Congo red degrading laccases from Bacillus amyloliquefaciens strains isolated from salt spring in Serbia. International Biodeterioration & Biodegradation, 91, 18-23. doi: 10.1016/j.ibiod.2014.03.008
Lorenzo, M., Moldes, D., Couto, S. R., & Sanromán, A. (2002). Improving laccase production by employing different lignocellulosic wastes in submerged cultures of Trametes versicolor. Bioresource Technology, 82(2), 109-113. doi: 10.1016/s0960-8524(01)00176-6
Margot, J., Bennati-Granier, C., Maillard, J., Blánquez, P., Barry, D. A. & Holliger, C. (2013). Bacterial versus fungal laccase: potential for micropollutant degradation. AMB Express, 3, 63. doi: 10.1186/2191-0855-3-63
Mishra, S. K., & Srivastava, S. K. (2016). Production of extracellular laccase from bacterial strain Bacillus subtilis MTCC 1039 using different parameter. Biosciences Biotechnology Research Asia, 13(3), 1645-1650. doi: 10.13005/bbra/2312
Mongkolthanaruk, W., Tongbopit, S., & Bhoonobtong, A. (2012). Independent behavior of bacterial laccases to inducers and metal ions during production and activity. African Journal of Biotechnology, 11, 9391-9398. doi: 10.5897/AJB11.3042
Moshtaghioun, S. M., Haghbeen, K., Lotfi, S. A., Legge, R. L., Khoshneviszadeh, R., & Farhadi, S. (2011). Direct spectrophotometric assay of laccase using diazo derivatives of guaiacol. Analytical Chemistry, 83(11), 4200-4205. doi: 10.1021/ac200501w
Murugesan, K., Dhamija, A., Nam, I. H., Kim, Y. M., & Chang, Y. S. (2007). Decolorization of reactive black 5 by laccase: optimization by response surface methodology. Dyes and Pigments, 75, 176-184. doi: 10.1016/j.enzmictec.2006.08.028
Muthukumarasamy, N. P., Jackson, B., Raj, A. J., & Sevanan, M. (2015). Production of extracellular laccase from Bacillus subtilis MTCC 2414 using agroresidues as a potential substrate. Biochemistry Research International, 2015, ID765190, 1-10. doi: 10.1155/2015/765190
Narayanan, P. M., Murugan, S., Eva, A. S., Devina, S. U., & Kalidass, S. (2015). Application of immobilized laccase from Bacillus subtilis MTCC 2414 on decolourization of synthetic dyes. Research Journal of Microbiology, 10(9), 421-432. doi: 10.3923/jm.2015.421.432
Neifar, M., Chouchane, H., Mahjoubi, M., Jaouani, A., & Cherif, A. (2016). Pseudomonas extremorientalis BU118: a new salt-tolerant laccase-secreting bacterium with biotechnological potential in textile azo dye decolorization. 3 Biotech, 6(1), 107. doi: 10.1007/s13205-016-0425-7
Peter, J. K., & Vandana, P. (2014). Congo red dye decolorization by partially purified laccases from Pseudomonas aeruginosa. International Journal of Current Microbiology and Applied Sciences, 3(9), 105-115.
Rajeswari, M., & Bhuvaneswari, V. (2016). Production of extracellular laccase from the newly isolated Bacillus sp. PK4. African Journal Biotechnology, 15, 1813-1826. doi: 10.5897/AJB2016.15509
Robles, A., Lucas, R., Cienfuegos, G. A., & Gálvez, A. (2000). Phenol-oxidase (laccase) activity in strains of the hyphomycete Chalara paradoxa isolated from olive mill wastewater disposal ponds. Enzyme and Microbial Technology, 26(7), 484-490. doi: 10.1016/s0141-0229(99)00197-0
Shah, A. I., & Jobanputra, J. B. (2017). Production and partial purification of laccase produced by Bacillus species Isolated from Contaminated Soil. International Journal of Innovative Research in Science, Engineering and Technology, 6(9), 17862-17869. doi: 10.15680/IJIRSET.2017.0609003
Sharma, P., Goel, R. & Capalash, N. (2007). Bacterial laccases. World Journal of Microbiology and Biotechnology, 23, 823-832. doi: 10.1007/s11274-006-9305-3
Sheikhi, F., Ardakani, M. R., Enayatizamir, N., & Rodriguez-Couto, S. (2012). The determination of assay for laccase of Bacillus subtilis WPI with two classes of chemical compounds as substrates. Indian Journal of Microbiology, 52, 701-707. doi: 10.1007/s12088-012-0298-3
Shraddha, R. S., Simran, S., Mohit, K., & Ajay, K. (2011). Laccase: microbial sources, production, purification, and potential biotechnological applications. Enzyme Researchm, 11,217861. doi: 10.4061/2011/217861
Sondhi, S., Sharma, P., George, N., Chauhan, P. S., Puri, N. & Gupta, N. (2015). An extracellular thermo-alkali-stable laccase from Bacillus tequilensis SN4, with a potential to biobleach softwood pulp. 3 Biotech, 5, 175-185. doi: 10.1007/s13205-014-0207-z
Stackebrandt, E., & Goebel, B. M. (1994). Taxonomic note: a place for DNA-DNA reassociation and 16s rRNA sequence analysis in the present species definition in bacteriology. Internatinal Journal of Systematic and Bacteriology, 44(4), 846-849. doi: 10.1099/00207713-44-4-846
Strong, P. J. (2011). Improved laccase production by Trametes pubescens MB89 in distillery wastewaters. Enzyme Research, 379176. doi: 10.4061/2011/379176
Wang, C., Zhao, M., Lu, L., Wei, Z., & Li, T. (2011). Characterization of spore laccase from Bacillus subtilis WD23 and its use in dye decolorization. African Journal of Biotechnology, 10, 2186-2192. doi: 10.5897/AJB10.937
Xu, R., Zhang, K., Liu, P., Han, H., Zhao, S., Kakade, A., … Li, X. (2018). Lignin depolymerization and utilization by bacteria. Bioresource Technology, 269, 557-566. doi: 10.1016/j.biortech.2018.08.118
Zhu, D., Zhang, P., Xie, C., Zhang, W., Sun, J., Qian, W. J., & Yang, B. (2017). Biodegradation of alkaline lignin by Bacillus ligniniphilus L1. Biotechnology for Biofuels, 10, 44. doi: 10.1186/s13068-017-0735-y
Notes