Comunicaciones
RAPID AND ACCURATE PCR-BASED AND BOILING DNA ISOLATION METHODOLOGY FOR SPECIFIC DETECTION OF Sclerotium cepivorum IN GARLIC (Allium sativum) CLOVES
UNA METODOLOGÍA RÁPIDA Y SEGURA, BASADA EN PCR Y AISLAMIENTO DE DNA POR HERVIDO, PARA LA DETECCIÓN ESPECÍFICA DE Sclerotium cepivorum EN DIENTES DE AJO (Allium sativum)
UMA METODOLOGIA RÁPIDA E SEGURA, BASEADA EM PCR E ISOLAMENTO DE DNA POR FERVIDO PARA A DETECÇÃO ESPECÍFICA DE Sclerotium cepivorum EM DENTES DE ALHO (Allium sativum)
RAPID AND ACCURATE PCR-BASED AND BOILING DNA ISOLATION METHODOLOGY FOR SPECIFIC DETECTION OF Sclerotium cepivorum IN GARLIC (Allium sativum) CLOVES
Interciencia, vol. 44, no. 2, pp. 71-74, 2019
Asociación Interciencia
Received: 14 May 2018
Corrected: 21 January 2019
Accepted: 21 January 2019
Funding
Funding source: The authors CONACYT (Grant Nº 133440), FODECYT (Grant Nº 193512) and Convenio de Desempeño Regional UTA-1795 for financial support of this research. Karla Cecilia Licona-Juárez also thanks CONACYT for fellowship support. Karla Cecilia Licona-Juárez and Gerardo Acosta-García contributed equally to this study.
Contract number: CONACYT (Grant Nº 133440), FODECYT (Grant Nº 193512) and Convenio de Desempeño Regional UTA-1795
Award recipient: Karla Cecilia Licona-Juárez, Gerardo Acosta-García, Humberto Ramírez-Medina, Wilson Huanca-Mamani eLorenzo Guevara-Olvera
Abstract: Allium white rot caused by the pathogenic fungus Sclerotium cepivorum is a devastating disease of garlic worldwide. In this study, we developed and evaluated a molecular assay for the detection of S. cepivorumin mycelia and infected garlic cloves, through PCR, using a specific primer set combined with a rapid boiling DNA extraction. The PCR products were amplified in all the samples infected with S. cepivorum, but not in other samples including phylogenetically closely related fungi. This methodology is specific and has a shorter execution time compared to standard methods.
Keywords: Allium White Rot , Garlic , ITS Sequence , PCR Detection , Sclerotium cepivorum .
Resumen: La pudrición blanca en Allium causada por el hongo patógeno Sclerotium cepivorum es una enfermedad devastadora del ajo en todo el mundo. En este estudio desarrollamos y evaluamos un ensayo molecular para la detección de S. cepivorumen micelios y en dientes de ajo infectados, mediante PCR, usando un set de primers específicos, combinado con una rápida extracción de ADN basada en ebullición. Los productos de PCR fueron amplificados en todas las muestras infectadas con S. cepivorum, pero no en otras muestras, incluyendo hongos filogenéticamente cercanos. Esta metodología es específica y tiene un tiempo de ejecución más corto en comparación con los métodos estándar.
Resumo: A podridão branca do Allium causada pelo fungo patogénico Sclerotium cepivorum é uma doença devastadora do alho em todo o mundo. Neste estudo, desenvolvemos e avaliamos um ensaio molecular para a detecção de S. cepivorumem micélios e em dentes de alho infectados, por PCR, utilizando um conjunto de primers específicos, combinado com uma rápida extração de DNA em ebulição. Os produtos de PCR foram amplificados em todas as amostras infectadas com S. cepivorum, mas não em outras amostras, incluindo fungos filogeneticamente próximos. Essa metodologia é específica e possui um tempo de execução menor em comparação aos métodos padrão.
Introduction
Allium white rot is prevalent in many regions where garlic is cultivated worldwide and can reduce crop yields drastically (Davis et al., 2007). Garlic (Allium sativum L) is an economically important crop in Mexico. However, this crop is affected by the fungal disease known as Allium white rot, caused by Sclerotium cepivorum Berk (Couch and Kohn, 2000), which is considered the main fungus disease in several states of the country (Velásquez-Valle and Reveles-Hernández, 2016) In the State of Guanajuato the losses may reach up to 100% (Pérez-Moreno et al., 1998). In the absence of an Allium host, S. cepivorum persists in soil as sclerotia that can survive for more than 20 years and which can germinate in response to sulfides of Allium species, producing an infective mycelium (Coley-Smith et al., 1990). Many approaches have been used to control Allium white rot, including application of fungicides, soil fumigants and soil solarization, biological control agents (Miñambres et al., 2010) and composted onion waste (Coventry et al., 2002). However, the efficacy of these methods depends on the assurance that the pathogen is S. cepivorum and on the stage of development of the disease, which is not evident until the infection is well advanced (Ulacio-Osorio et al., 2006; Xu et al., 2010).
Key to the control is the ability to detect the pathogen in seed cloves. The standard method to detect the pathogen is to use infected tissue or soil to collect sclerotia (Bakonyi et al., 2011). However, isolation onto agar media and morphological identification is required to confirm the presence of S. cepivorum, since sclerotia of similar size can be produced by other species (Xu et al., 2010). This approach is time-consuming, as isolation requires several days of incubation, and accurate identification of the resulting colonies based on their morphology requires specialist expertise.
Polymerase chain reaction (PCR) analysis has been used to detect several plant fungal pathogens. Its main advantages are: i) time effective, ii) versatile, and iii) sensitive in identifying specific products. Standard protocols for the genetic detection of plant pathogens are based on NaOH-SDS, CTAB or commercial kits. However, alternative strategies such as boiling methods have never been developed and tested for the detection of S. cepivorumin garlic cloves.
In this study we developed and tested a PCR-based method to detect S. cepivorum in both mycelia and infected garlic cloves (Allium sativum) using the boiling DNA extraction method.
Materials and Methods
Experimental materials
The experimental work was carried out on Sclerotium cepivorum strain C2 isolated from Cortazar, Guanajuato, México; S. cepivorumK1 isolated from Zacatecas, Zacate-mi hermanacas, México; Sclerotinia sclerotiorum, Fusarium oxysporum and Phytophthora capsici isolates, and Allium sativum Tacátzcuaro cloves, provided by the Instituto Nacional de Investigaciones Forestales, Agrícolas y Pecuarias (INIFAP Bajío). S. cepivorum was maintained on agar plates with Czapeck media at 16ºC; S. sclerotiorum and F. oxysporum were maintained on agar plates with PDA medium at 25º; and P. capsici was maintained on agar plates with V8 media at 25ºC. Liquid media were used to obtain individual mycelia, which were harvested by centrifugation after 5 days and frozen at -80ºC.
Allium sativum Tacátzcuaro infection
Garlic cloves were germinated individually in 50ml glass flasks containing 20ml double-distilled water and incubated for 72h at 16ºC, then one square centimeter of 7 days old S. cepivorum mycelia grown in Czapeck media was added to infect germinated garlic cloves and incubated for 72h at 16ºC. Garlic cloves were harvested using sterile forks and frozen at -80ºC.
DNA isolation
Frozen mycelia and garlic cloves were used to extract genomic DNA by the boiling method (Holmes and Quigley, 1981); 2-4mg of mycelium or 80-120mg of non-infected garlic tissue were transferred into an Eppendorf tube containing 200mg of glass beads of 500µm in diameter and 150µl TE (Tris-Base 50mM, EDTA 2mM), vortexed for 2min, boiled for 5min and vortexed for 1min. To each tube 150µl of phenol: chloroform (5:1 v:v) was added, vortexed for 1min and centrifuged at 12000rpm for 5min. Each supernatant was transferred to a sterile Eppendorf tube, adding 100µl chloroform, vortexed for 1min and centrifuged at 12000rpm for 5min. DNA was re-suspended in 50ul H2O. The Dellaporta method was used as a standard DNA extraction method (Dellaporta et al., 1983).
Primer design and PCR
In order to synthesize amplicons around of 200bp, two sets of primers, ITSKF/ITSKR and ITSLF/ITSLR, were designed from the ITS sequence with the OligoPerfect Designer program (Invitrogen) with reference to the ITS sequences of S. cepivorum (Z99683.1, Z99682.1, Z99681.1 and JX442 200.1). The PCR reaction was performed using as template approximately 50ng or 100ng of DNA extracted by the boiling and Dellaporta methods, respectively. The PCR reaction was carried out as follows: 4min at 94ºC, 30 cycles of 1min at 94ºC, 30s at 58ºC or 68ºC, 30s at 72ºC and a final extension of 4min at 72ºC. The PCR products were separated on 1.2% agarose gel (Fermentas) stained with GelRed (Botium), using the GeneRuler 100pb DNA Ladder (Life Technologies) as size marker.
Cloning and sequence analysis
The PCR product was purified from the agarose gel and cloned into the pCR4-TOPO vector according to the manufacturer (Invitrogen). Chemically competent cells of Escherichia coli TOP10 (Invitrogen) were transformed and grown on LB agar plates containing kanamycin at a final concentration of 50µg·ml-1. Transformed colonies were transferred into Falcon tubes (Corning) containing 3ml of LB broth with kanamycin and ampicillin added at a final concentration of 50µg·ml-1 for plasmid DNA extraction. Each plasmid DNA was digested with the enzyme EcoRI to verify the presence of the insert.
Plasmid DNA containing the insert was sent to LANGEBIO (National Laboratory of Genomics for Biodiversity) for sequencing. The sequences were compared to the database of the NCBI (National Center for Biotechnology Information).
Results
Design of new primers to produce amplicons of 200bp by PCR
In this study, two sets of nested primers ITSKF (19-mer)/ITSKR (20-mer) and ITSLF (22-mer)/ITSLR (23-mer) (Table I) were identified as highly specific and efficient primers. DNA isolated by the boiling method from Tacátzcuaro cloves inoculated with Sclerotium cepivorum strain C2 was used as template to amplify the S. cepivorum ITS region by PCR; the results are shown in Figure 1. The specific DNA fragments were approximately 200bp (Lanes 3 and 4). The ITS1SCF/ITS2SCR primers (Haq et al., 2003) were used as positive control (Lane 2) and as negative control DNA was isolated by the boiling method from Tacátzcuaro cloves not inoculated with S. cepivorum strain C2 and new primers ITSLF/ITSLR (Lane 5); this result confirms that the designed new primers specifically allow the amplification of a ITS region of 200bp by PCR.
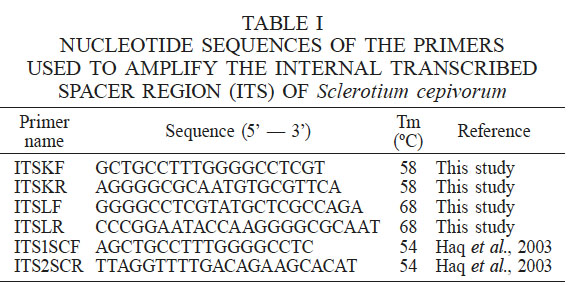
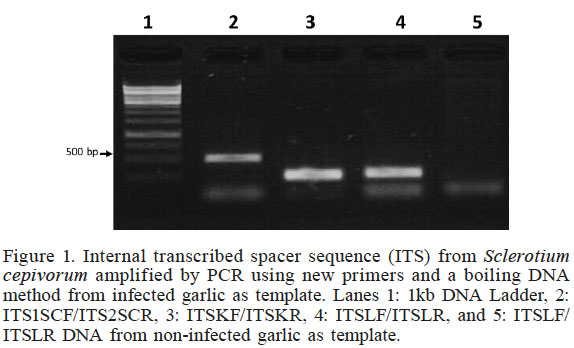
Figure 1
Internal transcribed spacer sequence (ITS) from Sclerotium cepivorum amplified by PCR using new primers and a boiling DNA method from infected garlic as template. Lanes 1: 1kb DNA Ladder, 2: ITS1SCF/ITS2SCR, 3: ITSKF/ITSKR, 4: ITSLF/ITSLR, and 5: ITSLF/ITSLR DNA from non-infected garlic as template
Identification and comparison of Sclerotim cepivorum strain C2 ITS sequence
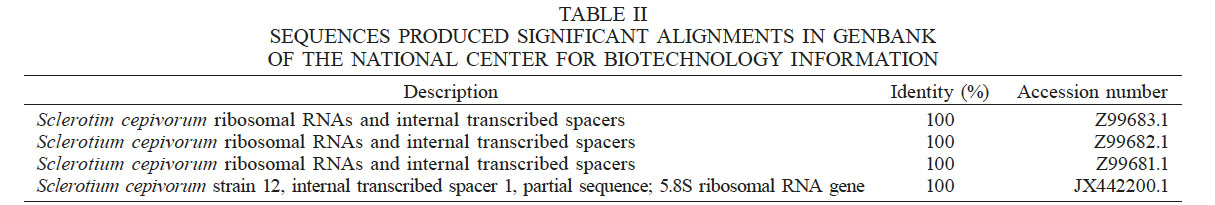
The 200bp fragments obtained by PCR were cloned into the vector pCR4-TOPO®TA, which was used to transform chemically competent Escherichia coli cells. Plasmid DNA was purified to obtain its sequence, which is shown in Figure 2. The sequence obtained was aligned with the NCBI database using the BLASTN algorithm (Altschul et al., 1990). The specific DNA fragment shared 100% identity with S. cepivorum ITS sequences Z99683.1, Z99682.1, Z99681.1 and JX442200.1, confirming the identification of S. cepivorum (Table II).

Figure 2
Partial internal transcribed spacer sequence (ITS) of Sclerotium cepivorum strain C2, using new ITSKF/ITSKR (underline) and ITSLF/ITSLR (double underline) primers (accession number KR108327.1).
Specificity of the new primer set for Sclerotium cepivorum ITS amplification
To demonstrate the specificity of the new designed primer ITSLF/ITSLR for S. cepivorum identification, PCR was performed using as template DNA isolated by the boiling method from several plant pathogens: Sclerotinia sclerotiorum, Fusarium oxysporum and Phytophthora capsici. The results are shown in Figure 3; no cross reaction was observed in PCR with any of the non-target species (Lanes 4, 5 and 6). PCR only achieved the synthesis of approximately 200pb amplicons for S. cepivorum strain C2 and S. cepivorum strain K1 (Lanes 2 and 3), confirming the high specificity of the PCR assay to detect the fungus causing white rot disease.
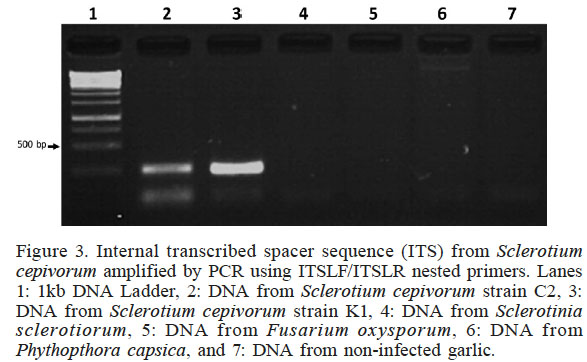
Figure 3
Internal transcribed spacer sequence (ITS) from Sclerotium cepivorum amplified by PCR using ITSLF/ITSLR nested primers. Lanes 1: 1kb DNA Ladder, 2: DNA from Sclerotium cepivorum strain C2, 3: DNA from Sclerotium cepivorum strain K1, 4: DNA from Sclerotinia sclerotiorum, 5: DNA from Fusarium oxysporum, 6: DNA from Phythopthora capsica, and 7: DNA from non-infected garlic
Discussion
Several tools have been used to differentiate fungi, including microbiological and phenotipic analysis (Bakonyi et al., 2013; Divya and Chouhan, 2014) and PCR (Haq et al., 2003; Xu et al., 2010). Detection of fungal pathogens by PCR is rapid and sensitive compared to other techniques. In this study we developed and tested a PCR-based assay for specific identification of Sclerotium cepivorum in both mycelia and infected garlic cloves, using rapid DNA extraction by the boiling method and new specific primers ITSKF/ITSKR and ITSLF/ITSLR. The boiling method for DNA extraction has not been used previously to identify S. cepivorumin garlic cloves. The boiling method is performed in ~30min, whereas the standard Dellaporta method, highly recommended for DNA extraction from plant tissue, requires about 2 to 10h (if overnight incubation is included) to be completed.
The amplicons obtained were approximately 200bp (Figure 1), which were confirmed by sequencing (Figure 2) and correspond to S. cepivorum C2. No amplification was obtained of DNA extracted from an noninfected garlic clove by the boiling method and using ITSKF/ITSKR or ITSLF/ITS LR primers (Figures 1 and 3), confirming the specificity of these primers to detect the presence of S. cepivorum.
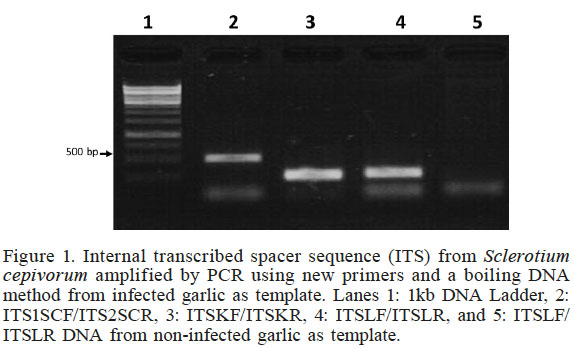
Figure 1
Internal transcribed spacer sequence (ITS) from Sclerotium cepivorum amplified by PCR using new primers and a boiling DNA method from infected garlic as template. Lanes 1: 1kb DNA Ladder, 2: ITS1SCF/ITS2SCR, 3: ITSKF/ITSKR, 4: ITSLF/ITSLR, and 5: ITSLF/ITSLR DNA from non-infected garlic as template.

Figure 2
Partial internal transcribed spacer sequence (ITS) of Sclerotium cepivorum strain C2, using new ITSKF/ITSKR (underline) and ITSLF/ITSLR (double underline) primers (accession number KR108327.1).
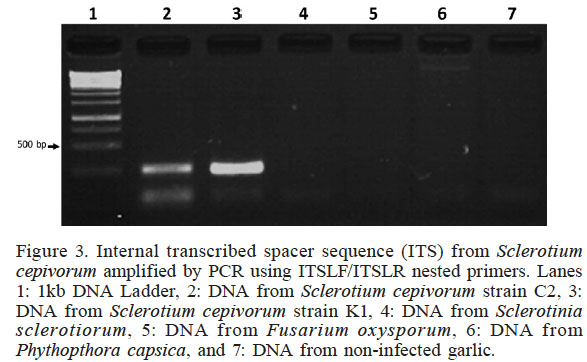
Figure 3
Internal transcribed spacer sequence (ITS) from Sclerotium cepivorum amplified by PCR using ITSLF/ITSLR nested primers. Lanes 1: 1kb DNA Ladder, 2: DNA from Sclerotium cepivorum strain C2, 3: DNA from Sclerotium cepivorum strain K1, 4: DNA from Sclerotinia sclerotiorum, 5: DNA from Fusarium oxysporum, 6: DNA from Phythopthora capsica, and 7: DNA from non-infected garlic
The PCR condition for new primers, for which the Tm are 58 and 68ºC, and annealing and synthesis times are 30sec, reduced significantly the total PCR-program time from 4h, used for primers ITS1SCF/ITS2SCR (Figure 1, Lane 2), to 2.5h using our primers. The PCR assay described here is highly specific for S. cepivorum(Figure 3, Lanes 2 and 3), because no PCR products were amplified for other plant pathogens such as Fusarium oxysporum and Phytophthora capsici, including a phylogenetically closely related fungus Sclerotinia sclerotiorum (Holst Jensen et al., 1998; Xu et al., 2010).
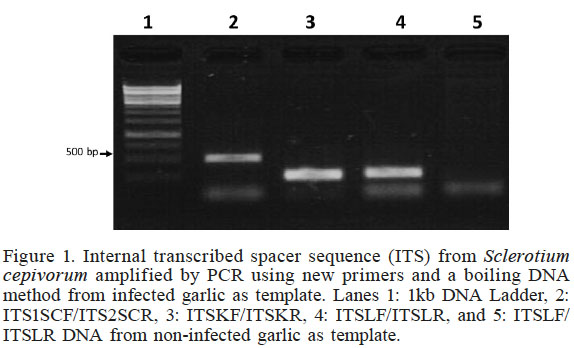
Figure 1
Internal transcribed spacer sequence (ITS) from Sclerotium cepivorum amplified by PCR using new primers and a boiling DNA method from infected garlic as template. Lanes 1: 1kb DNA Ladder, 2: ITS1SCF/ITS2SCR, 3: ITSKF/ITSKR, 4: ITSLF/ITSLR, and 5: ITSLF/ITSLR DNA from non-infected garlic as template.
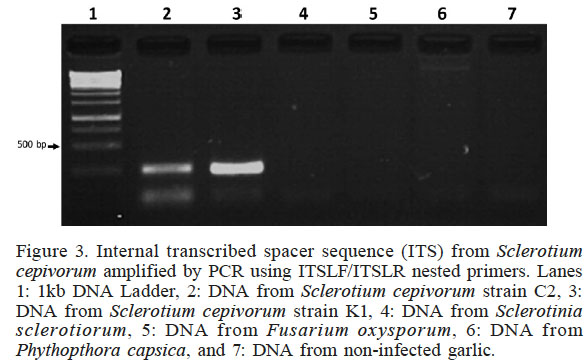
Figure 3
Internal transcribed spacer sequence (ITS) from Sclerotium cepivorum amplified by PCR using ITSLF/ITSLR nested primers. Lanes 1: 1kb DNA Ladder, 2: DNA from Sclerotium cepivorum strain C2, 3: DNA from Sclerotium cepivorum strain K1, 4: DNA from Sclerotinia sclerotiorum, 5: DNA from Fusarium oxysporum, 6: DNA from Phythopthora capsica, and 7: DNA from non-infected garlic
Conclusion
The methodology proposal in this study has two main major advantages over standard methods; i) it is highly specific because no other closely related fungi were falsely detected, thus avoiding problems associated with misidentification of samples contaminated with other fungi and ii) the total time required to establish the diagnosis was significantly reduced to ~3h compared to other methods that range from 6 to 14h. However, further studies with a larger sample are necessary to assess the efficacy of this method. The shorter execution time and accuracy of this strategy are very critical, especially in pathogens with a high potential of dispersion.
Acknowledgements
The authors CONACYT (Grant Nº 133440), FODECYT (Grant Nº 193512) and Convenio de Desempeño Regional UTA-1795 for financial support of this research. Karla Cecilia Licona-Juárez also thanks CONACYT for fellowship support. Karla Cecilia Licona-Juárez and Gerardo Acosta-García contributed equally to this study.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J. Mol. Biol. 215: 403-410.
Bakonyi J, Vajna L, Szeredi A, Tímár E, Kovács GM, Csősz M, Varga A (2011) First report of Sclerotium cepivorum causing white rot of garlic in Hungary. New Dis. Rep. 23: 5.
Coley-Smith JR, Mitchell CM, Sansford CE (1990) Long-term survival of sclerotia of Sclerotium cepivorum and Stromatinia gladioli. Plant Pathol. 39: 58-69.
Couch BC, Kohn LM (2000) Clonal spread of Sclerotium cepivorum in onion production with evidence of past recombination events. Phytopathology 90: 514-521.
Coventry E, Noble R, Mead A, Whipps JM (2002) Control of Allium White Rot (Sclerotium cepivorum) with composted onion waste. Soil Biol. Biochem. 34: 1037-1045.
Davis RM, Hao JJ, Romberg MK, Nunz JJ, Smith RF (2007) Efficacy of germination stimulants of sclerotia of Sclerotium cepivorum for management of White Rot of garlic. Plant Dis. 91: 204-208.
Dellaporta SL, Wood J, Hicks JB (1983) A plant DNA minipreparation version II. Plant Mol. Biol. Rep. 1(4): 19-21.
Divya KS, Chouhan JB (2014) Study of fungal diversity with reference to physical and chemical parameters. Int. J. Environ. Sci. 5: 401-406.
Haq MA, Collin HA, Tomsett AB, Jones MG (2003) Detection of Sclerotium cepivorum within onion plants using PCR primers. Physiol. Mol. Plant Pathol. 62: 185-189.
Holmes D, Quigley M (1981) A rapid boiling method for the preparation of bacterial plasmids. Analyt. Biochem. 114: 193-197.
Holst-Jensen A, Vaage M, Schumacher T (1998) An approximation to the phylogeny of Scle-rotinia and related genera. Nord. J. Bot. 18: 705-719.
Miñambres GG, Conles MY, Lucini EI, Verdelli RA, Meriles JM, Zygadlo JJ (2010) Application of thymol and iprodine to control garlic White Rot (Sclerotium cepivorum) and its effect on soil microbial communities. World J. Microbiol. Biotechnol. 26: 161-170.
Pérez-Moreno L, Olalde-Portugal V, Salinas-González JG, Sánchez-Pale JR, Entwistle AR (1998) Current known distribution of white rot Sclerotium cepivorum Berk. in Mexico. In: Entwistle, A.R., L. Pérez-Moreno (eds.) Proceedings of the 6th International Workshop on Allium white rot. Irapuato, Guanajuato, Mexico. pp. 6-9.
Ulacio-Osorio D, Zavaleta-Mejía E, Martínez-Garza A, Pedroza-Sandoval A (2006) Strategies for management of Sclerotium cepivorum Berk in garlic. J. Plant Pathol. 88: 253-261
Velásquez-Valle R, Reveles-Hernández M (2016) Effect of agents of alternative management on the garlic white rot development. Rev. Mex. Micol. 44: 41-47.
Xu Z, Harrington TC, Gleason ML, Batzer JC (2010) Phylogenetic placement of plant pathogenic Sclerotium species among teleomorph genera. Mycologia 102: 337-346.
Author notes
whuanca@uta.cl