Thermodynamics of DNA with “hump” Morse potential
Thermodynamics of DNA with “hump” Morse potential
Eclética Química, vol. 41, pp. 60-65, 2016
Universidade Estadual Paulista Júlio de Mesquita Filho

This work is licensed under Creative Commons Attribution 4.0 International.
Abstract: The thermal denaturation of DNA, i.e. the separation of the two strands is a phenomenon caused by the amplitude of the vibrations of the bases, therefore it is necessary to know how such separation is generated in order to implement alternative model of the melting behavior as a function of nucleotide sequence and therapies to combat the cancer. We propose to use the extended nonlinear Peyrard-Bishop(PB) model of DNA to include an anharmonic potential representing the aromatic stacking interaction between n- and (n-1)-th consecutive base pairs to treat the problem. We use Finite-difference methods for determine the mean value of the displacement for the “hump Morse” potential of the Peyrard-Bishop model of DNA. We show how the extended “pseudo- Schrödinger” combined with finite difference method can be used to obtain the mean value displacements for the thermal denaturation of DNA with “hump” Morse potential.
Keywords: Finite-difference methods, “pseudo-Schrödinger” equation, “hump” Morse potential.
Resumo: : A desnaturação térmica do DNA, ou seja, a separação das duas cadeias, é um fenômeno causado pela amplitude das vibrações das bases, portanto, é necessário saber como essa separação é gerada para implementar o modelo alternativo do comportamento de fusão como uma função da sequência de nucleotídeos e terapias para combater o câncer. Propomos usar o modelo de Peyrard-Bishop (PB) de DNA não linear estendido para incluir um potencial enarmônico que represente a interação de empilhamento aromático entre n- e (n-1)-enésimos pares de bases consecutivos para tratar o problema. Nós usamos métodos de diferença finita para determinar o valor médio do deslocamento para o potencial “corcunda de Morse” do modelo Peyrard-Bishop do DNA. Mostramos como o “pseudo-Schrödinger” estendido combinado com método de diferença finita pode ser usado para obter os deslocamentos do valor médio para a desnaturação térmica do DNA com o potencial Morse “da corcunda”.
Palavras-chave: Método das diferenças finitas, equação pseudo-Schrödinger, “corcunda” do potencial de Morse.
INTRODUCTION
We explore the thermodynamic properties solving the pseudo-Schrödinger equation of the transfer integral1,2,3,4,5,6,7 for the “hump Morse”. The mathematical tools used in the theoretical study of the problem are Finite Difference Method (FDM) and pseudo-Schrödinger equation with anharmonic stacking interactions for the DNA molecule.
THE MODELS
The Hamiltonian for the BP model is given by
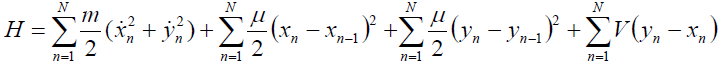 [Equation 1]
[Equation 1]or in the general form:
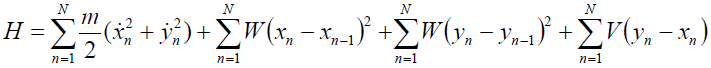 [Equation 1.1]
[Equation 1.1]where W is given by:
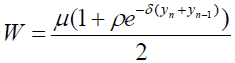 [Equation 1.2]
[Equation 1.2]where xn and yn denote the relative displacements of the nth nucleotide bases at each strand and N denotes the number of nucleotides. This expression can be viewed as a non-harmonic interaction with the parameter ρ≠0. The potential V(yn – xn) in the original (PB) model is taken as a Morse potential.
Using a suitable change of coordinates (“stretching” and “center of mass”) in equation (1), defined by
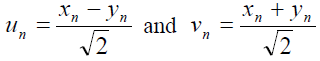 [Equation 2]
[Equation 2]the Hamiltonian (1) is divided in two parts, the acoustic (Hac) and optical (Hop) parts. They are given by
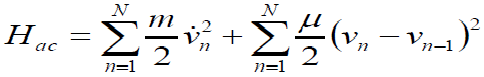 [Equation 3]
[Equation 3]and
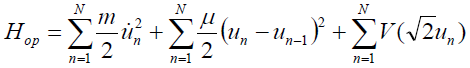 [Equation 4]
[Equation 4]respectively.
The general form is:
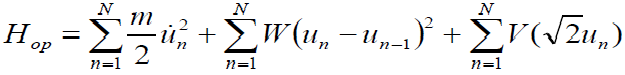 [Equation 4.1]
[Equation 4.1]The main interest here is in the optical Hamiltonian (4), because it contains the non- linearity of the problem. The equations of motion for equation (4) are a system of discrete nonlinear Klein-Gordon (KG) equations (n=1, 2... N):
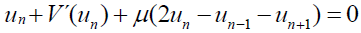 [Equation 5]
[Equation 5]with V'(un) being the derivative of the potential with respect the coordinate un.
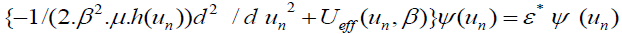 [Equation 5.1]
[Equation 5.1]The evaluation of the partition of equation (4.1) using the transfer integral operator method in the thermodynamic limit reduces to solving the “pseudo-Schrödinger” equation:
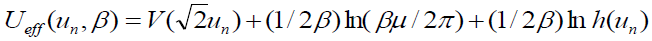 [Equation 5.2]
[Equation 5.2]where
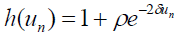 [Equation 5.3]
[Equation 5.3]We use the hump Morse potential28 in equation (4). The Morse Potential (PB model),
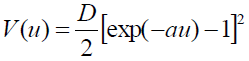 [Equation 6]
[Equation 6]was solved in32. The case for the symmetric Morse potential (SPB model) given by
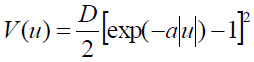 [Equation 7]
[Equation 7]was solved in29.
THERMODYNAMICS: THE MEAN VALUE OF THE DISPLACEMENTS
We can use the Finite difference methods for obtain the ground state wave function and the mean value of the displacement using the formulas of the literature7. We use the model (1) with the Morse Potential and the parameters μ =0.06, a= 4.45 and D=0.04. The Figure 1(b) shows that the mean of the amplitude is approximately 2.0 Å.
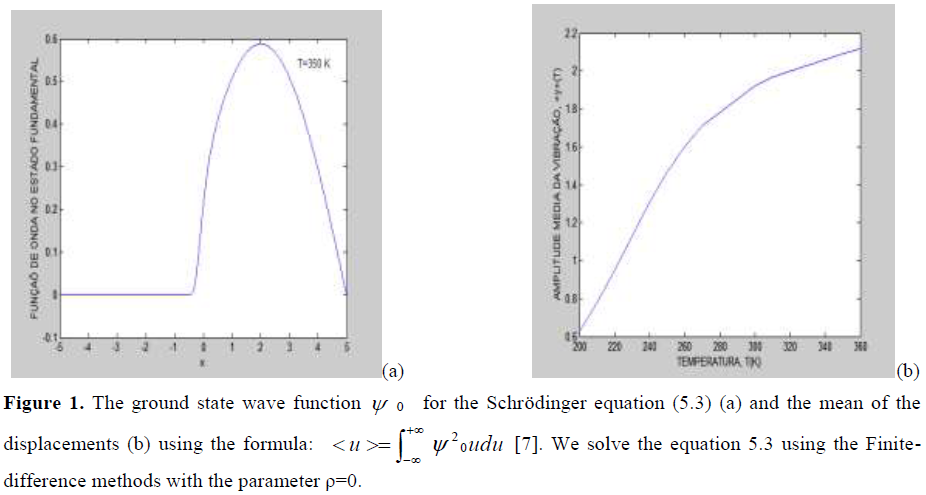
Figure 1
We can use the Finite difference methods for obtain the ground state wave function and the mean value of the displacement for the Hybrid potential (“hump Morse”)28 when the temperature is low and high. The results of this method is showed in Fig. 2(b).
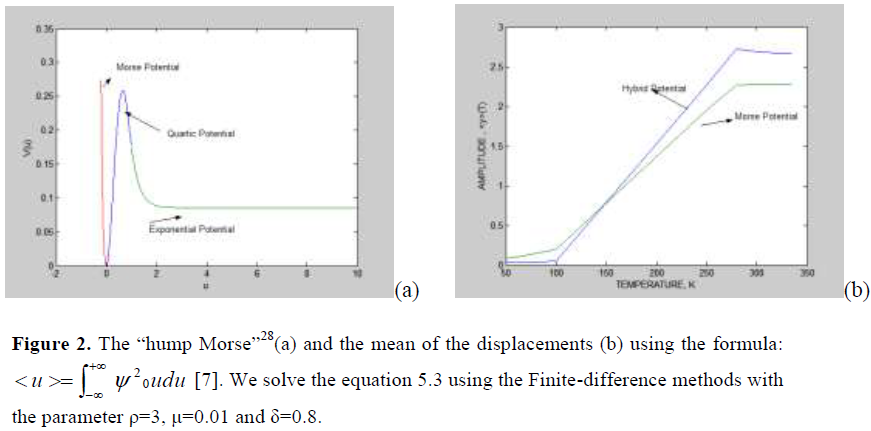
Figure 2
Also a model coupling vibrational and rotational motion for the DNA molecule30 extend the original PB model and the denaturation with anharmonic staking interaction (equation (1.2)) in DNA has the features of a first order phase transition. It is a consequence of the evaluation of the final average stretching using the partition function and the entropic barrier (the third term in equation (5.2)).
CONCLUSIONS
For the PB model we solve the “pseudo- Schrödinger” equation using the finite difference to determine eigenfunctions, the mean value of the displacements with a threshold value of 2.0 Å for the denaturation temperature of 350 K. The “Hump Morse” allow the model to describe the tunneling currents and the sharp thermal denaturation of DNA. Finally, it is important to emphasize the importance of the partition function in the numerical evaluation of the average stretching of the original PB model3 with rotational motions for nucleotides30 and anharmonic stacking for the existence of a first order phase transition.
REFERENCES
1. N. Theodorakopoulos, Phase transitions in one dimension: Are they all driven by domain walls?, Physica D, 2006, 216, 185-190.
2. L. Angelani, Relationship between phase transition and topological, Phys .Rev. E., 2005, 72, 1-9.
3. M. Peyrard, A.R. Bishop, Statistical Mechanics of a nonlinear model for DNA denaturation, Phys. Rev. Letters, 1989, 62, 2755-2758.
4. T. Dauxois, N. Theodorakopoulos, M. Peyrard, Thermodynamic Instabilities in OnDimension: Correlations, Scaling and Solitons, Journal of Statistical Physical, 2002, 107, 869-891.
5. J. Cuevas, Moving breathers in DNA model with competing short and longe-range dispersive interactions, Physica D, 2002, 163, 106-126.
6. Alvarez, J. F. R. Archilla, F. R. Romero, Dark breathers in Klein-Gordon Lattices. Band Analysis of their stability properties, New J. Phys., 2002, ., 1-72.
7. M. Peyrard, Nonlinear statistical physics of DNA¨, Nonlinearity, 2004, 17, 1-33.
8. J. de Luca, E. Drigo Filho, A. Ponno, J. R. Ruggiero, Energy localization in the Peyrard-Bishop model, Physical Review E, 2004, 70, 026213-1 to 026213-9.
9. M. Peyrard, The pathway to energy localization in nonlinear lattices, Physica D, 1998, 119, 184-199.
10. J. Cuevas, Moving discrete breather in a Klein-Gordon chain with an impurity, J. Phys. A, 2002, 35, 10519- 10530.
11. M. Hisakado, Breather trapping mechanism in piecewise homogeneous DNA, Physics Letters A, 1997, 227, 87-93.
12. M. Alvare, J. Cuevas, J. F. R. Archilla, Discrete moving breather collisions in a Kelin-Gordon chain oscillators, Physics Letters A, 2008, 372, 1256-1264.
13. F. C. Hoppensteadt, Analysis and Simulation of Chaotic Systems, Spring-Verlag, New York, 2000.
14. S. Aubry, Breathers in lattices: existence , linear stability, Physica D, 1997, 103, 201-250.
15. K. Forinash, M. Peyrard, B. Malomed, Interaction of discrete breathers with impurity modes, Phys. Rev. E, 1994, 49, 3400-3411.
16. R. S. Mackay, S. Aubry , Proof of existence of breathers for time-reversible for oscillators, Nonlinearity, 1994, ., 1623-1643.
17. J. L. Marin, S. Aubry, Breathers in nonlinear lattices : numerical calculation from the anticontinuous limit, Nonlinearity, 1996, ., 1501-1528.
18. H. Cortez, Modelo dinâmico e estatístico aplicado à transição de fase, Ph. D. Thesis, Universidade Estadual Paulista, São José do Rio Preto, Sao Paulo, Brasil, 2009.
19. S. Flach, C.R. Willis, Discrete breathers, Physics Reports, 1998, 295, 181-264.
20. S. Aubry, T. Cretegny, Mobility and Reactivity of Discrete Breathers, Physica D, 1998, 119, 34-46.
21. Y. Zhang, Theory of DNA melting based on the Peyrard-Bishop, Physical Review E, 1997, 56, 7100-7115.
22. J. Cuevas, Effect of the introduction of impurities on the stability of multibreathers at low coupling, Nonlinearity, 2005, 18, 769-790.
23. J. Cuevas, F. Palmero, J. F. R. Archilla, F. R. Romero, Moving discrete breather in a Klein-Gordon chain, J. Phys. A, 2002, 35, 10519-10530.
24. O. Bang, Generation of high-energy localized vibrational modes in nonlinear Klein-Gordon, Physical Review E, 1996, 53, 4143-4152.
25. J. Cuevas, Moving breathers in a bent DNA model, Physics Letters A, 2002, 299, 221-225.
26. J. L. Marin, J. C. Eibeck, F. M. Russell, Localized moving in a 2D lattice, Phys. Lett., 1998, A248, 225-229.
27. J. L. Marin, 2d breather, Nonlinear Science at the Dawn of the 21st Centure, Springer, Berlin, 2000, pp.293- 306.
28. M. Peyrard, Modelling DNA at the mesoscale: a challenge for nonlinear science, Nonlinearity, 2008, 21, 91- 100.
29. H. Cortez, E. Drigo, J. Ruggiero, Breather Stability in One Dimensional Lattices with a Symmetric Morse Potential, TEMA Tend. Mat. Apl. Comput., 2008, ., 205-212.
30. R. Silva, E. Drigo, J. Ruggiero, A model coupling vibrational and rotational motion for the DNA molecule, J. Biol. Physics, 2008, 34, 511-519.
31. M. Zoli, Path integral method for DNA denaturation, Physical Review E, 2009, 79, 021122-1 to 021122-5.
32. P. Augusto, E. Drigo, J. Ruggiero, Statistical Model to DNA melting, Eclet. Quim., 2001, 26, 77-85.
Author notes