Phytochemical and antibacterial investigation of Moringa oleifera seed: experimental and computational approaches
Phytochemical and antibacterial investigation of Moringa oleifera seed: experimental and computational approaches
Eclética Química, vol. 46, no. 2, pp. 17-25, 2021
Universidade Estadual Paulista Júlio de Mesquita Filho
Received: 31 March 2020
Accepted: 12 December 2020
Published: 01 April 2021
Abstract: The advent of antibiotic resistance and the growth of new strains of bacteria are of great concern to the world health sectors. Effective treatment of the infections caused by these human pathogens call for designing and development of new drugs with better efficiency. This study aimed at investigating the antibacterial activity of Moringa oleifera seed extract against some bacteria strains. The phytochemical and antibacterial activities of Moringa oleifera seed extract against both Gram-positive and Gram-negative bacteria were conducted using standard methods. The phytochemical analysis of the studied seed extract revealed the presence of alkaloids, tannins, saponins, phenols and flavonoids as secondary metabolites. The obtained result from the antibacterial study indicated that the extract exhibited high inhibition zones against all bacteria strain studied. Also, four molecular compounds from Moringa oleifera seed were selected based on their percentage yield and were optimized using density functional theory and they were used against five bacterial cell lines. It was observed that kaempferol and quercetin inhibited Escherichia coli. Kaempferol inhibited Klebsiella pneumonia, Pseudomonas aeruginosa, and Streptococcus pneumonia while Quercetin showed highest inhibition against Staphylococcus aureus when compared with other compounds. This showed the effectiveness of Moringa oleifera seed in eradicating some infections caused by bacteria.
Keywords: Moringa oleifera, antibacterial, phytochemical and multidrug resistant, DFT, Docking.
1. Introduction
Infectious diseases are syndromes that emanate from an infected person or animal to a susceptible host1. Infectious diseases remain the major cause of premature death in the world and it has been reported to be responsible for the death of about 50,000 people on a daily basis. The increase in the rate of bacterial infections has become alarming in the past few decades, thereby resulting into a severe health problem2. Staphylococcus aureus, Bacillus subtilis, Bacillus licheniformis and Esherichia coli have been reported as bacterial species for pathogenic infections of skin, septic arthritis, food intoxication food-borne illness, diarrhea and wounds3–5. Antimicrobial agents are very crucial in reducing the world encumbrance of infectious diseases6.
However, the multidrug resistance associated with pathogenic bacteria have led to serious health challenges7,8. Previous studies on rapid and extensive emergence of resistance to recent discovering of antimicrobial agents confirmed a short life expectancy of newly made antimicrobial agent9,10. The use of medicinal plants as source of drugs could be one of the most probable means of treating sickness and ailments in developing countries11. Moringa oleifera, popularly referred to as “horseradish tree”, is a medicinal plant found in the tropical and subtropical regions of the world and has been reported to possess antioxidant, anti-bacteria, anti-inflammatory, anti-cancerous activities12,13.
The discovery of some molecules used for protein–protein crossing point target have numerous challenges. Molecular docking revealed the connection between pharmacophore and receptor and it has the ability to provide assistance in detecting the appropriate binding site in the receptor. More so, calculated docking could be in the form of dock score, which is the arithmetic way of estimating the power of the relationship between docked compounds once the docking is accomplished14–16.
Thus, the compounds (catechin, epicatechin, kaempferol, quercetin) that were isolated from Moringa oleifera seed by Onuah et al.17, Sun et al.18, Ablajan and Tuoheti19, Saldanha et al.20 were theoretically studied via density functional theory and docking methods. Thus, in respect to the confirmation of the rapid global blowout of resistance to existing antibiotics, the need to find new anti-bacteria agents to reduce or end the epidemic health implication associated with multidrug resistant is of paramount importance.
2. Experimental
2.1 Sources of materials
Dry seed of Moringa oleifera, accession No. 266266, used in this study, were obtained from Obafemi Awolowo University Teaching and Research Farm (O.A.U.T. & R.F.), Ile-Ife, Nigeria, (7°33′N; 4°33′E).
The identification of the plant was done by a Plant Taxonomist in the Department of Botany, Obafemi Awolowo University. The pods enclosing the seed were removed. The seeds were air dried. Seeds of Moringa oleifera with good quality (Fig. 1) were selected and the seed coat and wings of the seeds were removed manually. The seeds were pulverized to fine powder, sieved and properly kept prior further analysis.

Figure 1.
Moringa oleifera seeds.
2.2 Extract preparation
About four hundred grams (400 g) of powdered seeds of Moringa oleifera were transferred into 1 L volumetric flask and 800 mL of methanol was added. The solution was allowed to stay for 72 h with vigorous agitation at 3 h interval. The extracting solvent was removed by decantation. The residue was filtered and concentrated on a digital rotary evaporator (Heidolph laborata 4010) to obtain 9.20 g crude extracts of Moringa oleifera.
2.3 Qualitative phytochemical analysis
The presence of alkaloids, reducing sugar, flavonoids, phenols, saponins and tannins in the seed extract of Moringa oleifera were qualitatively examined using standard methods21, as described below.
2.3.1 Determination of alkaloids
Zero point five gram (0.5 g) of Moringa oleifera seed extract was stirred with 5 mL of 1% HCl on water bath, it was filtered and a few drops of Meyer’s reagent was added to 1 mL of the filtrate. There was an occurrence of turbidity, which confirmed the present alkaloids.
2.3.2 Determination of tannins
Zero point three gram (0.3 g) of Moringa oleifera seed extract was dissolved in 10 mL of distilled water and filtered. Then freshly prepared FeCl3 solution was added to 5 mL of the filtrate. There was an observation of greenish dark coloration, which indicated the presence of tannins.
2.3.3 Determination of reducing sugar
Two drops of Fehling’s solution were added to a solution containing 0.5 g of Moringa oleifera seed extract in 5 mL distilled water and the solution was heated on water bath. The change in blue color of Fehling’s solution, which gave a red precipitation, indicated the presence of reducing sugar.
2.3.4 Determination of saponins
The formation of bubbles, which persisted for 20 min when Moringa oleifera seed extract dissolved in distilled water was vigorously shake, confirms the presence of saponins.
2.3.5 Determination of phenols
The intense coloration observed when FeCl3 solution was added to 0.4 g Moringa oleifera seed extract in distilled water confirmed the presence of phenols.
2.3.6 Determination of flavonoids
To the solution of Moringa oleifera seed extract, 5 mL of AlCl3, 2 mL methanol, 2 mL of concentrated HCl, 2 mL of KOH solution and few drops of magnesium turning was added. A pink coloration was observed which confirmed the presence of flavonoids.
2.4 Bacterial analysis
The antibacterial activity of the seed extract was examined using pure cultures of clinical isolates Gram-positive bacteria (Staphylococcus aureus and Streptococcus pneumonia) alongside with Gram-negative bacteria (Escherichia coli, Klebsiella pneumonia and Pseudomonas aeruginosa), obtained from the Department of Microbiology, Obafemi Awolowo University, Ile-Ife, Nigeria, were used for this study. The culturing and identification of the selected bacterial species was done using modified method given by Colle et al.22. The antibacterial sensitivity of the seed extract of Moringa oleifera against selected bacteria were examined with agar well diffusion method. The culture media was sterilized for 15 min at 121 °C. 0.1 mL standard inoculum of the test bacteria previously inoculated with bacterial suspension (100 mL of medium/1 mL of 107 CFU) to attain 105 CFU/mL of medium was introduced into the sterilized media. The inoculum was spread through the media. A sterile cork borer was used to bore a 6 mm well into the agar medium. Zero point one milliliter (0.1 mL) of 2, 4 and 6 mg mL-1 concentration obtained by re-dissolved 10 mg of Moringa oleifera seed extract in 5, 2.5 and 0.8 mL of methanol, respectively, was transferred into each of the well on the medium. The plates were allowed to stand for 1 h before incubation for another 24 h at 37 °C. The zones of inhibition on the plates were measured with a transparent ruler in millimeters. Filter paper discs loaded with 5 µg of ampicillin were used as control. The plates were kept in the fridge at 5 °C for 2 h to allow plant extracts diffusion then after incubation at 37 °C for 48 h, the zones of inhibition were measured. The effect of the seed extract of Moringa oleifera on bacteria strains was compared with standard control at concentrations of 2, 4 and 6 mg mL-1. The percentages of zones of inhibition were calculated to determine the antibacterial activity of the seed extract of Moringa oleifera. The antibacterial investigation was on triplicate analysis. The adopted formula from the study23 was used for calculating the percentage growth inhibition, as Eq. 1:
 (1)
(1)where PGI is percent growth inhibition, BDC is the bacteria colony diameter in control and BDT means bacteria colony diameter in treatment.
2.5 Theoretical calculation
Four (4) selected molecular compounds17–20 from molecules present in Moringa oleifera were used for this study. The selected compounds were chosen due to their highest percentage yield and the selected compounds were optimized using density functional theory at 6-31G* basis set and several molecular descriptors [EHOMO, ELUMO, band gap, dipole moment, hydrogen bond donor (HBD), hydrogen bond acceptor (HBA) log P, polarizability and Polar suface area (PSA)] which described anti-bacteria activity of the studied flavonoids were obtained using Spartan’14 software by wavefunction Inc. Also, the docking study were executed on the studied compounds and the studied enzymes (2J5O24, 2RQX25, 2W9T26, 1GRX27, 2M6U28) using Discovery studio, AutoDock Tool, AutoDock Vina and Pymol which serves as a post-dock software.
3. Results and discussion
3.1 Qualitative phytochemical analysis
The qualitative phytochemical analysis of Moringa oleifera seed extract confirmed the presence of alkaloids, tannins, saponins, phenols, flavonoids and the absence of reducing sugar (Tab. 1). However, the absence of reducing sugar indicated that the anti-bacteria potency of plant extracts against Gram-negative bacteria was due to the presence of phenolic compounds29. Findings from literature revealed that medicinal plants containing alkaloids, phenols and flavonoids as bioactive metabolites have good antibacterial properties30–33. Therefore, the presence of these metabolites in the seed extract of Moringa oleifera could render it as a good antibacterial agent.
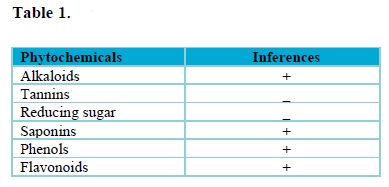
+ represents presence of bioactive compounds, while; – represents absence of bioactive compounds.
3.2 Antibacterial activities of Moringa oleifera seed extract against test bacteria
The percentage zones of inhibition exhibited by Moringa oleifera seed extract against selected bacterial strains are showed in Fig. 2. The antibacterial study revealed that Moringa oleifera seed extract showed some levels of inhibition against all the bacteria strain at various concentrations. The decreasing order of susceptibility of the bacteria strain to Moringa oleifera seed extract at 2 mg mL-1 was Staphylococcus aureus > Klebsiella pneumonia > Escherichia coli, Streptococcus pneumonia > Pseudomonas aeruginosa. Furthermore, when the percentage zones of inhibition of Moringa oleifera seed extract at 2 mg mL-1 was compared with the control, the percentage zones of inhibition observed was called “weak inhibition”, because percentage zones of inhibition were below the average of that of the control.
At concentration of 4 mg mL-1, an increase in percentage zones of inhibition was observed and was found higher than the average percentage zones of inhibition exhibited by the control when compared at same concentration. This was called “moderate inhibition”. The increasing order of potency of the extract against the bacteria strain was Klebsiella pneumonia > Staphylococcus aureus > Pseudomonas aeruginosa, Klebsiella pneumonia > Escherichia coli > Streptococcus pneumonia.
The highest percentage zones of inhibition were detected at concentration of 6 mg mL-1. The increasing order of effectiveness of Moringa oleifera seed extract against the test bacteria was given as Pseudomonas aeruginosa > Klebsiella pneumonia > Staphylococcus aureus > Escherichia coli > Streptococcus pneumonia. The zones of inhibition in this study were lower than that obtained from the study of Folorunso et al., on biosynthesis, characterization and antimicrobial activity of gold nanoparticles from leaf extracts of Annona muricata23 and higher than obtained from the study of Sarital et al., on the in vitro antimicrobial activity of some medicinal plants against human pathogenic bacteria34. This is an indication that Moringa oleifera seed extract has good inhibitory efficiency against bacteria and could possibly be used as an antibiotic for the treatment of infected diseases caused by bacteria.
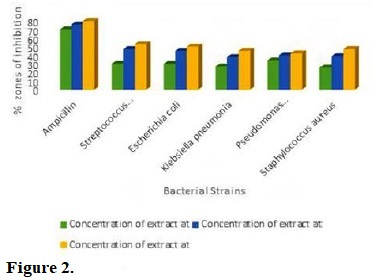
Figure 2.
Antibacterial activities of Moringa oleifera seed extract against test bacteria.
3.3 Docking calculations
Docking study was carried out on the flavonoids in Moringa oleifera seed and series of bacteria cell lines (2J5O, 2RQX, 2W9T, 1GRX, 2M6U). The obtained result, i.e., binding affinity between the studied compounds, were displayed in Tab. 2. As reported by Oyebamiji et al., molecules that have lower binding affinity show the molecule with tendency to inhibit well35.
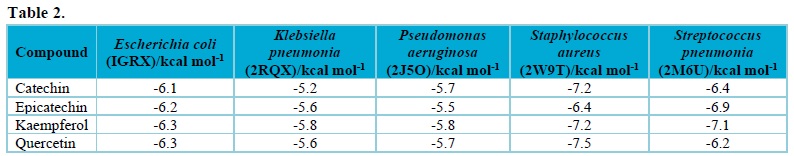
Thus, it was observed that kaempferol and quercetin are one of the vital flavonoids present in Moringa oleifera seed, as shown in Tab. 2. As displayed in Tab. 2, both kaempferol and quercetin inhibited Escherichia coli most. Kaempferol inhibited Klebsiella pneumonia, Pseudomonas aeruginosa, and Streptococcus pneumonia more than other studied compounds. Also, only quercetin possess the ability to inhibit Staphylococcus aureus more than other compounds used in this work. The residues involved in the interaction are shown in Figs. 3-8
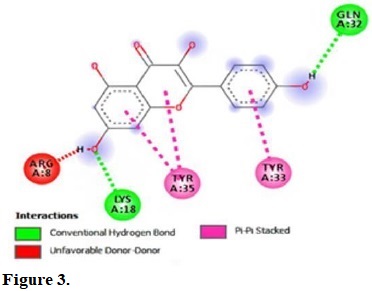
Figure 3.
Molecular interactions of kaempferol with the residue in the gouge of Escherichia coli (IGRX).
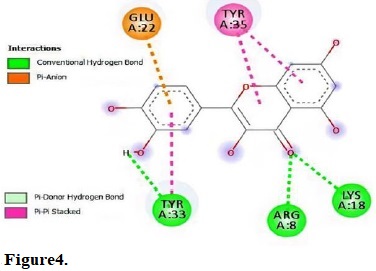
Figure 4.
Molecular interactions of quercetin with the residue in the gouge of Escherichia coli (IGRX).
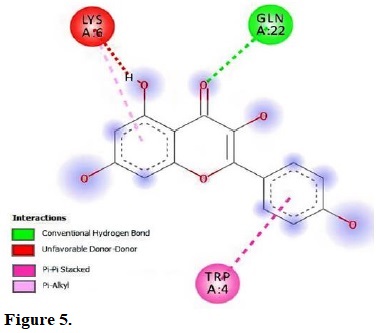
Figure 5.
molecular interactions of kaempferol with the residue in the gouge of Klebsiella pneumonia (2RQX).
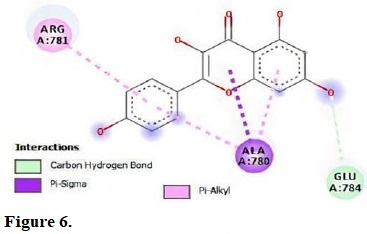
Figure 6.
Molecular interactions of kaempferol with the residue in the gouge of Pseudomonas aeruginosa (2J5O).
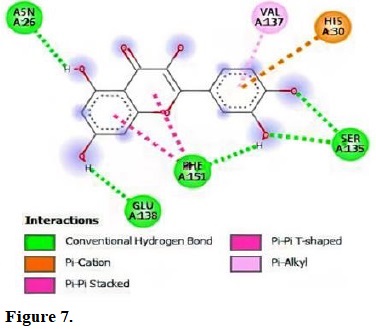
Figure 7.
Molecular interactions of quercetin with the residue in the gouge of Staphylococcus aureus (2W9T).
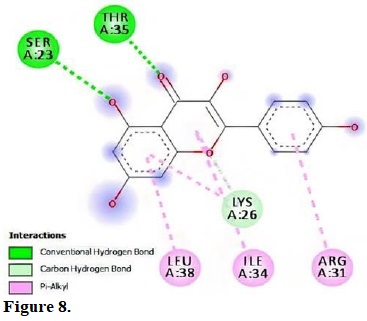
Figure 8.
Molecular interactions of kaempferol with the residue in the gouge of Streptococcus pneumonia (2M6U).
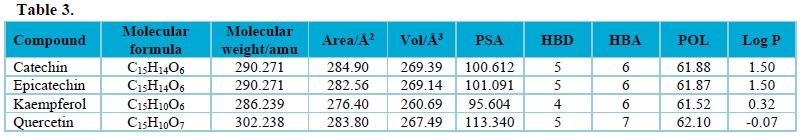
Moreover, the molecular descriptors obtained for the studied compounds, as shown in Tab. 3, were subjected to Lipinski rule of five (Molecular Weight ≤ 500, Log P ≤ 5, HBD ≤ 5, and HBA ≤ 10)36 and it was observed that the calculated molecular descriptors obtained from the studied compounds using density functional theory method obeyed the rule. This showed that all compounds used in this work have drug potential.
4. Conclusions
Moringa oleifera seed extract could be modified as antibiotic ingredient to combat both Gram-positive and Gram-negative bacterial infections, which could help in some way to reduce or overcome the health challenges caused by bacterial resistance to many locally or commercially available antibiotics. Therefore, isolation of active compounds from Moringa oleifera seed is highly recommended for the discoveries of proficient antibacterial agent that can serve as conventional antibiotics that can constrain or cure the infectious diseases caused by pathogenic bacterial strains. In this work, kaempferol and quercetin inhibited Escherichia coli more than other studied compounds. Also, kaempferol possess the ability to inhibit Klebsiella pneumonia, Pseudomonas aeruginosa and Streptococcus pneumonia more than other studied compounds, as well as Quercetin possess the capability to inhibit Staphylococcus aureus more than other compounds. The molecular descriptors were also in line with Lipinski rule.
Acknowledgments
We are grateful to Computational Chemistry Research Laboratory, Department of Pure and Applied Chemistry, Ladoke Akintola University of Technology for the computational resources and Mrs E.T. Oyebamiji as well as Miss Priscilla F. Oyebamiji for the assistance in the course of this study.
References
[1] Bhatia, R., Narain, J. P., The growing challenge of antimicrobial resistance in the South-East Asia Region - are we losing the battle? Indian Journal of Medical Research 132 (5) (2010) 482-486.
[2] El-Wafa, W. M. A., El-All, W. S. M. A., Effect of Moringa oleifera Seed Oil on Antimicrobial Activity of some Antibiotics against some Pathogenic Gram-Negative Bacteria, International Journal of Current Microbiology and Applied Sciences 4 (5) (2015) 140-151.
[3] Apetroaie-Constantin, C., Mikkola, R., Andersson, M. A., Teplova, V., Suominen, I., Johansson, T., Salkinoja-Salonen, M., Bacillus subtilis and B. mojavensis strains connected to food poisoning produce the heat stable toxin amylosin, Journal of Applied Microbiology 106 (6) (2009) 1976-1985. https://doi.org/10.1111/j.1365-2672.2009.04167.x.
[4] Karch, H., Tarr, P. I., Bielaszewska, M., Enterohaemorrhagic Escherichia coli in human medicine, International Journal of Medical Microbiology 295 (6-7) (2006) 405-418. https://doi.org/10.1016/j.ijmm.2005.06.009.
[5] Fazli, M., Bjarnsholt, T., Kirketerp-Møller, K., Jørgensen, A., Andersen, C., Givskov, M., Tolker-Nielsen, T., Quantitative analysis of the cellular inflammatory response against biofilm bacteria in chronic wounds, Wound Repair and Regeneration 19 (3) (2011) 387-391. https://doi.org/10.1111/j.1524-475X.2011.00681.x.
[6] Last, J. M., A Dictionary of Epidemiology, Oxford University Press, Oxford, 1998.
[7] Boucher, H. W., Talbot, G. H., Bradley, J. S., Edwards, J. E., Gilbert, D., Rice, L. B., Scheld, M., Spellberg, B., Bartlett, J., Bad Bugs, No Drugs: No ESKAPE! An Update from the Infectious Diseases Society of America, Clinical Infectious Diseases 48 (1) (2009) 1-12. https://doi.org/10.1086/595011.
[8] Giamarellou, H., Multidrug-resistant Gram-negative bacteria: how to treat and for how long, International Journal of Antimicrobial Agents 36 (Suppl 2) (2010) S50-S54. https://doi.org/10.1016/j.ijantimicag.2010.11.014.
[9] Coates, A., Hu, Y., Bax, R., Page, C., The future challenges facing the development of new antimicrobial drugs, Nature Reviews Drug Discovery 1 (11) (2002) 895-910. https://doi.org/10.1038/nrd940.
[10] Marasini, B. P., Baral, P., Aryal, P., Ghimire, K. R., Neupane, S., Dahal, N., Singh, A., Ghimire, L., Shrestha, K., Evaluation of Antibacterial Activity of Some Traditionally Used Medicinal Plants against Human Pathogenic Bacteria, BioMed Research International 2015 (2015) 265425. https://doi.org/10.1155/2015/265425.
[11] Kumar, V., Pandey, N., Mohan, N., Singh, R. P., Antibacterial & antioxidant activity of different extract of Moringa oleifera leaves - an in vitro study, International Journal of Pharmaceutical Sciences Review and Research 12 (1) (2012) 14. https://globalresearchonline.net/journalcontents/v12-1/014.pdf.
[12] Rani, N. Z. A., Husain, K., Kumolosasi, E., Moringa Genus: A Review of Phytochemistry and Pharmacology, Frontiers in Pharmacology 9 (2018) 108. https://doi.org/10.3389/fphar.2018.00108.
[13] Wang, L., Chen, X., Wu, A., Mini Review on Antimicrobial Activity and Bioactive Compounds of Moringa oleifera, Medicinal Chemistry 6 (9) (2016) 578-582. https://doi.org/10.4172/2161-0444.1000402.
[14] Oyebamiji, K. A., Semire, B., Studies Of 1, 4-Dihydropyridine Derivatives for Anti-Breast Cancer (MCF-7) Activities: Combinations Of DFT-QSAR And Docking Methods, New York Science Journal 9 (6) (2016) 58-66.
[15] Taylor, R. D., Jewsbury, P. J., Essex, J. W., A review of protein-small molecule docking methods, Journal of Computer-Aided Molecular Design 16 (3) (2002) 151-166. https://doi.org/10.1023/A:1020155510718.
[16] Duman, R., Ishikawa, S., Celik, I., Strahl, H., Ogasawara, N., Troc, P., Lowe, J., Hamoen, L. W., Structural and genetic analyses reveal the protein SepF as a new membrane anchor for the Z ring, PNAS 110 (48) (2013) E4601. https://doi.org/10.1073/pnas.1313978110.
[17] Onuah, C. L., Chukwuma, C. C., Ohanador, R., Chukwu, C. N., Iruolagbe, J., Quantitative Phytochemical Analysis of Annona muricata and Artocarpus heterophyllus Leaves Using Gas Chromatography-flame Ionization Detector, Trends in Applied Sciences Research 14 (2) (2019) 113-118. https://doi.org/10.3923/tasr.2019.113.118.
[18] Sun, J., Liang, F., Bin, Y., Li, P., Duan, C., Screening Non-colored Phenolics in Red Wines using Liquid Chromatography/Ultraviolet and Mass Spectrometry/Mass Spectrometry Libraries, Molecules 12 (3) (2007) 679-693. https://doi.org/10.3390/12030679.
[19] Ablajan, K., Tuoheti, A., Fragmentation characteristics and isomeric differentiation of flavonol .‐rhamnosides using negative ion electrospray ionization tandem mass spectrometry, Rapid Communications in Mass Spectrometry 27 (3) (2013) 451-460. https://doi.org/10.1002/rcm.6476.
[20] Saldanha, L. L., Vilegas, W., Dokkedal, A. L., Characterization of Flavonoids and Phenolic Acids in Myrcia bella Cambess. Using FIA-ESI-IT-MS. and HPLC-PAD-ESI-IT-MS Combined with NMR, Molecules 18 (7) (2013) 8402-8416. https://doi.org/10.3390/molecules18078402.
[21] Oyebamiji, A. K., Akintelu, S. A., Folorunso, A. S., Abiola, B. E., Ajayi, S. O., Abdusalam, I. O., Morakinyo, A. E., Computational and Experimental Studies on Antimicrobial Activity of the Bark of Annona muricata against Some Selected Human Pathogenic Bacteria and Fungi, International Journal of Modern Chemistry 11 (1) (2019) 9-27.
[22] Colle, J. G., Fraser, A. G., Marmion, B. P., Simmans, A., Mackie, McCartney, Practical Medical Microbiology, Elsevier, Amsterdam, 1996.
[23] Folorunso, A., Akintelu, S., Oyebamiji, A. K., Ajayi. S., Abiola, B., Abdusalam, I., Morakinyo, A., Biosynthesis, characterization and antimicrobial activity of gold nanoparticles from leaf extracts of Annona muricata, Journal of Nanostructure in Chemistry 9 (2019) 111-117. https://doi.org/10.1007/s40097-019-0301-1.
[24] Sivanathan, V., Allen, M. D., Bekker, C., Baker, R., Arciszewska, L. K., Freund, S. M., Bycroft, M., Löwe, J., Sherratt, D. J., The FtsK γ domain directs oriented DNA translocation by interacting with KOPS, Nature Structural & Molecular Biology 13 (2006) 965-972. https://doi.org/10.1038/nsmb1158.
[25] Luo, S.-C., Lou, Y.-C., Cheng, H.-Y., Pan, Y.-R., Peng, H.-L., Chen, C., Solution structure and phospho-PmrA recognition mode of PmrD from Klebsiella pneumonia, Journal of Structural Biology 172 (3) (2010) 319-330. https://doi.org/10.1016/j.jsb.2010.06.007.
[26] Heaslet, H., Harris, M., Fahnoe, K., Sarver, R., Putz, H., Chang, J., Subramanyam, C., Barreiro, G., Miller, J. R., Structural comparison of chromosomal and exogenous dihydrofolate reductase from Staphylococcus aureus in complex with the potent inhibitor trimethoprim, Proteins: Structure, Function, and Bioinformatics 76 (3) (2009) 706-717. https://doi.org/10.1002/prot.22383.
[27] Xia, T.-H., Bushweller, J.-H., Sodano, P., Billeter, M., Bjornberg, O., Holmgren, A., Wüthrich, K., NMR structure of oxidized Escherichia coli glutaredoxin: Comparison with reduced E. coli glutaredoxin and functionally related proteins, Protein Science 1 (3) (1992) 310-321. https://doi.org/10.1002/pro.5560010302.
[28] Achila, D., Liu, A., Banerjee, R., Li, Y., Martinez-Hackert, E., Zhang, J.-R., Yan, H., Structural determinants of host specificity of complement Factor H recruitment by Streptococcus pneumoniae, Biochemical Journal 465 (2) (2015) 325-335. https://doi.org/10.1042/BJ20141069.
[29] Marrufo, T., Nazzaro, F., Mancini, E., Fratianni, F., Coppola, R., de Martino, L., Agostinho, A. B., de Feo, V., Chemical Composition and Biological Activity of the Essential Oil from Leaves of Moringa oleifera Lam. Cultivated in Mozambique, Molecules 18 (9) (2013) 10989-11000. https://doi.org/10.3390/molecules180910989.
[30] Lewis, K., Ausubel, F. M., Prospects for plant-derived antibacterials, Nature Biotechnology 24 (12) (2006) 1504-1507. https://doi.org/10.1038/nbt1206-1504.
[31] Yu, T., Yamaguchi, H., Noshita, T., Kidachi, Y., Umetsu, H., Ryoyama, K., Selective cytotoxicity of glycyrrhetinic acid against tumorigenic r/m HM-SFME-1 cells: Potential involvement of H-Ras downregulation, Toxicology Letters 192 (3) (2010) 425-430. https://doi.org/10.1016/j.toxlet.2009.11.021.
[32] Cushnie, T. P. T., Lamb, A. J., Antimicrobial activity of flavonoids, International Journal of Antimicrobial Agents 26 (5) (2005) 343-356. https://doi.org/10.1016/j.ijantimicag.2005.09.002.
[33] Emiru, Y. K., Siraj, E. A., Teklehaimanot, T. T., Amare, G. G., Antibacterial Potential of Aloe weloensis (Aloeacea) Leaf Latex against Gram-Positive and Gram-Negative Bacteria Strains, International Journal of Microbiology 2019 (2019) 5328238. https://doi.org/10.1155/2019/5328238.
[34] Manandhar, S., Luitel, M., Dahal, R. K., In Vitro Antimicrobial Activity of Some Medicinal Plants against Human Pathogenic Bacteria, Journal of Tropical Medicine 2019 (2019) 1895340. https://doi.org/10.1155/2019/1895340.
[35] Oyebamiji, A. K., Oyedeji, F. O., Adejoro, I. A., Adeleke, B. B., Computational Studies of Some Hydrazone Derivatives as Antibacterial Agent: DFT and Docking Methods, The journal of Pure and Applied Chemistry Research 8 (1) (2019) 1-6. https://doi.org/10.21776/ub.jpacr.2019.008.1.433.
[36] Adejoro, I. A., Waheed, S. O., Adeboye, O. O., Molecular Docking Studies of Lonchocarpus cyanescens Triterpenoids as Inhibitors for Malaria, Journal of Physical Chemistry & Biophysics 6 (2) (2016) 1000213. https://doi.org/10.4172/2161-0398.1000213.
Author notes
abeloyebamiji@gmail.com