Communication

This work is licensed under Creative Commons Attribution-NonCommercial 4.0 International.
Received: 09 March 2018
Accepted: 24 October 2019
DOI: https://doi.org/10.5433/1679-0359.2019v40n1p503
Abstract: In recent years, anthropogenic factors such as pollution, overfishing, and construction of hydroelectric plants have significantly impacted natural fish populations. Research focusing on genetically evaluation of these impacts is necessary to objectively target conservation programs. The aim of this study was to evaluate the genetic diversity of Curimba (Prochilodus lineatus), Pacu (Piaractus mesopotamicus), and Piracanjuba (Brycon orbignyanus) populations from the Água Vermelha Reservoir, Rio Grande-SP. Microsatellite loci were amplified, producing 56, 24, and 26 alleles for the populations of the three species, respectively. The number of alleles per locus ranged from three to ten for P. lineatus, two to five for P. mesopotamicus, and two to four for B. orbignyanus. The observed heterozygosity (Ho) was higher in the P. lineatus population (0.547), relative to the P. mesopotamicus and B. orbignyanus populations (0.473 and 0.527, respectively). The mean values of Ho were lower than the average expected heterozygosity (He) in the three species, being corroborated by the positive inbreeding coefficient (Fis). Deviations from the Hardy-Weinberg equilibrium (HWE) were found in five, three, and two loci for P. lineatus, P. mesopotamicus, and B. orbignyanus, respectively. Wilcoxon tests revealed recent bottlenecks in the three species, evidenced by a significant excess of heterozygotes (p < 0.05) detected only in the Infinite Allele Model (IAM). In conclusion, adequate genetic variability was observed in the three populations with the presence of heterozygous deficits.
Keywords: Brycon orbignyanus, Conservation, Genetic variability, Microsatellites, Piaractus mesopotamicus, Prochilodus lineatus.
Resumo: Nos últimos anos, fatores antrópicos como poluição, sobrepesca e construção de hidrelétricas têm impactado significativamente as populações naturais de peixes. Pesquisas que permitam avaliar geneticamente esse impacto são necessárias para orientar objetivamente programas de conservação. O objetivo do presente estudo foi avaliar a diversidade genética de populações de Curimba (Prochilodus lineatus), Pacu (Piaractus mesopotamicus) e Piracanjuba (Brycon orbignyanus) provenientes do reservatório de Água Vermelha, Rio Grande-SP. Foram amplificados loci microssatélites que produziram 56, 24 e 26 alelos para as populações das três espécies, respectivamente. O número de alelos por locus variou de três a dez para P. lineatus, dois a cinco para P. mesopotamicus e dois a quatro para B. orbignianus. A heterozigosidade observada (Ho) foi mais elevada na população de P. lineatus (0,547) em relação às populações de P. mesopotamicus e B. orbignyanus (0,473 e 0,527, respectivamente). A média dos valores de Ho foram inferiores à média de He nas três espécies sendo corroborado pelo Fis positivo observado. Foi encontrado desvio do equilíbrio de Hardy-Weinberg (HWE) em cinco loci para P. lineatus, três loci para P. mesopotamicus e dois em B. orbignyanus. A análise de bottleneck revelou excesso de heterozigotos (p < 0,05) pelo teste de Wilcoxon nas três espécies apenas no modelo Infinite Allele Model (IAM). Foi observada adequada variabilidade genética nas três populações com a presença de déficit de heterozigotos.
Palavras-chave: Brycon orbignyanus, Conservação, Microssatélites, Piaractus mesopotamicus, Prochilodus lineatus, Variabilidade genética.
Reservoirs are constructed in most of the Brazilian hydrographic basins to meet the growing demand for energy. However, they represent a barrier to migratory species and may compromise the population structure of several aquatic species (REVALDAVES et al., 2016). Long-distance migratory fishes are especially affected since, during the reproductive period, they migrate to the spawning sites located upstream and thus require free river stretches (AGOSTINHO et al., 2008). Furthermore, due to the decrease in reproduction and population growth rate, the possibility of decreasing genetic variability by bottleneck effects, genetic drift, and inbreeding increases considerably (LAROCHE; DURAND, 2004).
Previous studies have shown that the fragmentation of rivers by the presence of dams may have negative effects on the genetic structure of Brazilian migratory fish populations (RAMOS et al., 2012; ASHIKAGA et al., 2015; RIBEIRO et al., 2016). However, notably, this is not the only factor that endangers the populations of these fishes, as they may also be affected by overfishing, pollution, river banks deforestation, and habitat destruction, among others (LOPERA-BARRERO et al., 2015, 2016).
In rivers with large numbers of reservoirs, such as the Rio Grande, in São Paulo (SP), Brazil, a decline in migratory fish populations has been detected in the past decades (AGOSTINHO et al., 2016). Among the native species of economic importance in this river, Pacu (Piaractus mesopotamicus), Curimba (Prochilodus lineatus), and Piracanjuba (Brycon orbignyanus) are the major long-distance migratory species (WAGNER et al., 2012; IBGE, 2015). Few studies have evaluated the genetic diversity of these species in the Rio Grande, for example, Lopera-Barrero et al. (2015) and Ribeiro et al. (2015), who evaluated P. lineatus and P. mesopotamicus, respectively. In addition, there has been no study on B. orbignyanus populations in this river. Thus, the objective of the present study was to evaluate the genetic diversity of P. lineatus, P. mesopotamicus, and B. orbignyanus populations in the Água Vermelha Reservoir, Rio Grande, SP.
Sixty-eight caudal fin samples of ~1 cm² were collected from P. lineatus (36 samples), P. mesopotamicus (25 samples), and B. orbignyanus (7 samples). All samples were obtained from the Água Vermelha Hydroelectric Power Plant Reservoir (19°52'04"S 50°20'43"W), located in the Rio Grande (São Paulo). Sampling was supervised by the company AES Brasil. The fin fragments were stored in microtubes with 70% alcohol until the analysis. The study was approved by the Committee on Ethics in Animal Experimentation (protocol nº 203/2012 of the State University of Londrina).
DNA was extracted following the NaCl extraction protocol described by Lopera-Barrero et al. (2008). Subsequently, DNA samples were quantified in PICODROP® (Picodrop Limited, Hinxton, UK) and diluted to a final concentration of 20 ng/μL. DNA integrity was evaluated on a 1% agarose gel stained with SYBR Safe™ DNA Gel Stain (Invitrogen, Carlsbad CA, USA).
The DNA was amplified to a final reaction volume of 15 μL, using 1X of Tris-KCl buffer, 2.0 mM of MgCl2, 0.8 μM of each primer (Forward and Reverse), 0.2 mM of each dNTP, one unit (for P. mesopotamicus and B. orbignyanus) or half unit (for P. lineatus) of Platinum Taq DNA Polymerase, and 20 ng of each sample. For P. lineatus DNA samples, PCR was performed according to the methodology described by Souza et al. (2018), using eight microsatellite loci: Par12, Par14, Par21, Par43, Par80 (BARBOSA et al., 2006, 2008), Pl01, Pli30, and Pli60 (YAZBECK; KALAPOTHAKIS, 2007). For B. orbignyanus, PCR was performed according to Castro et al. (2017), using eight microsatellite loci: BoM5, BoM13 (BARROSO et al., 2003), Bh5, Bh6, Bh8, Bh13, Bh16 (SANCHES; GALETTI JUNIOR, 2006), and Par80. Finally, the amplification conditions for P. mesopotamicus was according to the methodology described by Ribeiro et al. (2015), using seven loci: Pme2, Pme4, Pme5, Pme14, Pme20, Pme28, and Pme32 (CALCAGNOTTO et al., 2001). The reactions were performed on a Veriti® thermal cycler (Applied Biosystems®, Austin, TX, USA). Subsequently, the amplification products were submitted to electrophoresis on 10% polyacrylamide gel (acrylamide: bisacrylamide - 29:1), denatured in urea (6 M) and ran in 1X TBE buffer (90 mM of Tris-Borate and 2 mM of EDTA) at 180 V and 250 mA for 8 h. Allele visualization was achieved by staining with silver nitrate (BASSAM et al.,1991). The alleles size was calculated using a 100-bp DNA ladder (Invitrogen).
The observed (Ho) and expected (He) heterozygosities, as well as the Hardy-Weinberg equilibrium (HWE) were calculated using the POPGENE 1.31 software (YEH et al., 1999). The inbreeding coefficient or fixation index (Fis) was calculated by means of the FSTAT 2.9.3 software (GOUDET, 2002). The possible reduction of the effective population size was verified with the Bottleneck 1.2.02 software (CORNUET; LUIKART, 1996), considering the mutation models: IAM (Infinite Allele Model), TPM (Two-phase Mutation Model), and SMM (Stepwise Mutation Model). Statistical significance was evaluated with the Wilcoxon test.
The P. lineatus population produced a total of 56 alleles, ranging from 3 (Pli60) to 10 (Pli30) alleles per locus (Table 1). The fragments size ranged from 160 bp (Par12) to 305 bp (Pli30) and was in accordance with the sizes previously reported (BARBOSA et al., 2006, 2008; YAZBECK; KALAPOTHAKIS, 2007). Regarding the P. mesopotamicus population, 24 alleles were produced for the seven evaluated loci. The Pme32 locus produced the lowest number of alleles, with only two alleles, while the Pme28 produced the highest, with five alleles (Table 1). Fragment size ranged from 185 bp (Pme5) to 280 bp (Pme28), similar to that found in previous studies (CALCAGNOTTO et al., 2001; POVH et al., 2010). Finally, 26 alleles were obtained in the B. orbignyanus population, from 2 (Par80) to 4 (Bh8, Bh13 and Bh16) alleles per locus (Table 1), with fragments size ranging from 90 bp (BoM13) to 225 bp (Bh5). It is important to emphasize that, for this species, heterologous primers of related species were used. These primers were previously tested and produced good amplification and reproducibility standards, agreeing with earlier research (CASTRO et al., 2017).
Mean Ho values were lower than the mean He values in the three species, being corroborated by the observed positive Fis. Mean Fis was higher for P. lineatus (0.384), where with all loci presenting heterozygous deficits, and lower for B. orbignyanus (0.044), with only five loci showing heterozygous deficits. Deviations from the Hardy-Weinberg equilibrium (p < 0.05) were observed in five (Par12, Par14, Par21, Pli30, and Pli60), three (Pme5, Pme14 and Pme20), and two (BoM13 and Bh13) loci for P. lineatus, P. mesopotamicus, and B. orbignyanus, respectively (Table 1). This deviation may have been caused by inbreeding, bottleneck effects or a combination of both. The Bottleneck analysis showed heterozygotes excess (p < 0.05) in the three populations only in the IAM, with P. lineatus presenting the lowest p value (0.01) in the Wilcoxon test (Table 2). According to Cornuet and Luikat (1996), populations that have experienced a recent reduction in effective population size tend to have an Ho greater than the He in the equilibrium of mutation and drift. In view of this, it is probable that the P. lineatus population was more influenced by recent population reductions and/or inbreeding, which resulted in the deviation of HWE observed in most loci.
Intra-population diversity indexes in the three fish species captured in the Água Vermelha Reservoir.
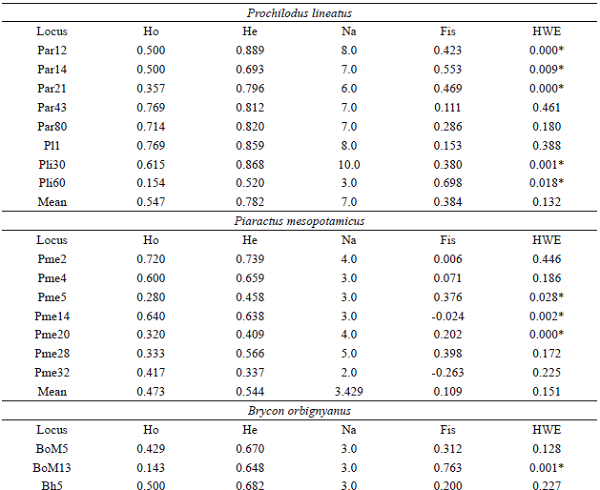
Intra-population diversity indexes in the three fish species captured in the Água Vermelha Reservoir.
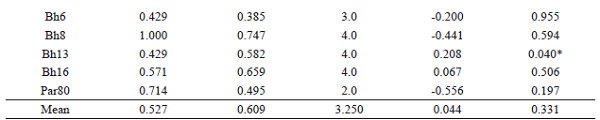
Ho: observed heterozygosityHe: expected heterozygosityNa: number of alleles Fis: coefficient of inbreedingHWE: Hardy-Weinberg equilibrium* Significant values (P < 0,05).
Despite the observed heterozygous deficit, the mean values of Ho were similar to those found by other authors in the Euclides da Cunha and Limoeiro reservoirs from the Rio Pardo, São Paulo (LOPERA-BARRERO et al., 2015) and slightly higher than those found by Carvalho-Costa et al. (2008) in Prochilodus costatus populations from the Três Marias Reservoir, Minas Gerais. Similarly, the Ho values of the P. mesopotamicus population was in accordance with that found in Pantanal populations of the same species (CALCAGNOTTO; DESALLE, 2009). Finally, the Ho of B. orbignyanus was also similar to that found by other studies for natural stocks (ASHIKAGA et al., 2015) or captive stocks (CASTRO et al., 2017). However, B. orbignyanus requires special attention since it is highly sensitive to environmental changes, which makes it endangered in certain regions (ASHIKAGA et al., 2015). Overall, it can be considered that the heterozygosity indexes were satisfactory, indicating the existence of genetic variability in the three populations of this reservoir. However, the results do not rule out the need for continuous genetic evaluations at this site, considering that the observed heterozygote deficit indicates a possible genetic decline in future generations.
Signal test for heterozygote excess/deficiency in the three fish species captured in the Água Vermelha Reservoir.
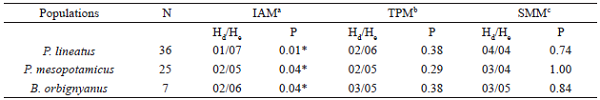
N: Number of individuals per populationa Infinite Allele Model b Two-phase Mutation Modelc Stepwise Mutation Model * Significant values from the Wilcoxon tests (P < 0.05).
The impact of anthropic factors over natural populations has been frequent in recent years (ASHIKAGA et al., 2015), with hydroelectric reservoirs being one of the causes of the reduction of natural fish stocks due to habitat fragmentation. Previous studies have shown that the presence of reservoirs may promote differences in fish genetic diversity due to the interruption of migration (BARROCA et al., 2012) while the time of dam implantation could influence the structure of these populations (LOPERA-BARRERO et al., 2016; FERREIRA et al., 2017). In addition, overfishing, pollution, and environmental destruction must also be highlighted as important factors that negatively impact these stocks and could lead to population declines. The Água Vermelha Hydroelectric Plant was inaugurated in 1978 and it is expected that, during the last 40 years of operation, geographic isolation has affected fish populations. However, further studies are needed to ascertain the magnitude of these impacts, as well as to make comparisons with samples collected downstream of the dam in order to ascertain the differentiation and genetic structure of these populations.
In conclusion, adequate genetic variability was observed in the three populations, and the presence of a heterozygous deficit was detected. Further studies are needed to evaluate the likelihood of future reductions in variability and changes in genetic structure.
References
AGOSTINHO, A. A.; GOMES, L. C.; SANTOS, N. C.; ORTEGA, J. C.; PELICICE, F. M. Fish assemblages in Neotropical reservoirs: colonization patterns, impacts and management. Fisheries Research, Amsterdam, v. 173, n. 1, p. 26-36, 2016.
AGOSTINHO, A. A.; PELICICE, F. M.; GOMES, L. C. Dams and the fish fauna of the Neotropical region: impacts and management related to diversity and fisheries. Brazilian Journal of Biology, São Carlos, v. 68, n. 4, p. 1119-1132, 2008. Supplement.
ASHIKAGA, F. Y.; ORSI, M. L.; OLIVEIRA, C.; SENHORINI, J. A.; FORESTI, F. The endangered species Brycon orbignyanus: genetic analysis and definition of priority areas for conservation. Environmental Biology of Fishes, Dordrecht, v. 98, n. 7, p. 1845-1855, 2015.
BARBOSA, A. C.; CORREA, T. C.; GALZERANI, F.; GALETTI, P. M.; HATANAKA, T. Thirteen polymorphic microsatellite loci in the Neotropical fish Prochilodus argenteus (Characiformes, Prochilodontidae). Molecular Ecology Resources, Oxford, v. 6, n. 3, p. 936-938, 2006.
BARBOSA, A. C.; GALZERANI, F.; CORRÊA, T. C.; GALETTI JUNIOR, P. M.; HATANAKA, T. Description of novel microsatellite loci in the Neotropical fish Prochilodus argenteus and cross-amplification in P. costatus and P. lineatus. Genetics and Molecular Biology, Ribeirão Preto, v. 31, n. 1, p. 357-360, 2008.
BARROCA, T. M.; SANTOS, G. B.; DUARTE, N. V. R.; KALAPOTHAKIS, E. Evaluation of genetic diversity and population structure in a commercially important freshwater fish Prochilodus costatus (Characiformes, Prochilodontidae) using complex hypervariable repeats. Genetics and Molecular Research, Ribeirão Preto, v. 11, n. 4, p. 4456-4467, 2012.
BARROSO, R. M.; HILSDORF, A. W. S.; MOREIRA, H. L. M.; MELLO, A. M.; GUIMARÃES, S. E. F.; CABELLO, P. H.; TRAUB-CSEKO, Y. M. Identification and characterization of microsatellites loci in Brycon opalinus (Cuvier, 1819) (Characiforme, Characidae, Bryconiae). Molecular Ecology Notes, Oxford, v. 3, n. 1, p. 297-298, 2003.
BASSAM, B. J.; CAETANO-ANOLLÉS, G.; GRESSHOFF, P. M. Fast and sensitive silver staining of DNA in polyacrylamide gels. Analytical Biochemistry, Bethesda, v. 196, n. 2, p. 80-83, 1991.
CALCAGNOTTO, D.; DESALLE, R. Population genetic structuring in pacu (Piaractus mesopotamicus) across the parana-paraguay basin: evidence from microsatellites. Neotropical Ichthyology, Maringá, v. 7, n. 4, p. 607-616, 2009.
CALCAGNOTTO, D.; RUSSELLO, M.; DESALLE, R. Isolation and characterization of microsatellite loci in Piaractus mesopotamicus and their applicability in other Serrasalminae fish. Molecular Ecology Notes, Oxford, v. 1, n. 4, p. 245-247, 2001.
CARVALHO-COSTA, L. F.; HATANAKA, T.; GALETTI JUNIOR, P. M. Evidence of lack of population substructuring in the Brazilian freshwater fish Prochilodus costatus. Genetics and Molecular Biology, Ribeirão Preto, v. 31, n. 1, p. 377-380, 2008.
CASTRO, P. L.; RIBEIRO, R. P.; SANTOS, S. C. A.; GOES, E. S. D. R.; SOUZA, F. P.; POVEDA-PARRA, A. R.; VARGAS, L.; URREA-ROJAS, A. M.; LOPERA-BARRERO, N. M. Cross-amplification of heterologous microsatellite markers in Piracanjuba. Ciência Rural, Santa Maria, v. 47, n. 12, p. 1-6, 2017.
CORNUET, J. M.; LUIKART, G. Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics, Rockville, v. 144, n. 4, p. 2001-2014, 1996.
FERREIRA, D. G.; SOUZA-SHIBATTA, L.; SHIBATTA, O. A.; SOFIA, S. H.; CARLSSON, J.; DIAS, J. H. P.; MAKRAKIS, S.; MAKRAKIS, M. C. Genetic structure and diversity of migratory freshwater fish in a fragmented Neotropical river system. Reviews in Fish Biology and Fisheries, London, v. 27, n. 1, p. 209-231, 2017.
GOUDET, J. FSTAT: a program to estimate and test gene diversities and fixation indices (version 2.9.3.2). Lausanne: University of Lausanne, Department of Ecology & Evolution, 2002. Disponível em: http://www2.unil.ch/popgen/softwares/fstat.htm. Acesso em: 26 jan. 2018.
INSTITUTO BRASILEIRO DE GEOGRAFIA E ESTATÍSTICA - IBGE. Produção da pecuária municipal 2015. Rio de Janeiro: IBGE, 2015. 49 p.
LAROCHE, J.; DURAND, J. D. Genetic structure of fragmented populations of a threatened endemic percid of the Rhône river: Zingel asper. Heredity, London, v. 92, n. 4, p. 329-334, 2004.
LOPERA-BARRERO, N. M.; POVH, J. A.; RIBEIRO, R. P.; GOMES, P. C.; JACOMETO, C. B.; LOPES, T. S. Comparación de protocolos de extracción de ADN con muestras de aleta y larva de peces: extracción modificada con cloruro de sodio. Ciencia e Investigación Agraria, Santiago, v. 35, n. 1, p. 76-86, 2008.
LOPERA-BARRERO, N. M.; SANTOS, S. C. A.; GOES, E. S. R.; CASTRO, P. L.; SOUZA, F. P.; POVEDA-PARRA, A. R.; CASSETA, J.; PONTILLO, B. G.; RIBEIRO, R. P. Monitoring and conservation genetics of Prochilodus lineatus wild populations of Pardo, Mogi Guaçu and Tiete rivers, São Paulo. Arquivo Brasileiro de Medicina Veterinária e Zootecnia, Belo Horizonte, v. 68, n. 6, p. 1621-1628, 2016.
LOPERA-BARRERO, N. M.; SANTOS, S. C. A.; RODRIGUEZ-RODRIGUEZ, M. P.; FORNARI, D. C.; ZANCHETA, C.; POVEDA-PARRA, A. R.; OLIVEIRA, C. A. L.; RIBEIRO, R. P. Genetic diversity of wild populations and broodstocks of curimba for restocking programs in the Tietê, Grande, Pardo and Mogi-Guaçu rivers (Brazil). Boletim do Instituto de Pesca, São Paulo, v. 41, n. 2, p. 287-304, 2015.
POVH, J. A.; RIBEIRO, R. P.; SIROL, R. N.; STREIT JUNIOR, D. P.; MOREIRA, H. L. M.; SIEWERDT, F.; LOPERA-BARRERO, N. M.; MANGOLIN, C. A.; VARGAS, L. Microsatellite analysis of the parental contribution of Piaractus mesopotamicus to the production of offspring in the semi-natural system of reproduction. Brazilian Archives of Biology and Technology, Curitiba, v. 53, n. 2, p. 389-396, 2010.
RAMOS, J. V. B.; SODRÉ, L. M. K.; ORSI, M. L.; ALMEIDA, F. S. Genetic diversity of the species Leporinus elongatus (Teleostei: Characiformes) in the Canoas Complex - Paranapanema River. Neotropical Ichthyology, Maringá, v. 10, n. 4, p. 821-828, 2012.
REVALDAVES, E.; RENESTO, E.; GOLD, J. R. Genetic variation of Prochilodus lineatus (Valenciennes, 1836) from Paraná, Baía, Miranda, and Corumbá rivers, Brazil. Genetics and Molecular Research, Ribeirão Preto, v. 15, n. 4, p. 1-11, 2016.
RIBEIRO, R. P.; LOPERA-BARRERO, N. M.; POVH, J. A.; RODRIGUEZ-RODRIGUEZ, M. P.; FORNARI, D. C.; BAUMGARTNER, G.; BAUMGARTNER, D.; SOUZA, F. P.; CASTRO, P. L.; POVEDA-PARRA, A. R. Genetic diversity of Salminus brasiliensis wild populations in downstream and upstream Cachoeira Branca, Verde River MS Brazil: a preliminary view. Semina: Ciências Agrárias, Londrina, v. 37, n. 1, p. 507-516, 2016.
RIBEIRO, R. P.; LOPERA-BARRERO, N. M.; SANTOS, S. C. A.; Rodriguez-Rodriguez, M. P.; OLIVEIRA, S. N.; ALEXANDRE FILHO, L.; VARGAS, L.; GOES, E. S. R.; CASTRO, P. L.; POVEDA-PARRA, A. R. Genetic diversity of pacu for restocking programs in the Tietê and Grande rivers, Brazil. Semina: Ciências Agrárias, Londrina, v. 36, n. 6, p. 3807-3826, 2015.
SANCHES, A.; GALETTI JUNIOR, P. M. Microsatellites loci isolated in the freshwater fish Brycon hilarii. Molecular Ecology Notes, Oxford, v. 6, n. 4, p. 1045-1046, 2006.
SOUZA, F. P.; CASTRO, P. L.; GOES, E. S. R.; RIBEIRO, R. P.; SANTOS, S. C. A.; LIMA, E. C. S.; FURLAN MURARI, P. J.; LOPERA-BARRERO, N. M. Genetic variability of Prochilodus lineatus in artificial and semi-natural reproduction. Italian Journal of Animal Science, Bologna, v. 17, n. 2, p. 321-325, 2018.
WAGNER, R. L.; MAKRAKIS, S.; CASTRO-SANTOS, T.; MAKRAKIS, M. C.; DIAS, J. H. P.; BELMONT, R. F. Passage performance of long-distance upstream migrants at a large dam on the Paraná River and the compounding effects of entry and ascent. Neotropical Ichthyology, Maringá, v. 10, n. 4, p. 785-795, 2012.
YAZBECK, G. M.; KALAPOTHAKIS, E. Isolation and characterization of microsatellite DNA in the piracema fish Prochilodus lineatus (Characiformes). Genetics and Molecular Research, Ribeirão Preto, v. 6, n. 4, p. 1026-1034, 2007.
YEH, F. C.; BOYLE, T. Y. Z.; XIYAN, J. M. POPGENE version 131: microsoft window-based freeware for population genetic analysis. Alberta: University of Alberta and Center for International Forestry Research, 1999. 29 p.
Author notes
nmlopera@uel.br
