ARTICLES
Primer design for complete sequencing of the Avastrovirus ORF2 gene
Desenho de primers para o sequenciamento completo do gene ORF2 em Avastrovirus
Primer design for complete sequencing of the Avastrovirus ORF2 gene
Semina: Ciências Agrárias, vol. 39, no. 1, pp. 411-418, 2018
Universidade Estadual de Londrina
Received: 30 March 2017
Accepted: 12 October 2017
Abstract: Avastrovirus infection is associated with enteric disease, nephritis, and hepatitis in birds. In this study, we present a protocol for the complete sequencing of the ORF2 gene of avian nephritis virus (ANV), chicken astrovirus (CAstV), and turkey astrovirus type 1 (TAstV-1) using a conventional Sanger technique. Previously and newly designed primer pairs targeting both the conserved flanking and internal regions of the ORF2 gene of these three viruses were used. The information derived from the astroviral sequences obtained in this study is fundamental for characterizing this virus and providing data regarding several aspects of disease epidemiology and prevention.
Key words: PCR, Sequencing, Avastrovirus, Avian, ORF2.
Resumo: As infecções por avastrovírus estão associados à doença entérica, nefrite e hepatite em aves. Aqui, nos presentamos um protocolo planejado para o sequenciamento completo do gene ORF2 em Avian Nephritis Vírus (ANV), Chicken Astrovirus (CAstV) e Turkey Astrovirus tipo 1 (TAstV-1), usando a técnica de sequenciamento convencional de Sanger. Foram usados primers previamente descritos e desenhados neste estudo, tendo como alvo as regiões conservadas flanqueadoras e internas dentro do gene ORF2 nos três vírus. O conhecimento destas sequencias é um elemento chave para caracterizar o vírus e prover de dados em diversos aspectos da epidemiologia e prevenção da doença.
Palavras-chave: PCR, Sequenciamento, Avastrovírus, Aves, ORF2.
Avastroviruses have been associated with enteric disease in avian species (DE BENEDICTIS et al., 2013). In addition to diarrhea, infection has been linked to kidney lesions and nephritis in chickens, fatal hepatitis in ducks, and poultry enteritis mortality syndrome (PEMS) in turkeys, chickens, and guinea fowl (PANTIN-JACKWOOD et al., 2011). The virus has been isolated worldwide, infecting different avian species (KOCI; SCHULTZ-CHERRY, 2002). In Brazil, avastroviruses have been shown to infect chickens with clinical symptoms of enteric diseases (METTIFOGO et al., 2014). Avastrovirus are non-enveloped, positive-sense, single-stranded RNA viruses, classified as a genus within the Astroviridae family (MACLACHLAN; DUBOVI, 2011; GUIX et al., 2013).
According to the International Committee on Taxonomy of Viruses (ICTV, 2014), the genus Avastrovirus can be divided into three species: Avastrovirus 1 (turkey astrovirus type 1, TAstV-1), Avastrovirus 2 (avian nephritis virus, ANV), and Avastrovirus 3 (duck astrovirus, DAstV; and turkey astrovirus type 2, TAstV-2). However, Chicken astrovirus (CAstV), a common enteric virus of broiler flocks (TODD et al., 2009), has yet to be regarded as a species within the ICTV classification.
More recently, an additional sub-division into genotypes has been suggested because of differences in the virus capsid amino acid sequence (BOSCH et al., 2011, GUIX et al., 2013). Thus far, seven genotypes of Avastrovirus have been recognized using the mean amino acid genetic distance (p-dist) calculated based on the viral capsid sequence as the cut-off; the range between and within genotypes is 0.414-0.728 and 0.047-0.299, respectively (BOSCH et al., 2011, GUIX et al., 2013). Based on this, ANV from chicken isolates belong to genotype 2 and 5, CAstV to genotype 4, and TAstV-1 (one cases of TAstV-1 infection in chickens has been reported) to genotype 1 (GUIX et al., 2013).
The ORF2 gene encodes a precursor protein, comprising of 668-742 amino acids (2007-2229 nt) and having a size of 73-81 kDa, known as the capsid protein (PANTIN-JACKWOOD et al., 2013). This protein is responsible for attachment and entry into host cells (BASS; UPADHAYAYULA, 1997), and it is a multi-domain protein containing a conserved N-terminal region (comprised of the basic S and P1 domains), a highly variable C-terminal region (spike or P2 domain), and acidic domains (KRISHNA, 2005), which are removed by host cell caspases (MENDEZ et al., 2004). Changes in receptor binding can play an important role in cross species transmission (DUBOIS et al., 2013).
Although ORF2 sequencing is a key element in the classification of Avastrovirus (BOSCH et al., 2011), PCR primers for complete gene amplification and sequence information are limited. In this study, we present a protocol for complete Sanger sequencing of the ORF2 gene of ANV, CAstV, and TAstV-1, combining new and previously designed primers.
Avastrovirus positive samples, ANV, CAstV, and TAsV-1 previously characterized by Chu et al. (2008) using ORF1b amplification and sequencing, were used for the complete amplification and sequencing of the ORF2 gene. Briefly, sixty pooled fecal samples (5-6 randomly selected fecal samples/pool; 1 pooled sample/flock) from apparent healthy broiler, layer hens, and breeders were collected from chicken flocks in Brazil and submitted for RNA extraction, using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol, and retro-transcribed with a reverse transcriptase enzyme (SuperScript III, Invitrogen). The ORF1b fragment amplification and sequencing primers and PCR conditions used for Avastrovirus species determination were as previously described by Chu et al. (2008).
For complete ORF2 gene amplification (2007-2229 nt), three different primer pairs were selected: ANV PreCAP / ANV PostCAP (TODD et al., 2011), CAstV PreCAP / CAstVPostCAP (SMYTH et al., 2012), and TAstV-1 PreCAP / TAstV-1 PostCAP; the latter was designed to pair to the flanking nucleotide regions of turkey astrovirus USA Strain (GenBank accession number NC002470; Table 1). The respective PCR mixes were prepared as follows: 1× PCR Buffer (Invitrogen), 0.2 mM of each dNTP, 0.5 µM of each primer (Table 1), 1.5 mM MgCl2, and 1 U of Platinum Taq DNA Polymerase (Invitrogen), in a 22.5 µL final volume. A total of 2.5 µL of each cDNA (ANV, CAstV, and TAstV-1) was added to each tube and submitted to 94ºC/2 min followed by 40 cycles of 94ºC/15 sec, 57ºC/30 sec, 50ºC/30 sec, or 60ºC/30 sec (ANV, CAstV, and TAstV-1, respectively), and 72ºC/2.5 min (increasing 5 sec from the cycle 25) and a final extension at 72ºC/10 min. The amplicons were analyzed by agarose gel (2%) electrophoresis, stained with SYBR Safe DNA Gel Stain (Invitrogen, Paisley, UK) according to the manufacturer’s recommendations, and visualized using UV light. The expected amplicon size is approximately 2,000-2,500 bp for ANV and CAstV and 2,000 for TAstV-1.
| Species | Primer | Sequence (5'- 3') | Annealing Tº | Size (nt) | Position1 | Gene | Reference |
|---|---|---|---|---|---|---|---|
| ANV | ANV PreCAP | ACCTTGAATCCCTGTGGGGCA | 57ºC | 4403-4423 | ORF1b / ORF2 / 3' UTR | Todd et al., 2011 | |
| ANV PostCAP | AAAAGTTAGCCAATTCAAAATTAATTC | 6915-6941 | |||||
| ANV InterCAPF | TCKCARTCRCCDGCTGA | * | 4985-5002 | ORF2 | This study | ||
| ANV InterCAPR | GTGTCNSTVRYRTYTTGCATTTG | 6114-6137 | |||||
| ANV InterCF | TCGCCWGCAGATKCGTATGG | * | 4990-5010 | ORF2 | This study | ||
| ANV Inter CR | TGCATTTGRAGAAARTAGTC | 6101-6121 | |||||
| CastV | CAstV PreCAP | TAGAGGGATGGACCGAAATATAGCAGC | 50ºC | 4980-5006 | ORF1b / ORF2 / 3' UTR | Smyth et al., 2009 | |
| CAstVPostCAP | TGCAGCTGTACCCTCGATCCTA | 7236-7257 | |||||
| CAstV InterCAPF | TWYTRMGAGGTGGCTGGTGGGT | * | 6028-6049 | ORF2 | This study | ||
| CAstV InterCAPR | TGTTTTTGCTGCTCAGGTCC | 7159-7178 | |||||
| TAstV-1 | TAstV-1 PreCAP | CGCCATGCCAAGGAAGCCGGC | 60ºC | 4771-4791 | ORF1b/ORF2 | This study | |
| TAstV-1 PostCAP | TCGATCCTACCCGGCGTGGCC | 6880-6900 |
The complete ORF2 gene PCR products of ANV and CAstV were gel excised and purified using the GeneJET Gel Extraction Kit (Thermo Scientific, CA, USA) and subsequently cloned using the InsTAclone PCR Cloning Kit (Thermo Scientific). Plasmids containing the respective inserts were sequenced using the Sanger technique in duplicates with the BigDye Terminator v.3.1 Cycle Sequencing Kit (Applied Biosystems, CA, USA), resolved using an ABI 3500 Genetic Analyzer (Applied Biosystems), and the obtained sequences were further edited with BioEdit v. 7.2.5 (HALL et al., 1995).
As the average read length obtained using the Sanger sequencing technique is approximately 800-1,000 nt (SAEED et al., 2009), the complete sequencing of the ANV and CAstV ORF2 gene required such a design of the internal primers which could help to obtain a set of overlapping products using a primer walking strategy (STUDIER, 1989; MARTIN-GALLARDO et al., 1992) (Table 1, Figure 1). Two internal primers pairs (ANV InterCAPF/ANV InterCAPR and ANV InterCF/ANV InterCR) were designed for ANV ORF2 sequencing because of the high heterogeneity observed in the obtained sequences (Table 1, Figure 1), while for CAstV, only one pair was designed (CAstV InterCAPF/CAstV InterCAPR). For TAstV-1, the ORF2 gene could be successfully sequenced with a flanking primer pair designed in this study (the total size of the TAstV-1 ORF2 gene is approximately 2050 bp). These primers were submitted to BLAST/n analysis to assess their identity and specificity, and the dimer and hairpin formations were evaluated using the Oligo Analyzer 3.1 software tool (IDT, Integrated DNA Technologies, Inc., Coralville, IA, USA). Each of the primers was directly added to the sequencing reaction mix and the previously obtained ORF2 gene plasmid inserts were used as the template. Sequences were edited using BioEdit v. 7.2.5 (HALL et al., 1995) and sequenced fragments of the same gene were assembled using CAP (Contig Assembly Program) (HUANG, 1992).
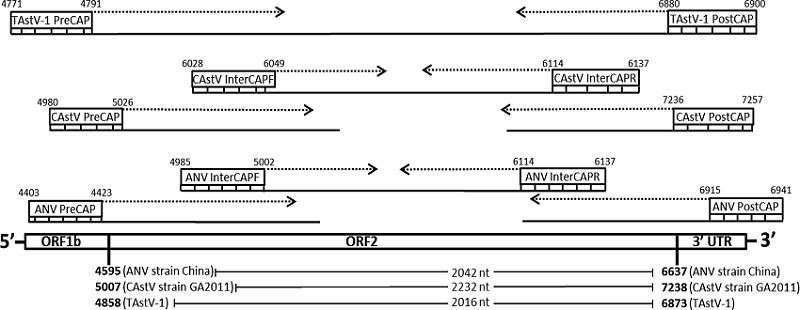
Figure 1
Schematic representation of the complete ORF2 sequencing strategy. Arrows represents the direction of nucleotide addition for each new DNA strand created. Dashed lines represent the estimated fragment to be generated by the primer. Solid lines represents the DNA fragment generated.
Amplification and sequencing of ORF1b from the 60 chicken fecal samples enabled ANV (21/60), TAstV-1 (6/60), and CAstV (2/60) species identification. Complete ORF2 gene amplification was achieved in 8/21 ANV-, 1/6 TAstV-1-, and 1/2 CAstV-positive samples. In addition, detection of ANV ORF2 polymorphic genes in the same sample was possible (via gene cloning) and up to four different ORF2 sequences were observed in the ANV-positive samples (except for two samples in which two different sequences were observed). ORF2 complete gene analysis (p-dist) showed that the ANV sequences belonged to genotype 5, previously described in chickens, and confirmed the CAstV and TAstV-1 fit with genotype 4 and 1, respectively. The p-dist percent within each genotype detected in this study was 0.027 ± 0.011, 0.018 ± 0.02, and 0.043 ± 0.01 for Genotype 5, 4, and 1, respectively (the cut off value within each genotype was 0.047-0.299). The ANV, CAstV, and TAstV-1 ORF2 sequences generated in this study were deposited with the GenBank database under accession numbers ANV (KU711050-KU711077), CAstV (KU711078), and TAstV-1 (KU711079).
In silico analysis of the designed primers using the BLAST online tool and BioEdit v. 7.2.5 software (HALL et al., 1995), demonstrated that the primers designed in this study enabled the amplification of all previously obtained Brazilian ANV, CAstV, and TAstV-1 sequences, as well as others recovered from the GenBank database. However, considering the large number of genotypes and high genetic diversity of these viruses, continuous analysis of Avastrovirus in chickens and other bird species should be performed. As expected, the BLAST/n results matched their respective viral species, with identities ranging from 65-100%, and these species showed no similarity with other enteric viruses. These relatively low values were generated mostly for the 5´ ends of the primer sequences. With regard to dimers and hairpins, although some of the primers were presented as theoretical hairpin or (self/cross) dimer formation, their respective ∆G (Free Gibbs Energy) was high, and therefore, their performance was not affected.
The previously described primers for the complete ORF2 gene (TODD et al., 2011; SMYTH et al., 2012) allowed for the amplification of 8/21 ANV- and 1/2 CAstV- positive samples. These primers, designed based on ANV and CAstV European and Asian sequences, demonstrated reduced specificity with South American samples, which can be explained by the high heterogeneity and diversity of Brazilian ANV and CAstV sequences. The complete TAstV-1 ORF2 gene was successfully amplified, although the primers were designed based on the single TAstV-1 sequence available in the GenBank database. The limited information available in the database regarding sequences and primers for ORF2 amplification restricts the analysis of this gene in Avastrovirus.
The application scope of the designed ANV primers enabled the amplification of sequences, including European, Brazilian, and Asian strains from the GenBank database, classified as genotypes 2 and 5, both previously described in chickens (IMADA et al., 2000; ZHAO et al., 2011), However, there is limited genetic information pertaining to CAstV and TAstV-1 ORF2 in the database, and reports of these viruses in Brazil and South America are very limited; thus, we cannot speculate regarding of the designed primers scope , although during standardization, we did successfully amplify and sequence one CAstV and TAstV-1 strain.
The complete sequencing of ORF2 by primer walking is a well-known strategy used in ANV and CAstV strains (TODD et al., 2011; SMYTH et al., 2012). The 3´ Rapid Amplification of cDNA Ends (RACE) technique has also been used for the complete sequencing of ORF2 in ANV strains (CHAMINGS et al., 2015). In fact, the unique complete TAstV-1 ORF2 sequence available in GenBank was obtained by RACE methodology (JONASSEN et al., 2001). Primer walking has the advantage of enabling the direct sequencing of unknown DNA, such as the ORF2 insert in a plasmid vector (MARTIN-GALHARDO et al., 1992). Moreover, the RACE technique is relatively more expensive.
The approach presented in this study provide an method to be used for genotypic and diversity studies of Avastrovirus in chickens, which are essential for the elucidation of the epidemiology of this virus.
References
BASS, D. M.; UPADHAYULA, U. Characterization of human serotype 1 astrovirus-neutralizing epitopes. Journal of Virology, Baltimore, v. 71, n. 11, p. 8666-8671, 1997.
BOSCH, A.; GUIX, S.; KRISHNA, N. K.; MÉNDEZ, E.; MONROE, S. S.; PANTIN-JACKWOOD, M.; SCHULTZ-CHERRY, S. Family Astroviridae. In: KING, A. M. Q.; LEFKOWITZ, E.; ADAMS, M. J.; CARSTENS, E. B. Virus taxonomy: classification and nomenclature of viruses (Ninth Report of the International Committee on the Taxonomy of Viruses). New York, NY: Elsevier Academic Press, 2011. p. 953-959.
CHAMINGS, A.; HEWSON, K. A.; O’ROURKE, D.; IGNJATOVIC, J.; NOORMOHAMMADI, A. H. High-resolution melt curve analysis to confirm the presence of co-circulating isolates of avian nephritis virus in commercial chicken flocks. Avian Pathology, London, v. 44, n. 6, p. 443-451, 2015.
CHU, D. K. W.; POON, L. L. M.; GUAN, Y.; PEIRIS, J. S. M. Novel astroviruses in insectivorous bats. Journal of Virology, Baltimore, v. 82, n. 18, p. 9107-9114, 2008.
DE BENEDICTIS, P.; SCHULTZ-CHERRY, S.; BURNHAM, A.; CATTOLI, G. Astrovirus infections in humans and animals - molecular biology, genetic diversity, and interspecies transmissions. Infection, Genetic and Evolution, Amsterdam, v. 11, n. 7, p. 1529-1544, 2013.
DUBOIS, R. M.; FREIDEN, P.; MARVIN, S.; REDDIVARI, M.; HEATH, R. J.; WHITE, S. W.; SCHULTZ-CHERRY, S. Crystal structure of the avian astrovirus capsid spike. Journal of Virology, Baltimore, v. 87, n. 14, p. 7853-7863, 2013.
GUIX, S.; BOSCH, A.; PINTO, R. Astrovirus taxonomy. In: SCHULTZ-CHERRY, S. (Ed.). Astrovirus research: essential ideas, everyday impacts, future directions. New York, NY: Springer Science+Business Media, 2013. p. 97-118.
HALL, A. T. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series, London, v. 41, n. 1, p. 95-98, 1995.
HUANG, X. A contig assembly program based on sensitive detection of fragment overlaps. Genomics, San Diego, v. 14, n. 1, p. 18-25, 1992.
INTERNATIONAL COMMITTEE ON TAXONOMY OF VIRUSES - ICTV. Paris, 2014. Available in <https://talk.ictvonline.org/>. Accessed at: 08 nov 2015.
IMADA, T.; YAMAGUCHI, S.; MASE, M.; TSUKAMOTO, K.; KUBO, M.; MOROOKA, A. Avian nephritis virus (ANV) as a new member of the family Astroviridae and construction of infectious ANV CDNA. Journal of Virology, Baltimore, v. 74, n. 18, p. 8487-8493, 2000.
JONASSEN, C. M.; JONASSEN, T. O.; SAIF, Y. M.; SNODGRASS, D. R.; USHIJIMA, H.; SHIMIZU, M.; GRINDE, B. Comparison of capsid sequences from human and animal astroviruses. Journal of General Virology, London, v. 82, n. 5, p. 1061-1067, 2001.
KOCI, M. D.; SCHULTZ-CHERRY, S. Avian astroviruses. Avian Pathology, London, v. 31, n. 3, p. 213-227, 2002.
KRISHNA, N. K. Identification of structural domains involved in astrovirus capsid biology. Viral Immunology, New York, v. 18, n. 1, p. 17-26. 2005.
MACLACHLAN, N. J.; DUBOVI, E. J. Fenner’s veterinary virology. 4th ed. San Diego, California: Academic Press, 2011, 499 p.
MARTIN-GALLARDO, A.; MC COMBIE, W. R.; GOCAYNE, J. D.; FITZGERALD, M. G.; WALLACE, S.; LEE, B. M. B.; LAMERDIN, J.; TRAPP, S.; KELLEY, J. M.; LIU, L. I.; DUBNICK, M.; JOHNSTON-DOW, L. A.; KERLAVAGE, A. R.; DE JONG, P.; CARRANO, A.; FIELDS, C.; VENTER, J. C. Automated DNA sequencing and analysis of 106 kilobases from human chromosome. Nature Genetics, New York, v. 1, n. 1, p. 34-39, 1992.
MENDEZ, E.; SALAS-OCAMPO, E.; ARIAS, C. F. Caspases mediate processing of the capsid precursor and cell release of human astroviruses. Journal of Virology, Baltimore, v. 78, n. 16, p. 8601-8608, 2004.
METTIFOGO, E.; NUÑEZ, L. F.; CHACÓN, J. L.; SANTANDER PARRA, S. H.; ASTOLFI-FERREIRA, C. S.; JEREZ, J. A.; JONES, R. C.; PIANTINO FERREIRA, A. J. Emergence of enteric viruses in production chickens is a concern for avian health. The Scientific World Journal, New York, v. 14, n. 1, p. 1-6, 2014.
PANTIN-JACKWOOD, M. J.; STROTHER, K. O.; MUNDT, E.; ZSAK, L.; DAY, J. M.; SPACKMAN, E. Molecular characterization of avian astroviruses. Archives of Virology, Wien, v. 156, n. 2, p. 235-244, 2011.
PANTIN-JACKWOOD, M.; TODD, D.; KOCI, M. D. Avian astroviruses. In: SCHULTZ-CHERRY, S. (Ed.). Astrovirus research: essential ideas, everyday impacts, future directions. New York, NY: Springer Science+Business Media, 2013. p. 151-180.
SAEED, F.; KHOKHAR, A.; ZAGORDI, O.; BEERENWINKEL, N. Multiple Sequence alignment system for pyrosequencing reads. In: RAJASEKARAN, S. (Ed.). Bioinformatics and computational biology: first international conference BICoB. New Orleans: Springer-Verlag Berlin Heidelberg, 2009. p. 361-375.
SMYTH, V. J.; TODD, D.; TRUDGETT, J.; LEE, A.; WELSH, M. D. Capsid protein sequence diversity of chicken astrovirus. Avian Pathology, London, v. 41, n. 2, p. 151-159, 2012.
STUDIER, F. W. A strategy for high-volume sequencing of cosmid DNAs: random and directed priming with a library of oligonucleotides. Proceedings of the National Academy of Sciences of the USA, Washington, v. 86, n. 18, p. 6917-6921, 1989.
TODD, D.; TRUDGETT, J.; SMYTH, V. J.; DONNELLY, B.; MCBRIDE, N.; WELSH, M. D. Capsid protein sequence diversity of avian nephritis virus. Avian Pathology, London, v. 40, n. 3, p. 249-259, 2011.
TODD, D.; WILKINSON, D. S.; JEWHURST, H. L.; WYLIE, M.; GORDON, A. W.; ADAIR, B. M. A seroprevalence investigation of chicken astrovirus infection. Avian Pathology, London, v. 38, n. 4, p. 301-309, 2009.
ZHAO, W.; HUA, X. G.; YUAN, L.; CUI, L.; SHAN, T. L.; DAI, X. Q.; ZHU, C. X.; YANG, Z. B. Sequence analyses of the representative Chinese-prevalent strain of avian nephritis virus in healthy chicken flocks. Avian Diseases, New York, v. 55, n. 1, p. 65-69, 2011.
Author notes
*Author for correspondence