Article
Synthesis, Characterization, and Molecular Docking of Casiopeinas® with Dipeptides as Secondary Ligand; Potential Inhibitors of SARS-Cov-2 Transcendental Proteins
Síntesis, caracterización y acoplamiento molecular de Casiopeinas® con dipéptidos como ligando secundario; inhibidores potenciales de proteínas trascendentales del SARS-Cov-2
Synthesis, Characterization, and Molecular Docking of Casiopeinas® with Dipeptides as Secondary Ligand; Potential Inhibitors of SARS-Cov-2 Transcendental Proteins
Journal of the Mexican Chemical Society, vol. 68, no. 1, pp. 29-55, 2024
Sociedad Química de México A.C.
Received: 11 August 2022
Accepted: 15 March 2023
Abstract: In this work, the synthesis and characterization of fourteen Casiopeinas® are presented, whose general formulae is [Cu(N-N)(L-L)]NO3, where N-N are 2,2´-bipirydine and 1,10-phenanthroline and some of its methylated derivatives, L-L represent the dipeptides L-Tyrosil-Glycinate or Glycil-L-Tyrosinate. Spectroscopic characterization and DFT studies determine the square planar geometry for the coordination compounds, as well as the influence of the dipeptide on the molecular arrangement of ternary copper(II) compounds. In addition, a molecular docking study was carried out against transcendental proteins of the SARS-CoV-2 virus such as main protease (Mpro) and the RBD Spike-ACE2 complex. Docking studies indicate that all compounds can produce stable adducts with Mpro, obtaining ΔGU values (-9.57 to -6.62 Kcal/mol) similar and superior to those presented by the reference inhibitors [boceprevir (-8.44 Kcal/mol) and remdesivir (-6.62 kcal/mol)], while for the RBD Spike-ACE2 complex obtaining ΔGU values of five (-6.69 to -4.61 in C-terminal region) and three (-8.27 to -6.34 in central region) orders of magnitude higher than those presented by the controls (Boceprevir: ΔGU=-1.98 in C-terminal, ΔGU=-4.97 in central region, Remdesivir: ΔGU=Non interactions in C-terminal, ΔGU=-3.37 in central region). π-alkyl interactions, π -cation, π -stacking, as well as hydrogen bonds and salt bridge bonds occur between the proteins and Casiopeinas®. In Mpro, interactions occur in aminoacids that are part of the enzymes catalytic site. Casiopeinas® interact at the interface of the RDB Spike-ACE2 complex in both, C-terminal and central regions. The obtained results position Casiopeinas® as potential candidates protein inhibitors of the virus that causes COVID-19.
Keywords: SARS-COV2, Casiopeinas®, metallodrugs, molecular docking, peptides, copper.
Resumen: En este trabajo, se presenta la síntesis y caracterización de 14 Casiopeinas®, cuya fórmula general es [Cu(N-N)(L-L)]NO3, donde N-N son 2,2´-bipiridina y derivados metilados o 1,10-fenantrolina y análogos con grupos metilo, L-L representan a los dipéptidos L-Tirosil-Glicinato o Glicil-L-Tirosinato. Mediante estudios espectroscópicos y de DFT determinan la geometría cuadrada de los compuestos sintetizados, así como la influencia del dipéptido en el arreglo molecular de los compuestos ternarios de cobre(II). Complementariamente, se realizó un estudio de docking molecular ante proteínas trascendentales del virus SARS-CoV-2 como lo son la proteasa principal (MPro o nsps-3) y el complejo RBD Spike-ACE2. Estudios de docking molecular con la MPro se obtuvieron valores de ΔGU (-9.57 a -6.629) kcal/mol, valores que son similares y superiores a los presentados por los inhibidores de referencia [boceprevir (-8.44 kcal/mol) y remdesivir (-6.62 kcal/mol)], mientras que para el complejo RBD Spike-ACE2 se obtuvieron valores de ΔGU de cinco (-6.69 to -4.61 en región C-terminal) y tres (-8.27 to -6.34 en región central) órdenes de magnitud superiores respectivamente a los presentados por los inhibidores de referencia (Boceprevir: ΔGU=-1.98 en C-terminal, ΔGU=-4.97 en region central, Remdesivir: ΔGU=Sin interacciones en C-terminal, ΔGU=-3.37 en region central). Interacciones π-alquilo, π-catión, apilamiento π, así como enlaces puentes de hidrogeno y puentes de sal se producen entre las proteínas y Casiopeinas® estudiadas. En Mpro, las interacciones ocurren en aminoácidos que forman parte del sitio catalítico de la enzima. Las Casiopeinas® interactúan en la interfase del complejo RBD Spike-ACE2 tanto en la región C-terminal como en la región central. Los resultados obtenidos, posicionan a las Casiopeinas® como potenciales candidatos a inhibidores proteicos del virus causante de la COVID-19.
Palabras clave: SARS-COV2, Casiopeinas®, metalofármacos, acoplamiento molecular, péptidos, cobre.
Introduction
In December 2019 in Wuhan China, a disease like pneumonia of idiopathic origin emerged [1]. Shortly after the causative agent was identified, that illness is transmitted by a virus and spreads in the air. This pathogen belongs to the β-coronavirus family, specifically called SARS-CoV-2 for its 89.1 % analogy with SARS-CoV [2], SARS-CoV-2 produce the COVID-19 disease, which was propagated throughout the world, on March 11, 2020, WHO being declared a pandemic. As of the date of writing, the pandemic still has about 761,071,826 cases and 6,879,677 deaths worldwide [3].
Coronaviruses are single-stranded, positive-sense, enveloped, unsegmented RNA-viruses [4] in which the genomic analysis determined for SARS-CoV-2 presents the following order of genes (5´to 3´): Replicase ORF1a1b, Spike (S), Envelope (E), Membrane (M) and Nucleocapsid (N). ORF1ab encodes 16 non-structural proteins (nsps) and 8 accessory proteins (3a, 3b, p6, 7a, 7b, 8b, 9b and orf14), the genes S, E, M and N encodes structural proteins with the same name. [5].
Considering the natural history of the infection process, the Spike protein is transcendent in the pathogenic process, since it is the antigenic determinant present in SARS-CoV-2, which must interact with the angiotensin-converting enzyme type II (ACE2) to be able to enter the host cells, where it will subsequently replicate the genomic material for the synthesis of viral RNA, structural and non-structural proteins [6]. Spike is a homotrimeric glycoprotein with two subunits (S1 and S2) in each monomer, the S1 subunit contains the receptor binding domain (RBD) that is expressed on the surface of the viral membrane and participates in the cellular recognition of the ACE2. ACE2 is a single-pass transmembrane protein involved in the regulation of vasoconstriction and blood pressure, ACE2 is expressed in cells of the lungs, kidneys, heart, and enterocytes of the small intestinal. [7] Particularly, the contact zones between the Spike protein RBD and ACE2 are divided into three clusters, the N-terminal region Gln498, Thr500 and Asn501 of Spike that bind via hydrogen bridge bond with Tyr41, Gln42, Lys353 and Arg 356 of ACE2. In the central region, Lys417 and Tyr453 of Spike interact with Asp30 and His34 of ACE2. Finally, in the C-terminal region, Gln474 and Phe486 of the RBD join Gln24 and Met 82 of angiotensin-converting enzyme type II [8].
The gene expression of ORF1a1b generates 16 non-structural proteins (nsps). One of them, nsps-5, also known as main protease [MPro], or 3C-like protease [3CLPro], is essential for the viral cycle. The Mpro is responsible for the digestion of 11 conserved sites of two polyproteins (pp1a and pp1b) [9]. The functional importance of this enzyme is fundamental to the viral cycle, it is needed for SARS-CoV-2 replication and primary transcription of the viral genetic material [10]. The Mpro active site contains a catalytic dyad (His41 and Cys145) and a substrate union triad (Met49, His163 and Gln189) [11]. This protein has been located at SARS-CoV, human coronavirus (HCoV229E) as well as porcine transmissible gastroenteritis virus (TGEV) [12].
Given the lack of drugs that can inhibit or even eliminate the virus, the scientific community have proposed strategies to be used in viral chemotherapy considering the various molecular targets involved in the viral cycle, both in vitro [13] and in silico [14]. Several drugs can be used under recycling to be directed against viral elements such as Mpro [15], Spike protein [16], and ACE2, as well as other non-structural proteins such as PLPro [17] and RNA-dependent of RNA polymerase (RdRp) [18]. A powerful tool for the analysis of candidates to be used as potential anti-SARS-CoV-2 drugs is molecular docking in which a diversity of molecules such as boceprevir (ΔGU=-8.3 Kcal/mol IC50 =8.0 μM, Ki=1.19 μM) [19-20] as Mpro active sites inhibitor [19], remdesivir (ΔGU=-10.1 Kcal/mol in RdRp) [21-22] (moldock score=-111.07 in Spike Protein central region) [22], Losartan [23], peptide derivatives [24], as well as coordination [25] and organometallics [26] compounds have been analyzed. The importance of targeting molecules to transcendent proteins is crucial to found molecules able to eliminate the SARS-CoV-2, as an example of this kind of compounds is PF-07321332 (PAXLOVID), the first approved drug by FDA, that is a Mpro inhibitor [27].
Several authors agree that coordination compounds represent a versatile option for drug development, since it physicochemical features can be tuned depending on the metal center, its oxidation number, the characteristics of ligand, coordination number, geometry around the metal, stereochemistry of the species formed to the aim to improve its biological activity. Medicinal inorganic chemistry has contributed to the development of metallodrugs with compounds that in their chemical structure present the metallic elementals to treat various ailments such as cancer, schizophrenia, stomach upset, diabetes, antimicrobials, antiparasitic and antiviral agents [28]. For instance, several metallodrugs have been tested against the proteins involved in the SARS-CoV-2 viral cycle. Gold compounds [29] have presented evidence of inhibition of ACE2 (IC50=16.2-25.0) μM and PLPro (IC50= 0.96 - >100 μM). Ebselen [30], a compound of selenium is capable to inhibit MPro (IC50=0.67±0.09 μM) and PLPro (IC50=2.4 μM), while rhenium compounds [31] work as Mpro inhibitors in the range of 7.5-9.1 μM.
On the other hand, Casiopeinas® are ternary copper(II) compounds with a general formulae [Cu(N-N)(L-L)]n+(NO3)n n=1 or 2, where primary donor (N-N fragment) is an aromatic substituted diimine (2,2´-bipyridine or 1,10-phenanthroline) and secondary donors (L-L groups) are several bidentate chelates. These compounds have presented evidence of antitumoral [32], antiparasitic [33] and antimicrobial activity [34]. In vitro, in vivo [35] and in silico studies [36], suggest that Casiopeinas® present diverse mechanisms of action such as cytotoxic and low genotoxic damage, induction of apoptosis [37] and autophagy [38] mediated by interactions with biomolecules as DNA [39], glutathione [40], albumin [32] as well as the generation of reactive oxygen species due to changes in the oxidation number between the redox CuII/CuI pair [41].
Recently we have published a work with the interaction of several Casiopeinas® with proteins such as Mpro of SARS-CoV-2. [42] The interactions with biomolecules can be potentiated by incorporating a secondary ligand that have both donor and acceptor groups. For that purpose, dipeptides [43] can be an interesting option, since their physicochemical features can provide structural diversity with the pH-dependent coordination modes [44]; likewise, the nature of the side chain provides donor/acceptor groups capable to enhance their interaction efficiency [45], as well as changes in the metal center oxidation state when interacting with active redox centers [46], favoring several π interactions in residues with aromatic groups (as tyrosine), aliphatic groups (as glycine), cation, stacking [47], they can also present hydrogen bridge and salt bridge bonds [48]. All these interesting features have promoted the flourishing of the field of copper(II)-peptide compounds. These compounds have been developed for diverse therapeutic purposes such as anticancer and antimicrobial agents, care for chronic degenerative diseases such as Alzheimer and Parkinson [49], as well as theranostic agents for imaging studies [50].
In this work, the synthesis and characterization of fourteen compounds with copper(II), diimines derived from 1,10-phenanthroline or 2,2´-bipyridine with the ligands L-Tyrosil-Glycinato (YG) or Glycil-L-Tyrosinate (GY) are presented. Density functional theory (DFT) and molecular docking analysis were performed to investigate the Mpro and Spike-ACE2 complex inhibitor behavior of the Casiopeinas®. In the Mpro, the catalytic and union sites were analyzed and for Spike-ACE2 complex, the C-terminal and central regions were studied. Table 1 contains Casiopeinas® presented in this work.
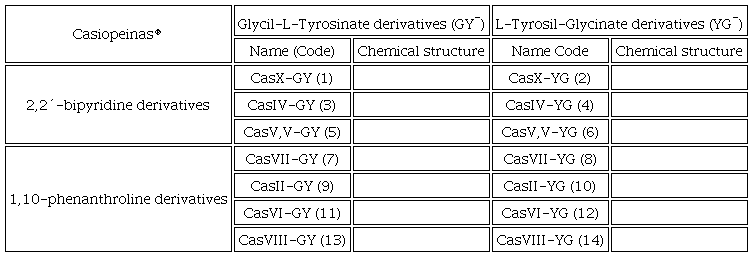
In all synthesized coordination compounds, nitrate was used as a counterion.
Experimental
Materials and methods
The copper(II) nitrate hemi(pentahydrate) - Cu(NO3)2•2.5H2O-, Glycyl-L-Tyrosine -C11H14N2O4- (GY), L-Tyrosil-Glycine -C11H14N2O4- (YG), 2,2´-bypiridine -C10H8N2-, 4,4´-dimethyl-2,2´-bipyridine -C12H12N2-, 5,5´-dimethyl-2,2´-bipyridine -C12H12N2-, 1,10-phenanthroline -C12H8N2-, 4,7-dimethyl-1,10-phenanthroline -C14H12N2-, 5,6-dimethyl-1,10-phenanthroline -C14H12N2-, 3,4,7,8-tetramethyl-1,10-phenanthroline -C16H16N2-, sodium hydroxide and solvents were purchased from Merck were used without further purification.
The FT-IR* spectra of coordination compounds were recorded in KBr pellets in the 4000-400 cm-1 range on Thermo Nicolet Avatar 320 FT-IR Spectrometer. The conductivity of the complexes in water (10-3 M) solutions was measured at 298K on a conductivity meter Jenway Conductivity and pH meter 4330, the conductivity cell constant was 1.0 cm-1. FAB (+) mass spectra were recorded on the MStation JMS-700 NBA solutions as a solvent. Magnetic susceptibility measurements were performed on powder samples at 298K on Sherwood-Scientific MK magnetic balance using Gouy´s method. The electronic spectra (UV-Vis) were recorded using Cary 60 UV-Vis Spectrophotometer in 200-1000 nm range using water as solvent. Electronic paramagnetic resonance (EPR) spectra were performed on a Jeol JES-TE300, X-band spectrophotometer (ν=9.60GHz), center field=300.00±75.00mT, PW=1mW, width=0.1mT, amplitude=250, sweep time=2min, all samples for EPR studies were prepared in methanol solutions glass at 77 K in a cold finger, with a copper(II) ternary compounds concentration of 10-3 M.
General procedure for synthesis of Casiopeinas®
For the synthesis of ternary copper(II) compounds, a modification was made to the method reported by Ruiz-Azuara and coworkers[51], in which the amount equivalent to 1 mmol of copper(II) nitrate hemihydrate was weighed, which was dissolved in 10 mL of methanol, the equivalent amount to 1 mmol of the diimine was weighed, it was dissolved in 10 mL of methanol. Diimine solution was added under slow drip and stirring at room temperature to the copper(II) nitrate solution, (Reaction mixture A). On the other hand, the equivalent amount to 1mmol of the dipeptide (GY or YG) was weighed, which was dissolved in 10 mL of methanol, under agitation. 1 mL was taken from a 1 M solution of NaOH, which was added under drip and gentle stirring to the dipeptide, to obtain the sodium salt of the dipeptide (Reaction mixture B). Once the reaction mixtures A and B were obtained, we proceeded under stirring and dripping, to add mixture B to mixture A. The solution was vacuum filtered and washed with cold water (3 times with 3 mL). The general synthesis of the complexes 1-14 is shown in Scheme 1.
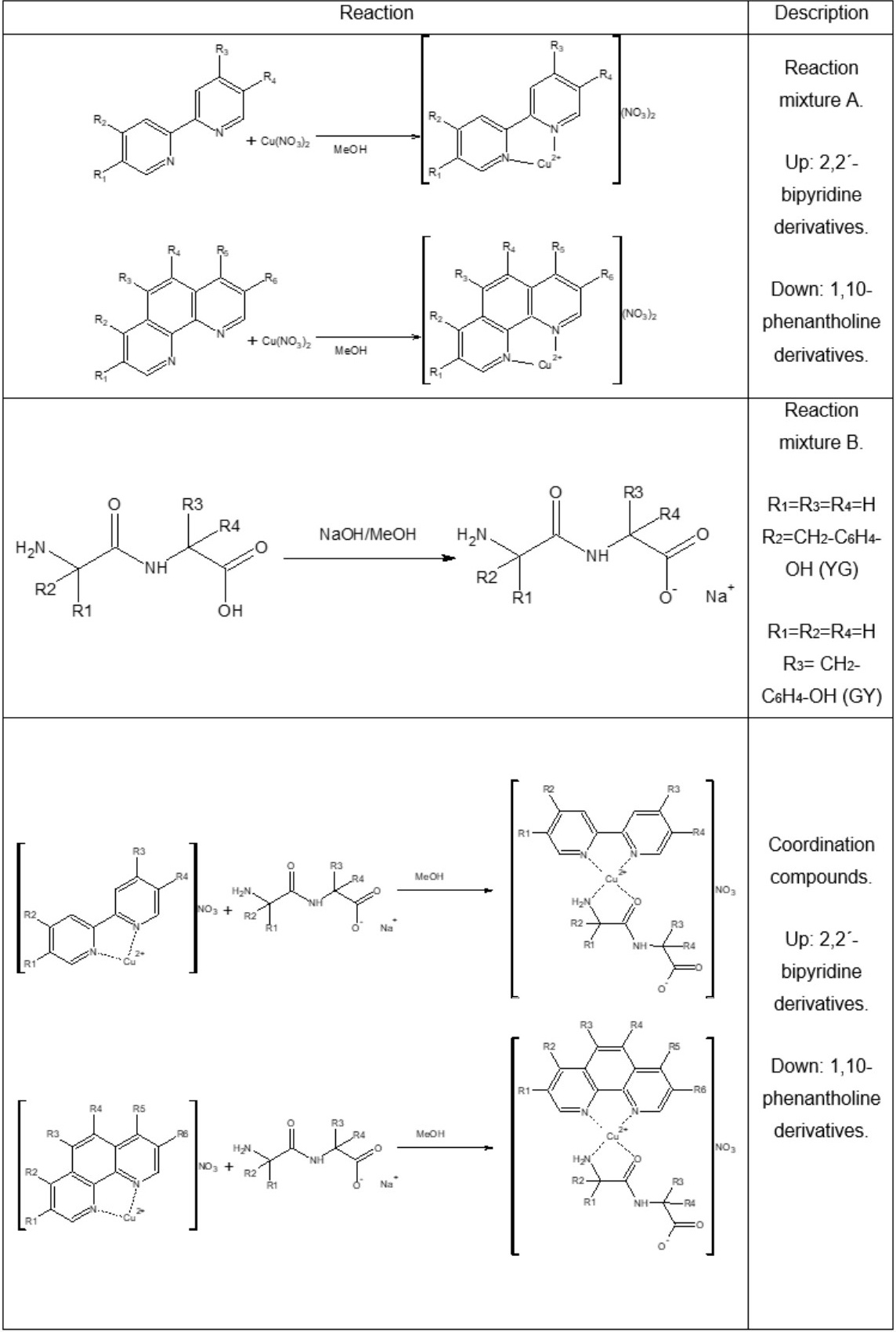
Scheme 1
Synthesis of Casiopeinas®.
Synthesis of 1. (2,2´-bipyridine) (Glycyl-L-Tyrosinate) copper(II) nitrate. (CasX-GY). Aqua blue powder; 90 % yield; IR KBr pellets λmax= 3427 (OH), 3249 (NH2, NH), 3016, 2948 (CH2, CH3), 1608 (C=C), 1685 (CONR), 1631 (C=N), 835, 721 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z (%Int) 458 [M]+ ; Λ (H2O) 132.63 Scm2mol-1; μeff: 1.86 B.M; UV-Vis (H2O): λmax (ε) 226 nm (29007 M-1cm-1) π(π*, 247 nm (25213 M-1cm-1) π(π*, 311 nm (16074 M-1cm-1) MLCT, 635 nm (105.52 M-1cm-1) b2g(b1g*; EPR parameters g‖=2.194, g┴=2.058, A‖=153.46x10-4 cm-1, A┴=84.24x10-4 cm-1; elemental analysis calculated for [CuC21H21N4O4]NO3•H2O (%). 46.97 C, 4.32 H, 13.04 N; found: 46.68 C, 4.39 H, 12.74 N; molecular weight 536.98 g/mol.
Synthesis of 2. (2,2´-bipyridine) (L-Tyrosil-Glycinate) copper(II) nitrate. (CasX-YG). Blue powder; 93 % yield; %yield; IR KBr pellets λmax= 3430 (OH), 3252, 3222 (NH2, NH), 3043, 2950 (CH2, CH3), 1602 (C=C), 1687 (CONR), 1631 (C=N), 827, 729 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z 458 [M]+; Λ (H2O) 141.85 Scm2mol-1; μeff: 1.86 B.M; UV-Vis (H2O): λmax (ε) 201 nm (611591 M-1cm-1) π(π*, 250 nm (147324 M-1cm-1) π(π*, 311 nm (151624 M-1cm-1) MLCT, 623nm (48.95 M-1cm-1) b2g(b1g*; EPR parameters g‖=2.189, g┴=2.059, A‖=136.31x10-4 cm-1, A┴=115.44x10-4 cm-1; elemental analysis calculated for [CuC21H21N4O4]NO3•2H2O (%). 45.45 C, 4.54 H, 12.62 N; found: 45.46 C, 4.61 H, 12.14 N; molecular weight 554.99g/mol.
Synthesis of 3. (4,4´-dimethyl-2,2´-bipyridine) (Glycyl-L-Tyrosinate) copper(II) nitrate. (CasIV-GY). Blue powder; 87 % yield; %yield; IR KBr pellets λmax= 3419 (OH), 3251 (NH2, NH), 3026, 2939 (CH2, CH3), 1602 (C=C), 1685 (CONR), 1622 (C=N), 827, 729 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z 485 [M]+; Λ (H2O) 141.49 Scm2mol-1; μeff: 2.02 B.M; UV-Vis (H2O): λmax (ε) 207 nm (131656 M-1cm-1) π(π*, 228 nm (58866 M-1cm-1) π(π*, 297 nm (24253 M-1cm-1) n(π*, 307 nm (22622 M-1cm-1) MLCT, 631nm (43.70 M-1cm-1) b2g(b1g*; EPR parameters g‖=2.186, g┴=2.077, A‖=123.05x10-4 cm-1, A┴=126.22x10-4 cm-1; elemental analysis calculated for [CuC23H25N4O4]NO3•H2O (%). 48.85 C, 4.81 H, 12.39 N; found: 48.85 C, 4.81 H, 12.39 N; Molecular weight 565.03 g/mol.
Synthesis of 4. (4,4´-dimethyl-2,2´-bipyridine) (L-Tyrosil-Glycinate) copper(II) nitrate. (CasIV-YG). Blue powder; 85 % yield; IR KBr pellets λmax= 3467 (OH), 3259, 3222 (NH2, NH), 2950 (CH2, CH3), 1618 (C=C), 1685 (CONR), 1654 (C=N), 827, 727 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z 485 [M]+; Λ (H2O) 152.99 Scm2mol-1; μeff: 1.90 B.M; UV-Vis (H2O): λmax (ε) 208 nm (126201 M-1cm-1) π(π*, 248 nm (26504 M-1cm-1) π(π*, 314 nm (29583 M-1cm-1) MLCT, 637nm (47.34 M-1cm-1) b2g(b1g*; EPR parameters g‖=2.121, g┴=2.058, A‖=139.77x10-4 cm-1, A┴=101.21x10-4 cm-1; elemental analysis calculated for [CuC23H25N4O4]NO3•H2O (%). 48.85 C, 4.81 H, 12.39 N; found: 48.62 C, 4.97 H, 12.62 N; Molecular weight 565.03 g/mol.
Synthesis of 5. (5,5´-dimethyl-2,2´-bipyridine) (Glycyl-L-Tyrosinate) copper(II) nitrate. (CasV,V-GY). Blue green powder; 92 % yield; IR KBr pellets λmax= 3421 (OH), 3251 (NH2, NH), 3043, 2948 (CH2, CH3), 1608 (C=C), 1691 (CONR), 1623 (C=N), 827, 725 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z 485 [M]+; Λ (H2O) 152.72 Scm2mol-1; μeff: 1.81 B.M; UV-Vis (H2O): λmax (ε) 203 nm (128211 M-1cm-1) π(π*, 256 nm (26607 M-1cm-1) π(π*, 319 nm (29583 M-1cm-1) MLCT, 634nm (37.06 M-1cm-1) b2g(b1g*; EPR parameters g‖=2.176, g┴=2.059, A‖=142.78x10-4 cm-1, A┴=77.17x10-4 cm-1; elemental analysis calculated for [CuC23H25N4O4]NO3•CH3OH (%). 49.78 C, 5.05 H, 12.09 N; found: 49.51 C, 4.63 H,11.93 N; molecular weight 579.06 g/mol.
Synthesis of 6. (5,5´-dimethyl-2,2´-bipyridine) (L-Tyrosil-Glycinate) copper(II) nitrate. (CasV,V-YG). Green powder; 88 % yield; IR KBr pellets λmax= 3473 (OH), 3253, 3222 (NH2, NH), 3010, 2942 (CH2, CH3), 1618 (C=C), 1683 (CONR), 1654 (C=N), 827, 727 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z 485 [M]+; Λ (H2O) 158.37 Scm2mol-1; μeff: 2.04 B.M; UV-Vis (H2O): λmax (ε) 206 nm (120689 M-1cm-1) π(π*, 257 nm (38606 M-1cm-1) π(π*, 319 nm (25904 M-1cm-1) MLCT, 620 nm (49.41 M-1cm-1) b2g(b1g*; EPR parameters g‖=2.258, g┴=2.023, A‖=149.59x10-4 cm-1, A┴=109.61x10-4 cm-1; elemental analysis calculated for [CuC23H25N4O4]NO3•H2O (%). 48.85 C, 4.81 H, 12.39 N; found: 48.45 C, 4.47 H, 12.35 N; molecular weight 565.03 g/mol.
Synthesis of 7. (1,10-phenanthroline) (Glycyl-L-Tyrosinate) copper(II) nitrate. (CasVII-GY). Dark blue powder; 85 % yield; IR KBr pellets λmax= 3428 (OH), 3253 (NH2, NH), 3012, 2925 (CH2, CH3), 1610 (C=C), 1685 (CONR), 1625 (C=N), 827, 721 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z 482 [M]+; Λ (H2O) 139.40 Scm2mol-1; μeff: 1.86 B.M; UV-Vis (H2O): λmax (ε) 204 nm (175182 M-1cm-1) π(π*, 222 nm (166119 M-1cm-1) π(π*, 272 nm (87823 M-1cm-1) n(π*, 294 nm (27718 M-1cm-1) MLCT, 631nm (59.78 M-1cm-1) b2g(b1g*; EPR parameters g‖=2.135, g┴=2.062, A‖=152.28x10-4 cm-1, A┴=68.72x10-4 cm-1; elemental analysis calculated for [CuC23H21N4O4]NO3•0.5H2O (%). 50.04 C, 4.02 H, 12.69 N; found: 49.94 C, 4.03 H, 12.95 N; molecular weight 551.99 g/mol.
Synthesis of 8. (1,10-phenanthroline) (L-Tyrosil-Glycinate) copper(II) nitrate. (CasVII-YG). Green jade powder; 92 % yield; IR KBr pellets λmax= 3442 (OH), 3252, 3223 (NH2, NH), 3025, 2946 (CH2, CH3), 1631 (C=C), 1685 (CONR), 1631 (C=N), 848, 721 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z 482 [M]+; Λ (H2O) 157.40 Scm2mol-1; μeff: 2.01 B.M; UV-Vis (H2O): λmax (ε) 204 nm (164928 M-1cm-1) π(π*, 225 nm (102435 M-1cm-1) π(π*, 276 nm (59436 M-1cm-1) n(π*, 296 nm (21031 M-1cm-1) MLCT, 631nm (59.78 M-1cm-1) b2g(b1g*; EPR parameters g‖=2.144, g┴=2.062, A‖=153.13x10-4 cm-1, A┴=87.20x10-4 cm-1; Elemental analysis calculatedd for [CuC23H21N4O4]NO3•2H2O(%). 47.71 C, 4.35 H, 12.10 N; found: 47.95 C, 4.73 H, 12.18 N; molecular weight 579.02 g/mol.
Synthesis of 9. (4,7-dimethyl-1,10-phenanthroline) (Glycyl-L-Tyrosinate) copper(II) nitrate. (CasII-GY). Blue green powder; 91 % yield; IR KBr pellets λmax= 3419 (OH), 3257 (NH2, NH), 2998, 2923 (CH2, CH3), 1613 (C=C), 1683 (CONR), 1622 (C=N), 827, 723 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z 510 [M]+; Λ (H2O) 136.10 Scm2mol-1; μeff: 1.80 B.M; UV-Vis (H2O): λmax 204 nm (120466 M-1cm-1) π(π*, 226 nm (60819 M-1cm-1) π(π*, 274 nm (56496 M-1cm-1) n(π*, 307 nm (10069 M-1cm-1) MLCT, 658nm (42.55 M-1cm-1) b2g(b1g*; EPR parameters g‖=2.198, g┴=2.060, A‖=155.22x10-4 cm-1, A┴=76.99x10-4 cm-1; elemental analysis calculated for [CuC25H25N4O4]NO3•2H2O (%). 49.96 C, 4.82 H, 11.66 N; found: 50.23 C, 4.83 H, 12.02 N; molecular weight 607.07 g/mol.
Synthesis of 10. (4,7-dimethyl-1,10-phenanthroline) (L-Tyrosil-Glycinate) copper(II) nitrate. (CasII-YG). Blue green powder; 93 % yield; IR KBr pellets λmax= 3436 (OH), 3265 (NH2, NH), 3014, 2950 (CH2, CH3), 1610 (C=C), 1685 (CONR), 1636 (C=N), 848, 723 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z 510 [M]+; Λ (H2O) 144.80 Scm2mol-1; μeff: 2.08 B.M; UV-Vis (H2O): λmax (ε) 208 nm (97547 M-1cm-1) π(π*, 223 nm (54800 M-1cm-1) π(π*, 274 nm (46290 M-1cm-1) n(π*, 305 nm (8989 M-1cm-1) MLCT, 631nm (50.44 M-1cm-1) b2g(b1g*; EPR parameters g‖=2.189, g┴=2.059, A‖=156.37x10-4 cm-1, A┴=85.27x10-4 cm-1; elemental analysis calculated for [CuC25H25N4O4]NO3•2.5H2O (%). 48.74 C, 4.91 H, 11.37 N; found:48.68 C, 4.76 H, 11.44 N; molecular weight 616.08 g/mol.
Synthesis of 11. (5,6-dimethyl-1,10-phenanthroline) (Glycyl-L-Tyrosinate) copper(II) nitrate. (CasVI-GY). Pale blue powder; 85 % yield; IR KBr pellets λmax= 3419 (OH), 3245 (NH2, NH), 3012, 2929 (CH2, CH3), 1604 (C=C), 1700 (CONR), 814, 734 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z 510 [M]+; Λ (H2O) 140.39 Scm2mol-1; μeff: 2.14 B.M; UV-Vis (H2O): λmax (ε) 245 nm (84722 M-1cm-1) π(π*, 287 nm (70841 M-1cm-1) n(π*, 315 nm (21602 M-1cm-1) MLCT, 628nm (51.45 M-1cm-1) b2g(b1g*; EPR parameters g‖=2.177, g┴=2.076, A‖=140.56x10-4 cm-1, A┴=83.09x10-4 cm-1; elemental analysis calculated for [CuC25H25N4O4]NO3•2CH3OH (%). 51.06 C, 5.24 H, 11.03 N; found: 51.21 C, 5.38 H, 10.67 N; molecular weight 635.12 g/mol.
Synthesis of 12. (5,6-dimethyl-1,10-phenanthroline) (L-Tyrosil-Glycinate) copper(II) nitrate. (CasVI-YG). Cobalt blue powder; 89 % yield; IR KBr pellets λmax= 3429 (OH), 3259, 3216 (NH2, NH), 3016, 2944 (CH2, CH3), 1616 (C=C), 1685 (CONR), 1633 (C=N), 810, 727 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z 510 [M]+; Λ (H2O) 158.45 Scm2mol-1; μeff: 1.98 B.M; UV-Vis (H2O): λmax (ε) 207 nm (172965 M-1cm-1) π(π*, 245 nm (73651 M-1cm-1) π(π*, 285 nm (68066 M-1cm-1) n(π*, 307 nm (27157 M-1cm-1) MLCT, 627nm (49.41 M-1cm-1) b2g(b1g*; EPR parameters g‖=2.193, g┴=2.059, A‖=134.69x10-4 cm-1, A┴=118.27x10-4 cm-1; elemental analysis calculated for [CuC25H25N4O4]NO3•2H2O (%). 49.46 C, 4.81 H, 11.54 N; found: 49.73 C, 4.41 H, 12.32 N; molecular weight 607.07 g/mol.
Synthesis of 13. (3,4,7,8-tetramethyl-1,10-phenanthroline) (Glycyl-L-Tyrosinate) copper(II) nitrate. (CasVIII-GY). Green powder; IR KBr pellets λmax= 3428 (OH), 3249 (NH2, NH), 3016, 2933 (CH2, CH3), 1616 (C=C), 1678 (CONR), 1636 (C=N), 823, 721 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z 538 [M]+; Λ (H2O:MeOH 99:1) 151.28 Scm2mol-1; Λ (MeOH) 102.29 Scm2mol-1; μeff: 2.16 B.M; UV-Vis (H2O): λmax (ε) 211 nm (118383 M-1cm-1) π(π*, 228 nm (94981 M-1cm-1) π(π*, 284 nm (68173 M-1cm-1) n(π*, 307 nm (17808 M-1cm-1) MLCT; EPR parameters g‖=2.266, g┴=2.007, A‖=145.35x10-4 cm-1, A┴=79.81x10-4 cm-1; elemental analysis calculated for [CuC27H29N4O4]NO3•H2O (%). 52.55 C, 5.06 H, 11.35 N; found: 52.37 C, 5.11 H, 11.67 N; molecular weight 617.10 g/mol.
Synthesis of 14. (3,4,7,8-tetramethyl-1,10-phenanthroline) (L-Tyrosil-Glycinate) copper(II) nitrate. (CasVIII-YG). Green powder; 87 % yield; IR KBr pellets λmax= 3454 (OH), 3279, 3232 (NH2, NH), 3072, 3000 (CH2, CH3), 1609 (C=C), 1675 (CONR), 1625 (C=N), 812, 723 (CHar), 1384 (NO3-) cm-1: FAB(+)-MS m/z 538 [M]+; Λ (H2O:MeOH 99:1) 158.46 Scm2mol-1: Λ (MeOH) 98.27 Scm2mol-1; μeff: 2.06 B.M; UV-Vis (H2O): 212 nm (29705 M-1cm-1) π(π*, 227 nm (19940 M-1cm-1) π(π*, 284 nm (16744 M-1cm-1) n(π*, 312 nm (12080 M-1cm-1) MLCT; EPR parameters g‖=2.164, g┴=2.078, A‖=136.74x10-4 cm-1, A┴=95.16x10-4 cm-1; elemental analysis calculated for [CuC27H29N4O4]NO3•CH3OH (%). 53.28 C, 5.27 H, 11.10 N; found: 53.14 C, 5.05 H, 10.86 N; molecular weight 631.14 g/mol.
Computational methods
Geometry optimization and atomic charge estimation
Computational calculations were conducted in Gaussian 09 [52] using density functional theory (DFT) calculations. 2nd generation of Minnesota M06 functional [53] and Los Alamos LanL2DZ [54] basis set were used to compute all compounds. X-ray diffraction preliminary results for [CasVII-YG] (compound 8) were considered, planar arrangements were observed, those were used as a starting point geometry. Remaining compounds were constructed manually in GaussView6. To simulate solvent (H2O) effects, a SMD Model [55] was achieved. No imaginary frequencies were detected confirming that the optimized geometry correspond to a local minimum on the potential energy surface. NBO analysis was used to estimate the required atomic charges for molecular docking experiments. Molar volume was obtained as single point calculations from optimized geometries.
Molecular docking
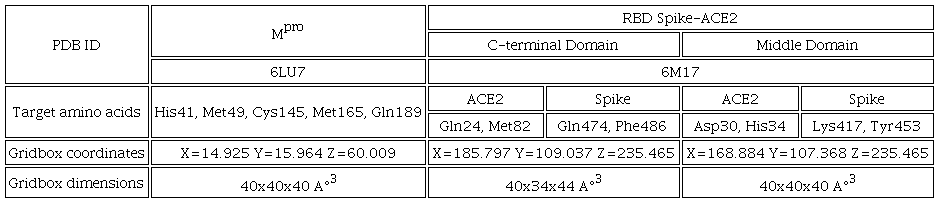
The crystallographic structures of main protease (Mpro) and RBDSpike-ACE2 complex were obtained from Protein Data Bank with the ID: 6LU7 [11], 6M17 [7] respectively. These proteins were used as controls because their inhibition capacity was previously reported as well as the interactions it presents. [19, 22] Prior to docking studies Mpro, and complex RBD Spike-ACE2 and Casiopeinas® were prepared as follows. In the proteins, extra units, water molecules, cofactors and inhibitors were removed then, polar hydrogens and Gasteiger charges were added using MGL Tools 1.5.6 software [55]. The modified structure was saved as pdbqt file. The optimized structures of copper complexes, remdesivir and bocepevir were employed as ligands, and the estimated atomic charges (QM: M06-LanL2DZ) were added manually in the Autodock4 pdbqt files. Constraints of copper atom as VdW radii, solvation volume, VdW well dept were included in the software parameters to handle the metal ion in the simulations. The docking studies were carried out using AutoDock 4.2 software [56]. A genetic algorithm study inside of the complex protein-Casiopeina® centered as seen in the table 2, 150 individuals in population with 2.5x103 evaluations to result in 10 docked poses. For the most stable conformation, energy stabilization (ΔGU) and inhibition constant (Ki) were reported. Finally, the docked conformation was analyzed with Discovery Studio 2021 [57] graphic interface.
Results and discussion
Synthesis of ternary copper(II) compounds
The preparation of the coordination compounds is a modification to the synthetic methodology of the Casiopeinas® [51, 58]. The change consists of deprotonate the dipeptide in a separate way, and then add this solution to mixture reaction, those was made considering the pH dependence of the dipeptides, since they can present in union modes to metallic center and the charge of the complexes [59], these ensuring coordination sphere that resembles to Casiopeinas®. The compounds are soluble in water, except 13 and 14, all compounds are soluble in MeOH, MeCN and DMSO. Coordination compounds decompose above 120 °C. The yields obtained were in the range of 85-93%. The powders obtained present colors in the scale from blue to green, being mostly blue when the primary ligand is bipyridine derivatives, while in the compounds that present 1,10-phenanthroline derivatives they are mostly green.
Physicochemical characterization
The synthesized compounds 1-14 (Table 1) present in the elementary analysis a minimum formula that agrees with the proposed formulas adjusted with solvent reaction molecules such as a methanol and water. Comparative analysis of the selected vibration frequencies in the FT-IR spectrum shows the presence of the characteristic functional groups. In the spectrum (Fig. S1 and Table S1) there are changes in the vibration frequencies attributed to the coordination of the central metal to the ligands used in this work, shifts in the vibration frequencies attributed to the carbonyl and amine groups are appreciable, so that in the IR spectra it is considered that coordination bonding occurs by the above mention functional groups, being noticeable in the signals belonging to the C=N (1610-1654 cm-1), O-H (3419-3473 cm-1), C-Har (723-734 and 810-848 cm-1), N-H and NH2 (3222-3279 cm-1) C=O (1675-1700 cm-1), C-Haliph (2923-3072 cm-1) and NO3- groups (1384 cm-1) [60]. In the FAB (+) spectra, the m/z signal corresponds to the cation complex. The conductivity measurements (Table S2) indicate 1:1 electrolytic ratio (H2O 132.63-158.45 Scm2mol-1 range), (MeOH 80.00-115.00 Scm2mol-1 range) type electrolyte [61], observed in various coordination compounds similar. The techniques of FAB (+) (Fig. S2 and table 2) and conductivity are consistent with the proposed molecular formulas. Magnetic susceptibility values are in the range of 1.80-2.16 B.M (Table S2), values that characteristic for an electronic configuration d9 (CuII) with an unpaired electron [62]; UV-Vis spectra have various absorbance maxima that are determined by the presence of various types of bond that make up the chemical structures of the synthesized compounds, such as the electronic transitions π(π* (201-257nm) presented mainly by aromatic groups, n(π* transitions (272-287nm) presented in groups with pairs of non-bonding electrons such as C=O and NH, metal to ligand charge transfer (MLCT) absorption band in the range of 294-219nm (d(π*), as well a maximum absorption in the visible region (620-637nm) assigned for the b2g(b1g transition in square planar geometry around the metal ion in the coordination compounds [63], this geometry would imply that the coordination bonding to the metal center is performed both by the nitrogen atoms of the diimines, as well as by the terminal amino and carbonyl groups of the peptide bond. Fig. 1. Show the UV-Vis spectrum for compound 8. The coordination compounds are stable in 1x10-3 M aqueous solution. Stability tests (Fig. S3) for compounds 3, 4, 9 and 10 were performed at 0.0, 0.5, 1.0, 2.0, 4.0, 6.0, 12.0, 24.0, 48.0 and 72.0 hours. These show that no change around the metal center involving a change in the coordination were observed. The decrease of absorbance values may be due to slight precipitation of the compounds, and then a dilution effect is seen in the electronic spectra.
![UV-Vis spectrum of 8. Solvent: water. Acquired UV region [0.1-1.0] mM; Vis region (top) acquired [1.0-3.0] mM.](../1870-249X-jmcs-68-01-29-gf2.jpg)
Fig. 1
UV-Vis spectrum of 8. Solvent: water. Acquired UV region [0.1-1.0] mM; Vis region (top) acquired [1.0-3.0] mM.
The EPR spectra obtained in methanol at 77K at 9.6GHz (X-Band), present the characteristic pattern of copper(II) compounds, in which an axial type spectrum is observed where g‖>g┴, with a hyperfine coupling in the g‖ region that has a multiplicity (M=2nI+1) of four due to nuclear spin (I=3/2) 63Cu or 65Cu; in the g┴ region [60-62], it can be seen in Fig. 2, belonging to CasX-GY (1) and CasVII-GY (7) that the signals presents a multiplicity of 7 due to superhyperfine coupling with 14N(I=1) [63]. Fig. 2 shows the comparative EPR spectrum for compounds 1 and 7, as well as shows the Aiso vs giso diagram [64-66] (Fig 3) in which Casiopeinas® synthesized in the region of the square planar geometry are located agree with the values of giso (Equation 1) and Aiso (Equation 2) obtained, a square planar geometry reported in the literature [67-68]. Table 3 shows Hamiltonian spin parameters for the compounds 1-14. The experimental and simulated EPR spectra of compound 1 are presented in Fig. S4.
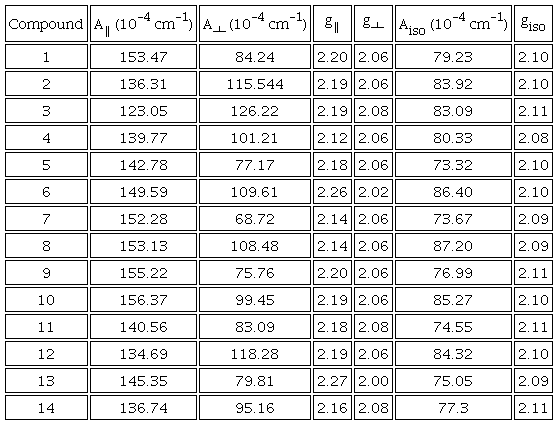
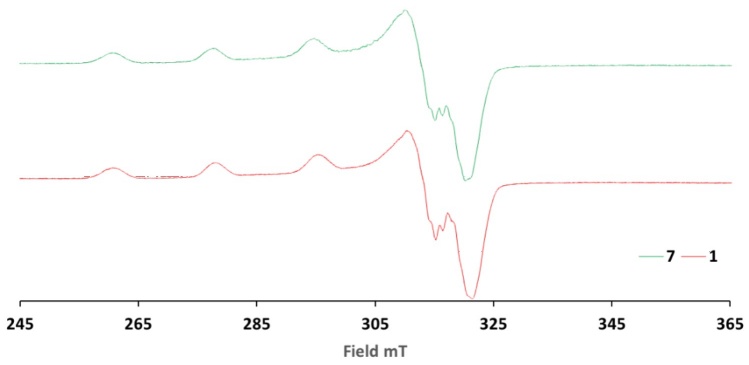
Fig. 2
Comparative EPR spectra for the compounds 1 and 7. EPR spectra were acquired at 77K from a MeOH solution of final concentration 1mM. EPR spectra acquired using X-Band.
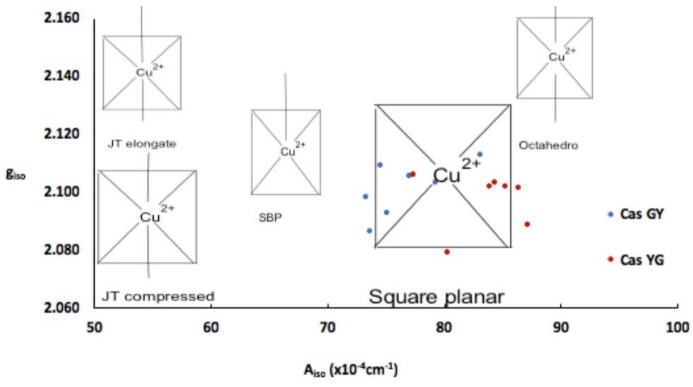
Fig. 3
Aiso vs giso diagram. Comparison of several copper(II) coordination compounds with different geometries, compounds studied in this work appear as square planar.
Considering the analysis performed in the spectroscopic techniques (IR, UV-Vis and EPR), it is proposed that the dipeptide is functionalized as a dipeptide ligand with the amino and carbonyl groups bonded to the copper(II) atom forming a 5-membered rings; the basic characteristics as a Pearson soft base of the primary amine and the peptide carbonyl, would imply a higher affinity to the metallic center that other donor groups present in the dipeptide as the secondary amine and carboxylate groups, which are Pearson hard bases.
Geometry optimization
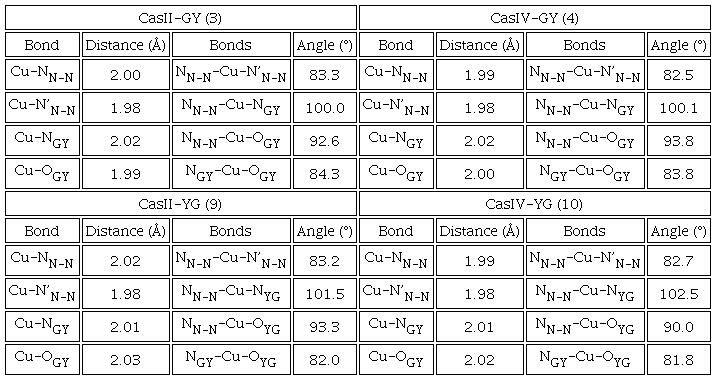
Prior to molecular docking studies, all Casiopeinas® were optimized using M06 functional and LanL2DZ pseudopotential. All copper(II) coordination compounds were optimized as [Cu(N-N)(L-L)]+ square planar complexes and were optimized employing DFT protocols. Finally, boceprevir and remdesivir were also optimized with the same computational methodology. Optimized geometries of CasII-GY, CasIV-GY and their analogues of YG are presented in Table 4. The formation of coordination bonds occurs through the nitrogen’s of the diimine, as well as the amino terminal of the dipeptide and the carbonyl group belonging to the peptide bond. This coordination mode agrees with the FT-IR and EPR characterization. DFT results suggests that the side chain present in tyrosine has an important effect on the chemical structure, as well as on the intramolecular interactions that the ternary copper(II) compounds can present. Since in the YG family the aromatic ring present in the dipeptide is in a position far from the copper atom, while in GY, the chemical structure is twisted so that there is an approach and a π-cation interaction (≈3.5 Å) between the phenyl and the central metal. Due to the structural similarity of both peptides, there is not separate cause for this behavior. Nevertheless, the steric hindrance, generated by the tertiary carbon of GY peptide, can be diminished by the formation of these intramolecular interaction. This trend is repeated in all studied Casiopeinas®. All Casiopeinas® (1-14) present a square planar arrangement around the metal center. In some cases, geometrical deviations were observed. Angles and distances estimated are in the range of the reported ones for some Casiopeinas® [72]. Generally, Casiopeinas® present an octahedral or square pyramidal geometries in solid state [73], completing with nitrate (NO3-) donors in the axial positions. However, in physiological conditions, the axial ligands can be substituted by aquo or other relevant biological molecules as nucleotides [74], glutathione or protein heteroatoms [75]. The optimizations presented here emulated the solution behavior of the coordination compounds and they are in according to the EPR experiments and the aqueous behavior of these class of compounds [32]. Energies, atomic coordinates and the remaining optimized geometries can be found in Table S3 and Scheme S1 of Supporting Information File.
Molecular docking
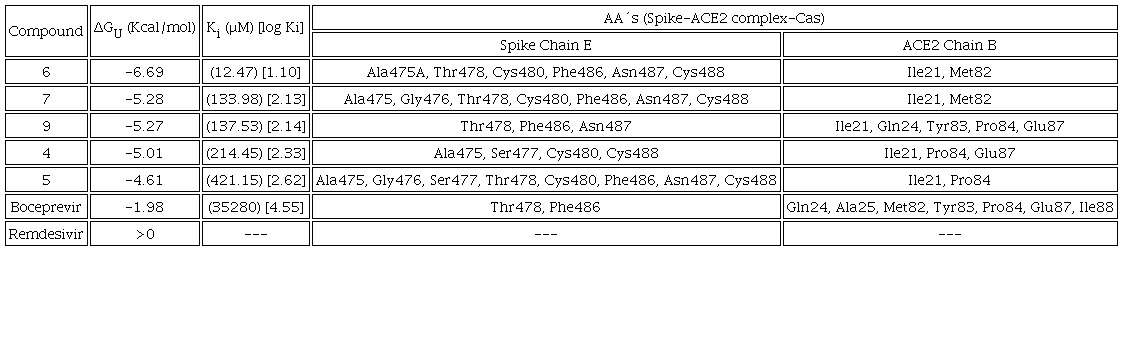
All compounds were docked to the Mpro enzyme as well as the RBDSpike-ACE2 complex in the RBD C-terminal and middle region. The values of binding energy (ΔGU), inhibition (Ki) for the 5 best rated compounds were analyzed and boceprevir and remdesivir were presented for comparative purposes. The major inhibitor systems are those that have a more negative ΔGU value, which is indicative of a stable interaction, a lower Ki value corresponds to the minimum amount to inhibit the protein efficiently. The objective is to focus on the Casiopeinas® that presents a greatest potential to be used as metallodrugs, particularly as anti-SARS-CoV-2 agents. In addition, the protein residues that interact with the coordination compounds are presented and the interactions obtained from the best molecular docking simulation were analyzed.
Mpro:Cas adduct
Table 5 presents the predicted ΔGU to evaluate the affinity to Mpro and the Ki for compounds 14, 10, 5, 3, 8, which exert the protein, as well as the interacting amino acids. Results with Mpro for compounds 3 and 9 as well as Casiopeinas® controls with amino acids (CasII-Tyr, CasIV-Tyr, CasII-gly and CasIV-gly) and without amino acids (CasIII-ia) were previous published [41]. Fig. 4 shows the comparative graph for all compounds analyzed. ΔGU values for the better pose are in the range of (-6.62 to -9.57) Kcal/mol. The calculated values of Ki present a range of 0.10-14.00 μM. For a wide range, log Ki is a better descriptor of the inhibition of Mpro. In general, Casiopeinas® can produce stable complexes with Mpro, that is expected from the structural diversity of the compounds, i. e., Casiopeinas® primary and secondary ligands are capable to form stabilizing interactions with the key residues of Mpro catalytic site. 14 and 10 have better ΔGU and Ki values. 5, 3, 8 have values in the same magnitude order and slightly less stable compared to boceprevir [76]. The remain copper compounds have higher stabilization and inhibition values compared to remdesivir. Considering that stable adducts are predicted from docking simulations, the catalytic site of the protein is blocked with a high specificity. As observed in Fig. 2, the Ki values are less than 10 μM, even presenting values in nanomolar order. Remained Mpro:Cas ΔGU, Ki and interactions can be found in Scheme S2 of supporting information file.
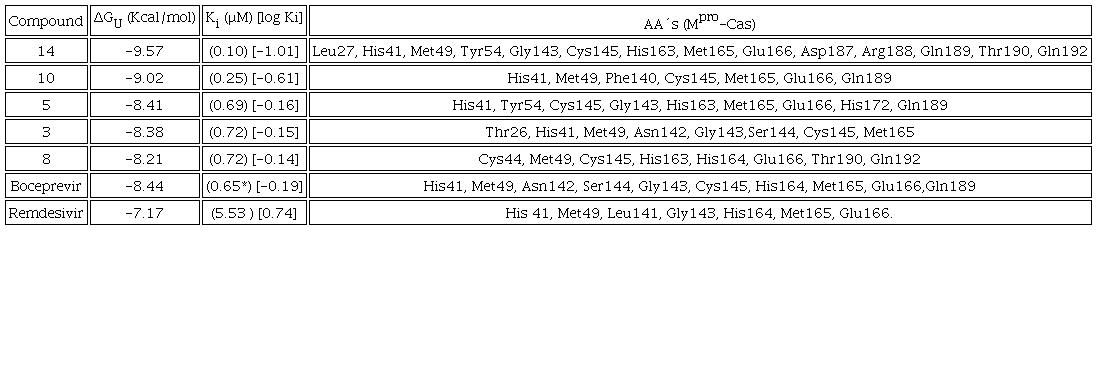
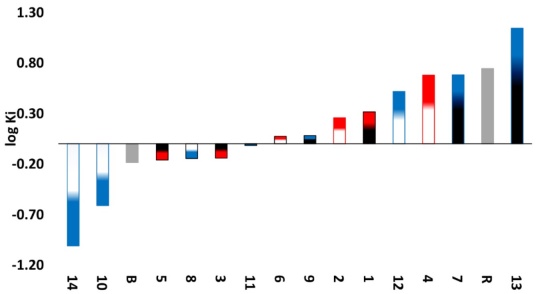
Fig. 4
Graph of predicted values of Mpro inhibition constants for all compounds. Red: Compounds derived from 2,2´-bipyridine. Blue: Compounds derived from 1,10-phenanthroline. White: Compounds derived from YG. Black: Compounds derived from GY. Grey. Controls. B: boceprevir. R: remdesivir.
Several interactions can stabilize the Casiopeina-Mpro adducts. The complex predicted between Mpro and 14, the best rated compound, is shown in Fig. 5. In the same figure, the pocket close-up and 2D diagrams can be observed to facilitate the localization of the stabilizing interactions.
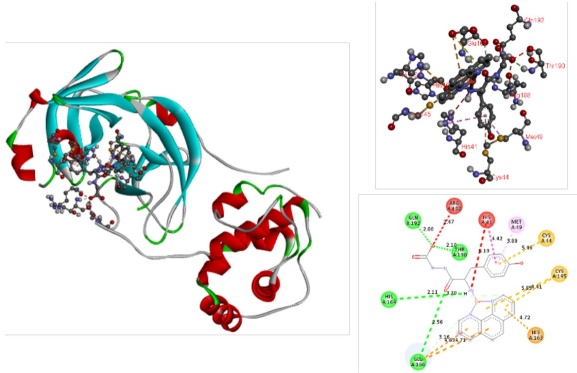
Fig. 5
Docked simulations of 14-Mpro complex. In the left side can be founded the complete complex. In the top of right side is showed a 3D diagram with the principal stabilizing interactions. 2D diagram is in the bottom of the right side.
According to DFT optimizations compounds 1-14 have square planar geometries, with copper atom in the center of the square, N-N ligands and L-L ligands are in the vertex. This arrangement allows the interaction with the amino acids responsible of the catalytic activity of the main protease, the catalytic dyad (His41, Cys145) and union triad (Met49, His163 and Gln189) [10]. Compound 14 interact with catalytic site and other residues belonging to the polypeptide chain of the protein. π-cation interactions are presented by the aromatic N-N ligand with His41, also π-π stacking between this residue and the tyrosine fragment of L-L ligand can be observed. Several π-alkyl interactions between the methyl substituents of 1,10-phenanthroline and Leu27, His41, His163. Hydrogen bonds between the amino and carboxylate groups of the dipeptide with Glu166, Gln189, Thr190, Gln192 were founded. The π-sulfur interaction of the diimine ligand with Met49, also occurs. Finally, π-sulfur interaction and hydrogen bond between the N-N ligand and Cys145 were observed. Similar interactions have been reported between Mpro and several species such as organic [77] and inorganic compounds [78], particularly with N3 peptide [79], as well as with rhenium and selenium coordination compounds [80] respectively. Compounds 10 and 5 presented in Fig. 6 offer value information regarding to the interactions provided by both the diimine ligand and the dipeptide moiety. Similar interactions occur due to the chemical structure of these 14 Casiopeinas® that favor the interaction with Mpro, and its potential inhibition. Inhibition of this protein may be satisfactory to prevent viral replication and transcription [81].
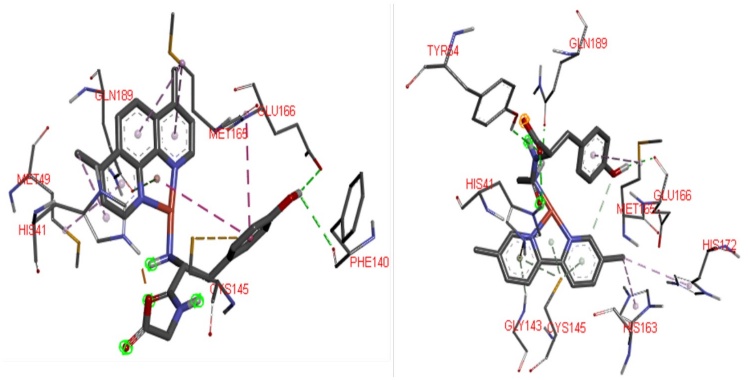
Fig.6
Possible 3D interaction diagrams of 10:Mpro (Left) and 5:Mpro (right) adducts, To recognize principal interactions, most the protein was removed. Only the main interactions predicted were presented.
Spike-ACE2 complex
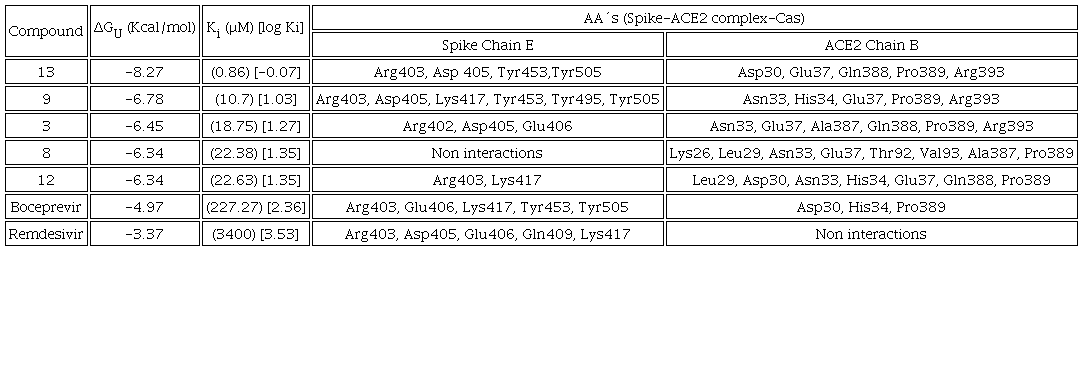
The study of the molecular docking of the Spike-ACE2 complex was carried out to determine if the Casiopeinas® can block the formation of protein complex. All compounds can interact with the C-terminal and central regions. Tables 6 (RBD C-terminal) and 7 (RBD-bridge) present the values of predicted ΔGU, Ki, as well as the amino acids of the B chain belonging to the Spike protein and the residues of the E chain of the human ACE2 protein. It is worth mentioning that in several cases they bind to the amino acids that participate in the assemble of the complex. That provokes that the virus enters to the human cells. The amino acids involved in assemble are Gln24 and Met82 located in the E chain of the ACE2, Gln474 and Phe486 of the Spike protein, belong to the C-terminal region. In the central region the targets are Asp30 and Hys34 from the ACE2, Lys417 and Tyr453 for Spike [7]. The details of the interactions are broken down separately for each region of the RBD Spike-ACE2 complex. The test compounds present significantly better values of ΔGU and Ki (Tables 6 and 7) than those shown by the controls, they are comparable with those evidenced by other potential inhibitors of RBDSpike-ACE2 complex [82]; there is currently no drug that has this protein complex as a molecular target [23]. Remained Spike-ACE2-Cas ΔGU, Ki and interactions can be found in Scheme S3 (C-terminal region) and Scheme S4 (central region) of Supporting Information File.
a) C-terminal region
The inhibition of this zone of the RBD of the Spike-ACE2 complex is centered in the amino acids Gln24 and Met82 of the ACE2, Gln474 and Phe486 of Spike protein [7]. The predicted values of ΔGU and Ki presented in table 6, which can be seen its visual representation in Fig. 7. Remdesivir control cannot inhibit the complex formation. On the other hand, Boceprevir control has a Ki value of 35.28 mM (log Ki=4.55), which is very high compared to that observed by compound 6, the difference is 3 orders of magnitude (Ki=12.47 μM, log Ki=1.10) value, 6 has the highest affinity to the Spike-ACE2 complex in the RBD at C-terminal domain. Most compounds have log Ki values in the range of 2.00 to 4.00.
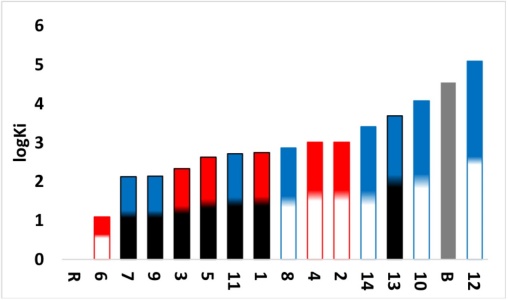
Fig. 7
Graph of predicted values of RBD-C terminal Spike-ACE2 complex inhibition constants for all compounds. Red: Compounds derived from 2,2´-bipyridine. Blue: Compounds derived from 1,10-phenanthroline. White: Compounds derived from YG. Black: Compounds derived from GY. Grey: Controls. B: Boceprevir. R: Remdesivir.
The potential high affinity of compound 6 to the Spike-ACE2 complex is due to highest number of potential interactions. The contacts presented this compound are Ile21 and Met82 (involved in the assemble of Spike-ACE2 complex), both in the ACE2, they are interactions of hydrogen bonds, salt bridge interaction between the carboxylate anion and the protonated amine of Ile21 (see Fig. 8). Regarding the E chain of the Spike, it should be noted that the amino acids that are part of the binding are with Gln474 and Phe486, the last amino acid interacts with Met82 of ACE2 to form the Spike-ACE2 complex. Compound 6 interacts with Phe486 (Spike) and Met82 (ACE2) by blocking the binding site to RBD; it interacts through a hydrogen bond, Phe486 forms the interaction by π-π stacking with one of the aromatic rings belonging of N-N ligand. Other interactions determined with the Spike E chain by molecular docking are π-sulfur (Cys480 and Cys488), π-alone pair (Asn487), π-donor hydrogen bond (Gly476 and Thr478) and π-alkyl (Ala475), between an aromatic ring of the 2,2´-bipyridine derivative with the residues described. 7 and 9 interacts in the same way that 6. The principal interactions of these compounds to be observed in Fig. 9. They allow the interaction with RBD-Spike-ACE2 C-terminal domain, as well as its potential inhibition.
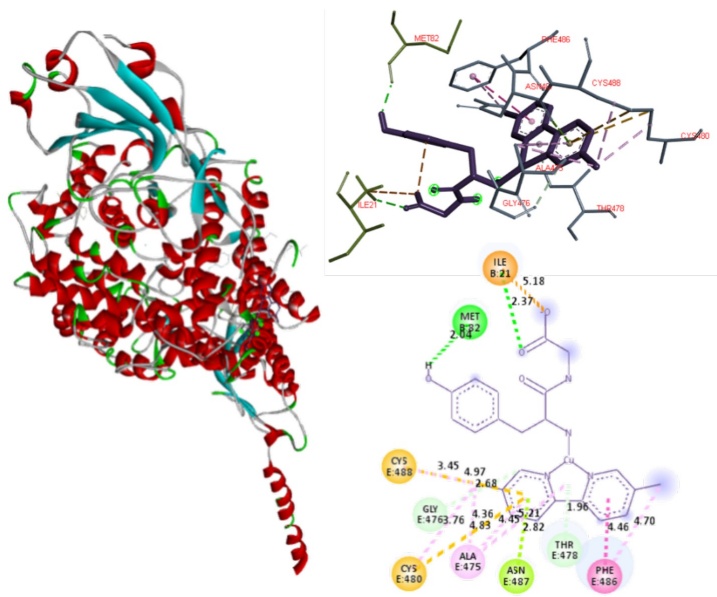
Fig. 8
Docked simulations of 6:Spike-ACE2 complex, C-terminal region. In the left side can be founded the complete complex. Red: Chain B of ACE2. Blue: Chain E of Spike. In the top right is showed a 3D diagram with the principal interactions between 6 and Spike-ACE2 complex. 2D diagrams is in the bottom of the right side.
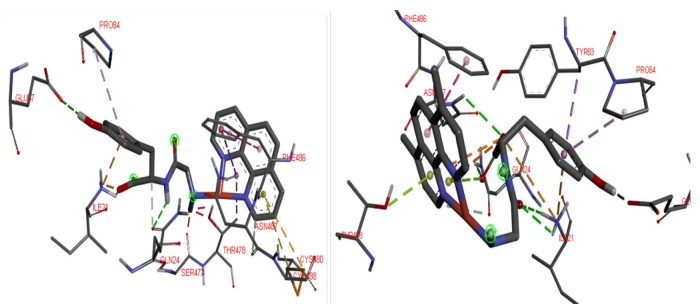
Fig. 9
Possible 3D interactions of 7-Complex Spike-ACE2 (left) and 9-Complex Spike-ACE2 (right) adducts. To recognize principal interactions, most the protein was removed. Only the main interactions were presented.
b) Central region
In this region, molecular docking is focused on the amino acids Asp30 and Hys34 of the ACE2 and the Lys417 and Tyr453 residues of the Spike protein, which are the amino acids that comprise the RBD Spike-ACE2 complex [7]. As denoted in table 7, Casiopeinas® present interaction with the residues of the E chain of the Spike protein and the B chain of the ACE2 with values of ΔGU -(4.76 to 8.27 Kcal/mol) and Ki (0.86 to 326.37 μM) better than those presented by the controls up to 4 orders of magnitude (Fig. 10), it should be mentioned that boceprevir presents interaction with the 4 amino acids that comprise the RBD Spike-ACE2, but the affinity is low compared to 13 of the proposed ternary copper(II) compounds. Compound 13 has the highest affinity to the Spike-ACE2 complex in the RBD at central region.
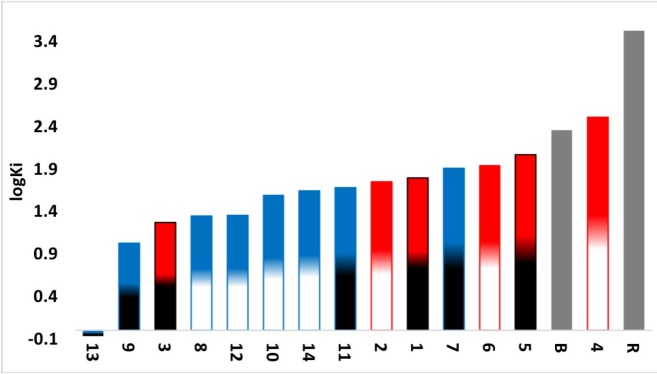
Fig. 10
Graph of predicted values of RBD-middle region Spike-ACE2 complex inhibition constants for all compounds. Red: Compounds derived from 2,2´-bipyridine. Blue: Compounds derived from 1,10-phenanthroline. White: Compounds derived from YG. Black: Compounds derived from GY. Grey: Controls. B: boceprevir. R: remdesivir.
The interactions that stabilized RBD SPIKE-ACE2 adducts involve the amino acids Asp30 and His34 interacts with Lys417 and Tyr453 respectively, the effect of blocking these contacts is important to inhibit the interaction of SARS-CoV-2 with the human host cells that express the ACE2 protein. As can be seen in table 7, compounds 3, 8, 9, 12 and 13 exhibits at least one of the four interactions mentioned, so inherently these compounds can inhibit the formation of the Spike-ACE2 complex in the bridge domain. Particularly, compound 13 has interactions with the amino acids Asp30 of the E chain of ACE2 and Tyr453 of the Spike protein [8], the inhibition action is performed through the formation of hydrogen bonds between carbonyl group of Asp30 and primary amine group of L-L. Other interaction occurs between hydroxyl group of Tyr453 and secondary amine on dipeptide. Other interesting interactions observed are due to the steric effects that can be allows the inhibition of RBD Spike-ACE2 in the bridge domain. These interactions are hydrogen bonds between Asp405 (Spike) and hydroxyl group of Tyrosine. Amide-π interactions (Arg393-N-N ligand and Arg403-Tyr of L-L ligand), π-alkyl interactions between the methyl groups with the amino acids Pro389 and Tyr505 in the ACE2 (Fig. 11) also were founded. Compounds 9 and 3 presented in Fig. 12 have similar interactions than 13, again all of these favors the interaction with RBD-Spike-ACE2 bridge domain and its potential inhibition.
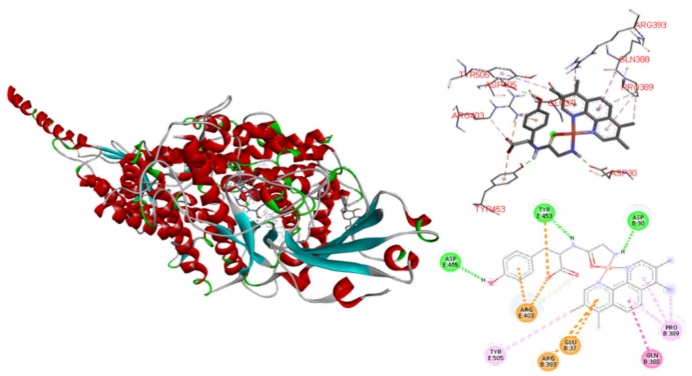
Fig. 11
Docked simulations of 13:Spike-ACE2 complex middle region. In the left side can be founded the complete complex, Red: Chain B of ACE2. Blue: Chain E of Spike. In the top right is showed a 3D diagram with the principal interaction between 6 and Spike-ACE2 complex. 2D diagrams is in the bottom of the right side.
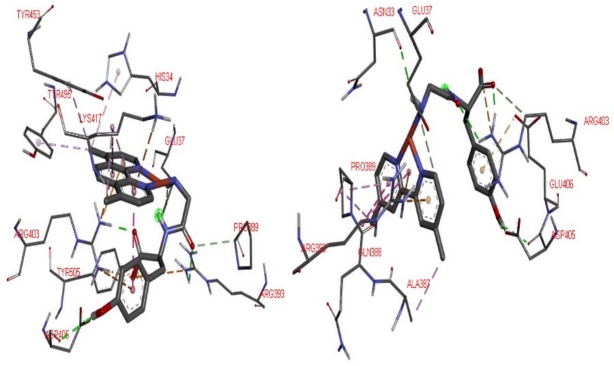
Fig. 12
3D interactions of 9-Spike-ACE2 complex (left) and 3-Spike-ACE2 complex (right) adducts. To recognize principal interactions, most the protein was removed. Only the main interactions were presented.
Conclusions
The 14 coordination compounds with diimine ligands and dipeptides Casiopeinas® synthesized were composed as monocationic coordination sphere according to the analysis obtained by the FAB(+), as well as with the values obtained in the conductivity. Ternary copper(II) compounds are paramagnetic with an unpaired electron. EPR studies suggest a planar square geometry and DFT calculations are in according with this geometry proposal and with the structural parameters informed for other Casiopeinas® and their solution behavior. The values obtained from ΔGU and Ki position the Casiopeinas® as potential inhibitory agents of SARS-CoV-2 transcendental proteins, since in the study with the Mpro, similar and even better values are presented with respect to the controls used in this work, in terms of the 2 regions analyzed in the RBD Spike-ACE2, the potential copper metallodrugs present significantly better values compared to those obtained for boceprevir and remdesivir. The potential inhibitory effect of Casiopeinas® may be due to the various components (copper[II], diamine and dipeptide) that form the ternary coordination compounds, wich interact with the amino acid residues present in SARS-COV2 proteins, through hydrogen bonding, various π-interactions, as well as by electrostatic attraction. Casiopeinas® may stabilize the formation of adducts with amino acids belonging to the catalytic site of Mpro or with those involved in the formation of RBD Spike-ACE2 in the domains analyzed.
Acknowledgements
This work was supported by UNAM-PAPIIT IN218013; Project Grant UNAM-PAPIIT 230020; UNAM-FQ 5000-9047; UNAM-DGTIC (projects LANCAD-UNAM-DGTIC-358 and UNAM-DGTIC-COVID-005). LGTC thanks to CONACYT-CVU 689679 for the PhD scholarship. To Dra. Laura Padierna and M. Karyna Pérez from the Laboratorios de Especialidades Inmunológicas S.A. de C.V. for the donating material. To M in C. Adrian Espinoza Guillén for the technical support in the LQIM. †MLVC died on April 28, 2009, her expertise in the chemical synthesis of peptides inspired this work, as well as she was a disciple of Professor Joaquín Tamaríz who is honored with this special issue. Dedicated to all people who died from this emerging disease.
References
Zhou, P.; Yang, X. L.; Wang, X. G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H. R.; Zhu, Y.; Li, B.; Huang, C. L.; Chen, H. D.; Chen, J.; Luo, Y.; Guo, H.; Jiang, R. D.; Liu, M. Q.; Chen, Y.; Shen, R.; Wang, X.; Zheng, X. S.; Zhao, K.; Chen, Q. J.; Deng, F.; Liu, L. L.; Yan, B.; Zhan, F. X.; Wang, Y. Y.; Xiao, G. F.; Shi, Z. L. Nature 2020, 579, 270-273. DOI: https://doi.org/10.1038/s41586-020-2012-7
Wu, F.; Zhao, S.; Yu, B.; Wang, W.; Song, Z. G.; Hu, Y.; Tao, Z. W.; Tlan, J. H.; Pel, Y. Y.; Yuan, M. L.; Zhang, Y. L.; Dal, F. H.; Liu, Y.; Yang, Q. M.; Zheng, J. J.; Xu, L.; Holmes, E. C.; Zhang, Y. Z. Nature 2020, 579, 265-269. DOI: https://doi.org/10.1038/s41586-020-2008-3
http://covid19.who.int/, accessed in March 2023.
Guarner, J. Am. J. Clin. Pathol, 2020, 153, 420-421. DOI: https://doi.org/10.1093/AJCP/AQAA029
V'koski, P.; Kratzel, A.; Steiner, S.; Stalder, H.; Thiel, V. Nat. Rev, Microbiol. 2020, 19, 155-170. DOI: https://doi.org/10.1038/s41579-020-00468-6
Wu, A.; Peng, Y.; Huang, B.; Ding, X.; Wang, X.; Niu, P.; Meng, J.; Zhu, Z.; Zhang, Z.; Wang, J.; Sheng, J.; Quan, L.; Xia, Z.; Tan, W.; Cheng, G.; Jiang, T. Cell Host Microbe. 2020, 27, 325-328. DOI: https://doi.org/10.1016/j.chom.2020.02.001
Yan, R.; Zhang, Y.; Li, Y.; Xia, L.; Guo, Y.; Zhou, Q. Science. 2020, 367, 1444-1449. DOI: https://doi.org/10.1126/science.abb2762
Lan, J.; Ge, J.; Yu, J.; Shan, S.; Zhou, H.; Fan, S.; Zhang, Q.; Shi, X.; Wang, Q.; Zhang, L.; Wang, X. Nature 2020, 581, 215-220. DOI: https://doi.org/10.1038/s41586-020-2180-5
Zhou, X.; Zhang, Y.; Zhong, F.; Lin, C.; McCormick, P. J.; Jiang, F.; Luo, J.; Zhou, H.; Wang, Q.; Fu, Y.; Duan, J.; Zhang, J. Chin. Sci. Bull. 2021, 66, 661-663. DOI: https://doi.org/10.106/j.scib.2020.10.018
Finkel, Y.; Mizrahi, O.; Nachshon, A.; Weingarten-Gabbay, S.; Morgenstern, D.; Yahalom- Ronen, Y.; Tamir, H.; Achdout, H.; Stein, D.; Israeli, O.; Beth-Din, A.; Melamed, S.; Weiss, S.; Israely, T.; Paran, N.; Schwartz, M.; Ginossar, S. Nature. 2020, 589,125-130. DOI: https://doi.org/10.1038/s41586-020-2739-1
Jin, Z.; Du, X.; Xu, Y.; Deng, Y.; Liu, M.; Zhao, Y.; Zhang, B.; Li, X.; Zhang, L.; Peng, C.; Duan, Y.; Yu, J.; Wang, L.; Yang, K.; Llu, F.; Jlang, R.; Yang, X.; You, T.; Liu, X.; Yang, X.; Bai, F.; Liu, H.; Liu, X.; W, Guddat.; Xu, W.; Xiao, G.; Qin, C.; Shi, Z.; Jlang, H.; Rao, Z.; Yang, H. Nature. 2020, 582, 289-293. https://doi.org/10.1038/s41586-020-2233-y
Xu, T.; Ooi, A.; Lee, H. C.; Wilmouth, R.; Liu, D. X.; Lescar, J. Acta Crystallogr., Sect. F: Struct. Biol. Cryst. Commun. 2005, F61, 964-966. DOI: https://doi.org/10.1107/s1744309105033257
Parvez, S. A.; Karim, A.; Hasan, M.; Jaman, J.; Karim, Z.; Tahsin, T.; Hasan, N.; Hozen, M. Int. J. Biol. Macromol. 2020, 163, 1787-1797. DOI: https://doi.org/10.1016/j.ijbiomac.2020.09.098
Naveja, J. J.; Madariaga-Mazón, A.; Flores-Murrieta, F.; Granados-Montiel, J.; Madariaga- Ceceña, M.; Duarte-Alaniz, V.; Maldonado-Rodríguez, M.; García-Morales, J.; Senosiain- Peláez, J. P.; Martínez-Mayorga, K. Drug Discovery Today. 2021, 26, 229-239. DOI: https://doi.org/10.1016/j.drudis.2020.10.018
Domínguez-Villa, F. X.; Durán-Iturbide, N. A.; Ávila-Zarraga, J. G. Bioorg. Chem. 2021, 106, 104497. DOI: https://doi.org/10.1016/j.bioorg.2020.104497
Wu, C.; Liu, Y.; Yang, Y.; Zhang, P.; Zhong, W.; Wang, Y.; Wang, Q.; Xu, Y.; Li, M.; Li, Z.; Zheng, M.; Chen, L.; Li, H. Acta Pharm. Sin. B. 2020, 10, 766-788. DOI: https://doi.org/10.1016/j.apsb.2020.02.008
Krumm, Z. A.; Lloyd, G. M.; Francis, C. P.; Nasif, L. H.; Mitchell, D. A.; Golde, T. E.; Giasson, B. I.; Xia, Y. Virol. J. 2021, 18. DOI: https://doi.org/10.1186/s12985-021-01526-y
Guedes, I. A., Costa, L. S. C., Dos Santos, K. N., Karl, A. L., Rocha, G. K., Texeira, I. M., Galheigo, M. M., Medeiros, V., Krempser, E., Custódio, F. L., Barbosa, H. J. C., Nicolas, M. F., Dardenne, L. E. Sci. Rep. 2021, 11, 5543. DOI: https://doi.org/10.1038/s41598-021-84700-0
Ahamad, S.; Kanipakam, H.; Birla, S.; Ali, M. S.; Gupta, D. Eur. J. of Pharm. 2021, 890, 173664. DOI: https://doi.org/10.1016/j.ejphar.2020.173664
Fu, L.; Ye, F.; Feng, Y.; Yu, F.; Wang, Q.; Wu, Y.; Zhao, C.; Shun, H.; Huang, B.; Niu, P.; Song, H.; Shi, Y.; Li, X.; Tan, W.; Qi, J.; Gao, G.F. Nat. Commun. 2020, 11, 4417, 1-8. DOI: https://doi.org/10.1038/s41467-020-18233-x
Ali, A.; Sepay, N.; Afzal, M.; Alarifi, A.; Shadid, M.; Ahmad, M. Bioorg. Chem. 2021, 110, 104772. DOI: https://doi.org/10.1016/j.bioorg.2021.104772
Eweas, A. F.; Alhossary, A. A.; Abdel-Moneim, A. S. Front. Microbiol. 2021, 11, 1-15. DOI: https://doi.org/10.3389/fmicb.2020.592908
Gil-Moles, M.; Türck, S.; Basu, U.; Pettenuzzo, A.; Bhattacharya, S.; Rajan, A.; Ma, X.; Büssing, R.; Wölker, J.; Burmeister, H.; Hoffmeister, H.; Schneeberg, P.; Prause, A.; Lippmann, P.; Kusi-Nimarko, J.; Hassell-Hart, S.; McGown, A.; Guest, D.; Lin, Y.; Notaro, A.; Vinck, R.; Karges, J.; Cariou, K.; Peng, K.; Qin, X.; Wang, X.; Skiba, J.; Szczupak, L.; Kowalski, K.; Schatzchneider, U.; Hemmert, C.; Gornitzka, H.; Milaeva, E. R.; Nazarov, A. A.; Gasser, G.; Spencer, J.; Ronconi, L.; Kortz, U.; Cinatl, J.; Bojkova, D.; Ott, I. Chem. Eur. J. 2021, 27, 17928-17940. DOI: https://doi.org/10.1002/chem.202103258
Pal, M.; Musib, D.; Roy, M. New. J. Chem. 2020, 10, 1039. DOI: https://doi.org/10.1039/D0NJ04578K
Geromichalou, E. G.; Trafalis, D. T.; Dalezis, P.; Malis, G.; Psomas, G.; Geromichalos, G. D. J. Inorg. Biochem. 2022, 231, 111805. DOI: https://doi.org/10.1016/j.jinorgbio.2022.111805
Karges, J.; Cohen, S. M. Chem. Biochem. 2021, 22, 1-9. DOI: https://doi.org/10.1002/cbic.202100186
Owen, D. R.; Allerton, C. M. N.; Anderson, A. S.; Aschenbrenner, L.; Avery, M.; Berrit, S.; Boras, B.; Cardin, R. D.; Carlo, A.; Coffman, K. J.; Dantonio, A.; Di, L.; Eng, H.; Ferre, R. A.; Gajiwala, K. S.; Gibson, S. A.; Greasley, S. E.; Hurst, B. L.; Kadar, E. P.; Kalgutkar, A. S.; Lee, J. C.; Lee, J.; Liu, W.; Mason, S. W.; Noell, S.; Novak, J. J.; Obach, R. S.; Ogilvie, K.; Patel, N. C.; Pettersson, M.; Rai, D. K.; Reese, M. R.; Sammons, M. F.; Sathish, J. G.; Singh, R. S. P.; Steppan, C. M.; Stewart, A. E.; Tuttle, J. B.; Updyke, L.; Verhoest, P. R.; Wei, L.; Yang, Q.; Zhu, Y. Science. 2021, 374, 1586-1593. DOI: https://doi.org/10.1126/science.abl4784
Anthony, E. J.; Bolitho, E. M.; Bridgewater, H. E.; Carter, O. W. L.; Donnelly, J. M.; Imberty, C.; Lant, E. C.; Lermyte, F.; Needham, R. J.; Palau, M.; Sadler, P. J.; Shi, H.; Wang, F. X.; Zhang, W. Y.; Zhang, Z. Chem. Sci. 2020, 11, 12888-12917. DOI: https://doi.org/10.1039/d0sc04082g
Cirri, D.; Pratesi, A.; Marzo, T.; Messori, L. Expert. Opin. Drug. Discovery. 2020. 16, 39-46. DOI: https://doi.org/10.1080/17460441.2020.1819236
Karges, J. Chem. Bio. Chem. 2020, 21, 1-4. DOI: https://doi.org/10.1002/cbic.202000397
Ioannou, K.; Vlasiou, M. Biometals. 2022. 35, 639-652. DOI: https://doi.org/10.1007/s10534-022-00386-5
Correia, I.; Borovic, S.; Cavaco, I.; Matos, C. P.; Roy, S.; Santos, H. M.; Fernández, L.; Capelo, J. L.; Ruiz-Azuara, L.; Costa Pessoa, J. J. Inorg. Biochem. 2017, 175, 284-297. DOI: https://doi.org/10.1016/j.jinorgbio.2017.07.025
Rufino-González, Y.; Ponce-Macotela, M.; García-Ramos, J. C.; Martínez-Gordillo, M. N.; Galindo-Murillo, R.; González-Maciel, A.; Reynoso-Robles, R.; Tovar-Tovar, A.; Flores- Álamo, M.; Toledano-Magaña, Y.; Ruiz-Azuara, L. J. Inorg. Biochem. 2019, 195, 83-90. DOI: https://doi.org/10.1016/j.jinorgbio.2019.03.012
Larios-Cervantes, A. A.; Chávez-Cortéz, E. G.; Martínez-Hernández, M.; Talavera- Contreras, L. G.; Espinoza-Guillen, A.; Carrillo-Ávila, B. A.; Ruiz-Azuara, L.; Álvarez-Pérez, M. A.; Martínez-Aguilar, V. M. Braz. J. Microbiol. 2022, 53, 179-184. DOI: https://doi.org/10.1007/s42770-021-00648-3
Ruiz-Azuara, L.; Bravo-Gómez, M. E. Curr. Med. Chem. 2010, 17, 3606-3615. DOI: https://doi.org/10.2174/092986710793213751
Ramírez-Palma, L. G.; García-Jacas, C. R.; García-Ramos, J. C.; Almada-Monter, R.; Galindo-Murillo, R.; Cortés, F. J. Molec. Struct. 2020, 1204, 127480. DOI: https://doi.org/10.1016/j.molstru.2019.127480
Trejo-Solís, C.; Jiménez-Farfan, D.; Rodríguez-Enriquez, S.; Fernández-Valverde, F.; Cruz- Salgado, A.; Ruiz-Azuara, L.; Sotelo, J. BMC. Cancer. 2012, 12, 156.
Gutiérrez, A. G.; Vázquez-Aguirre, A.; García-Ramos, J. C.; Flores-Alamo, M.; Hernández- Lemus, E.; Ruiz-Azuara, L.; Mejía, C. J. Inorg. Biochem. 2013, 126, 17-25. DOI: https://doi.org/10.1016/j.jinorgbio.2013.05.001
Figueroa-DePaz, Y.; Resendiz-Acevedo, K.; Dávila-Manzanilla, S. G.; García-Ramos, J. C.; Ortiz-Frade, L.; Serment-Guerrero, J.; Ruiz-Azuara, L. J. Inorg. Biochem. 2022, 231, 111772. DOI: https://doi.org/10.1016/j.jinorgbio.2022.111772
Kachadourian, R.; Brechbuhl, H. M.; Ruiz-Azuara, L.; Gracia-Mora, I.; Day, B. J. Toxicol.2010, 268, 176-183. DOI: https://doi.org/10.1016/j.tox.2009.12.010
García-Ramos, J. C.; Galindo-Murillo, R.; Tovar-Tovar, A.; Alonso-Sáenz, A. L.; Gómez- Vidales, V.; Flores-Álamo, M.; Ortiz-Frade, L.; Cortés-Guzmán, F.; Moreno-Esparza, R.; Campero, A.; Ruiz-Azuara, L. Chem. Eur. J. 2014, 20, 13730-13741. DOI: https://doi.org/10.1002/chem.201402775
Reyna, M.; Talavera Contreras, L. G.; Figueroa de Paz, Y.; Ruiz-Azuara, L.; Hernández Ayala, L. F. New. J. Chem., 2022, 46, 12500-12511. DOI: https://doi.org/10.1039/d2nj01480g
Erxleben, A. Coord. Chem. Rev. 2018, 360, 92-121. DOI: https://doi.org/10,1016/j.ccr.2018.01.008
Grenács, Á.; Kaluha, A.; Kállay, C.; Jószai, V.; Sanna, D.; Sóvago, I. J. Inorg. Biochem. 2013, 128, 17-25. DOI: https://dx.doi.org/10.1016/j.jinorgbio.2013.07.008
Oliveira-Brett, A. M.; Diculescu, V. C.; Enache, T. A.; Fernandes, I. P. G.; Chiorcea-Paquim, A. M.; Oliveira, S. C. B. Curr. Opin. Electrochem. 2019, 14, 173-179. DOI: https://doi.org/10.1016/j.coelec.2019.03.008
Arcos-López, T.; Qayyum, M.; Rivilas-Acevedo, L.; Miotto, M. C.; Grande-Aztatzi, R.; Fernández, C. O.; Hedman, B.; Hodgson, K. O.; Vela, A.; Solomon, E. I.; Quintanar, L. Inorg. Chem. 2016, 55, 2909-2922. DOI: https://doi.org/10.1021/acs.inorgchem.5b02794
Brasún, J.; Cebrat, M.; Sochacka, A.; Gladysz, O.; Swiatek-Kozloska, J. Dalton. Trans. 2008, 4978-4980. DOI: https://doi.org/10.1039/b807799a
Kotynia, A.; Czyznikowska, Z.; Cebrat, M.; Jaremko, L.; Gladysz, O.; Jaremko, M.; Marciniak, A.; Brasún, J. Inorg. Chim. Acta. 2013, 396, 40-48. DOI: https://dx.doi.org/10.10167j.ica.2012.09.035
Atrián-Blanco, E.; González, P.; Santoro, A.; Alies, B.; Faller, P.; Hureau, C. Coord. Chem. Rev. 2018, 371, 38-55. DOI: https://doi.org/10.1016/j.ccr.2018.04.007
McGivern, T. J. P.; Afsharpour, S.; Marmion, C. J. Inorg. Chim. Acta. 2018, 472, 12-39. DOI: https://doi.org/10.1016/j.ica.2017.08.043
Ruiz-Azuara, L. 07/628,628 Re 35,458, Feb 18 (1997) U.S. Patent 1992, in 1992. Ruiz-Azuara, L. 07/628,628: 5,576,326 U.S. Patent 1996 in 1996.
Frisch, M. J.; Trucks, G. W.; Schlegel, H. B.; Scuseria, G. E.; Robb, M. A.; Cheeseman, J. R.; Scalmani, G.; Barone, V.; Mennucci, B.; Petersson, G. A.; Nakatsuji, H.; Caricato, M.; Li, X.; Hratchian, H. P.; Izmaylov, A. F.; Bloino, J.; Zheng, G.; Sonnenberg, J. L.; Hada, M.; Ehara, M.; Toyota, K.; Fukuda, R.; Hasegawa, J.; Ishida, M.; Nakajima, T.; Honda, Y.; Kitao, O.; Nakai, H.; Vreven, T.; Montgomery, J. A.; Peralta, J. E.; Ogliaro, F.; Bearpark, M. J.; Kudin, K. N.; Staroverov, V. N.; Kobayashi, R.; Normand, J.; Raghavachari, K.; Rendell, A. P.; Burant, J. C.; Iyengar, S. S.; Tomasi, J.; Cossi, M.; Rega, N.; Millam, N. J.; Klene, M.; Knox, J. E.; Cross, J. B.; Bakken, V.; Adamo, C.; Jaramillo, J.; Gomperts, R.; Stratmann, R. E.; Yazyev, O.; Austin, A. J.; Cammi, R.; Pomelli, C.; Ochterski, J. W.; Martin, R. L.; Morokuma, K.; Zakrzewski, V. G.; Voth, G. A.; Salvador, P.; Dannenberg, J. J.; Dapprich, S.; Daniels, A. D.; Farkas, O.; Foresman, J. B.; Ortiz, J. V.; Cioslowski, J.; Fox, D. J. Gaussian 09, Gaussian Inc, Wallingford, 2009.
Zhao, Y.; Truhlar, D.G. Theor. Chem. Acc., 2008, 120, 215-241. DOI: https://doi.org/10.1007/s00214-007-0310-x
Dolg, M.; Wedig, U.; Stoll, H. J. Chem. Phys., 1989, 90, 1730-1734. DOI: https://doi.org/10.1063/1.456066
Marenich, A. V.; Cramer, C. J.; Truhlar, D. G. J. Phys. Chem. B, 2009, 113, 6378-6396. DOI: https://doi.org/10.1021/jp810292n
Morris, G.; Huey, R.; Lindstrom, W.; Sanner, M.; Belew, R.; Goodsell, D.; Olson, A. J. Comput. Chem., 2009, 30, 2785-2791. DOI: https://doi.org/10.1002/jcc.21256
BIOVIA, Dassault Systèmes, Discovery Studio 2021, V.21 1.0, San Diego: Dassault Systèmes, 2020.
Título de Marca Reg. 407543 SECOFI (1992), (2002), (2012).
Sóvágo, I.; Várnagy, K.; Lihi, N.; Gregnács, Á. Coord. Chem. Rev. 2016, 327-328, 43-54. DOI: http://dx.doi.org/10.1016/j.ccr.2016.04.015
Pretsch, E.; Bühlmann, P.; Badertscher, M. Structure determination of organic compounds. 4th Ed. Springer-Verlag, Berlin, 2009.
Pethybridge, A. D.; Spiers, D. J. J. Chem. Soc., Chem Commun. 1974, 423-424. DOI: https://doi.org/10.1039/C39740000423
Drago, R. S. Physical methods for chemists. 2nd Ed. W.B. Saunders Company, Philadelphia, 1977.
Hagen, W. H. Dalton Trans. 2006, 4415-4436. DOI: https://doi.org/10.1039/B608163K
Fielding, A. J.; Fox, S.; Millshauser, G. L.; Chattopadhyay, M.; Kroneck, P. M. H.; Fritz, G.; Eaton, G. R.; Eaton, S. S. J. Magn. Reson. 2006, 179, 92-104. DOI: https://doi.org/10.1016/j.jmr.2005.11.011
Garriba, E.; Micera, G. J. Chem. Ed. 2006, 83, 1229-1232. DOI: https://doi.org/10.1021/ed083p1229
Kivelson, D.; Neiman, R. J. Chem. Phys. 1961, 35, 149. DOI: http://dx.doi.org/10.1063/1.1731880
Sawada, T.; Fukumaru, K.; Sakurai, H. Chem. Pharm. Bull. 1996, 44, 1009-1016.
Kwik, W. L.; Ang, K. P. J. Inorg. Nucl. Chem. 1980, 42, 303-313.
Yokoi, H.; Addison, W. Inorg. Chem. 1977, 16, 1341-1349.
Peisach, J.; Blumberg, W. E. Arch. Biochem. Biophys. 1974, 165, 691-708. DOI: https://doi.org/10.1016/0003-9861(74)90298-7
Gembus, A.; Corzilius, B.; Eichel, R. A.; Dinse, K. P.; Immel, S.; Stumm, D.; Flauaus, M. & Plenio, H. J. Phys. Chem. B. 2006, 110, 15012-15020. DOI: https://doi.org/10.1021/jp062158x
Bravo-Gómez, M. E.; Dávila-Manzanilla, S.; Flood-Garibay, J.; Muciño-Hernández, M. A.; Mendoza, A.; García-Ramos, J. C.; Moreno-Esparza, R.; Ruiz-Azuara, L. J. Mex. Chem. Soc., 2012, 56, 85-92. DOI: https://doi.org/10.29356/jmcs.v56i1.280
Tovar-Tovar, A.; García-Ramos, J. C.; Flores-Álamo, M.; Ruiz-Azuara, L. Acta Cryst., 2011, E67, m1796-m1797. DOI: https://doi.org/10.1107/S1600536811047398
García-Ramos, J. C.; Tovar-Tovar, A.; Hernández-Lima, J.; Cortés-Guzman, F.; Moreno- Esparza, R.; Ruiz-Azuara, L. Polyhedron, 2011, 30, 2697-2703. DOI: https://doi.org/10.1016/j.poly.2011.07.022
Ugone, V.; Pisanu, F.; Sana, D.; Garriba, E. J. Inorg. Biochem., 2021, 224, 111566. DOI: https://doi.org/10.1016/j.jinorgbio.2021.111566
Ma, C.; Sacco, M. D.; Hurst, B.; Townsend, J. A.; Hu, Y.; Szeto, T.; Zhang, X.; Tarbet, B.; Marty, M. T.; Chen, Y.; Wang, J. Cell. Res. 2020, 30, 678-692. DOI: https://doi.org/10.1038/s41422-020-0356-z
Zhang, L.; Lin, D.; Sun, X.; Curth, U.; Drosten, C.; Sauerhering, L.; Becker, S.; Rox, K.; Hilgenfeld, R. Science. 2020, 368, 409-412. DOI: https://doi.org/10.1126/science.abb3405
Buitrón-González, I.; Aguilera-Durán, G.; Romo-Mancillas, A. Results. Chem. 2021, 3, 100094. DOI: https://doi.org/10.1016/j.rechem.2020.100094
Yuan, S.; Wang, R.; Fuk-Woo Chan, J.; Zhang, A. J.; Cheng, T.; Ka-Heng Chik, K.; Ye, Z. W.; Wang, S.; Chak-Yiu Lee, A.; Jin, L.; Li, H.; Jin, D. Y.; Yuen, K. Y.; Sun, H. Nat. Microbiol. 2020, 5, 1439-1448. DOI: https://doi.org/10.1038/s41564-020-00802-x
Griffin, J. W. D. J. Struct. Biol. 2020, 211, 107575. DOI: https://doi.org/10.1016/j.jsb.2020.07575
Weglarz-Tomczak, E.; Tomczak, J. M.; Talma, M.; Burda-Grabowska, M.; Giurg, M.; Brul, S. Sci. Rep. 2021, 11, 3640. DOI: https://doi.org/10.1038/s41598-021-83229-6
Alkhatip, A. A. A. M. M.; Georgakis, M.; Montero Valenzuela, L. R.; Hamza, M.; Farag, E.; Hodgkinson, J.; Hosny, H.; Kamal, A. M.; Wagih, M.; Naguib, A.; Yassin, H.; Algameel, H.; Elayashy, M.; Abdelhaq, M.; Younis, M. I.; Mohamed, H.; Abdulshafi, M.; Elramely, M. A. Int.J.Mol. Sci. 2021, 22, 2977. DOI: https://doi.org/10.3390/ijms22062977
Gil-Moles, M.; Basu, U.; Büssing, R.; Hoffmeister, H.; Türck, S.; Varchmin, A.; Ott, I. Chem. Eur. J. 2020, 26, 15140-15144. DOI: https://doi.org/10.1002/chem.202004112
Author notes
*Corresponding author: lenar701@gmail.com, lenar701@quimica.unam.mx; Phone: +52(55)56223529.
