Artículos
Registration frameworks for aligning brain images
Esquemas de registro para alinear imágenes del cerebro
Registration frameworks for aligning brain images
Archivos Venezolanos de Farmacología y Terapéutica, vol. 39, núm. 4, pp. 521-525, 2020
Sociedad Venezolana de Farmacología Clínica y Terapéutica

Esta obra está bajo una Licencia Creative Commons Atribución-SinDerivar 4.0 Internacional.
Recepción: 28 Mayo 2020
Aprobación: 15 Junio 2020
Publicación: 07 Julio 2020
Resumen: En este artículo, se presentan los resultados obtenidos por el proceso de registro de los volúmenes de imágenes cerebrales obtenidos por resonancia magnética (MRI) y resonancia magnética funcional (fMRI) utilizando dos marcos computacionales diferentes. El objetivo es comparar el rendimiento de cada marco, enfocando esta comparación en la medición de error obtenida por el registro de volúmenes cerebrales. La comparación involucra el registro intramodal e intramodalidad (MRI–MRI y fMRI–fMRI). El Statistical Parametric Mapping (SPM) y el Insight Segmentation and Registration Toolkit (ITK) se eligen como marcos de registro. La metodología propuesta considera la generación de conjuntos de datos, planificación de pruebas, diseño de casos de prueba, ejecución de pruebas y evaluación. Finalmente, estos resultados son analizados. La correspondencia entre los volúmenes registrados y el volumen objetivo usando el marco ITK es mayor que la obtenida con el marco SPM.
Palabras clave: Imágenes del cerebro, MRI, fMRI, registro de imágenes, SPM, ITK.
Abstract: In this paper, the results obtained by the registration process of brain image volumes obtained by magnetic resonance imaging (MRI) and functional magnetic resonance imaging (fMRI) using two different computational frameworks are presented. The objective is to compare the performance of each framework, focusing this comparison in the error measurement obtained by brain volumes registration. The comparison involves the intra patient and intra modality (MRI–MRI and fMRI–fMRI) registration. Statistical Parametric Mapping (SPM) and Insight Segmentation and Registration Toolkit (ITK) are chosen as registration frameworks. The proposed methodology considers the data sets generation, test planning, designing test cases, tests execution and evaluating. Finally, these results are analysed. The correspondence between the volumes registered and the target volume using the ITK framework is greater than that obtained with the SPM framework.
Keywords: Brain images, MRI, fMRI, image registration, SPM, ITK.
Introduction
Image registration is an image processing technique very important in medical imaging. This technique can be defined as the process of finding the spatial transformation that allows to align points between two images acquired at different times, from different points of view, or from the same or different energy sources1,2. Figure 1 shows a diagram of the image registration process which consists of the following basic components: a target image, a source image, a transformation, a metric, an interpolator and an optimizer.
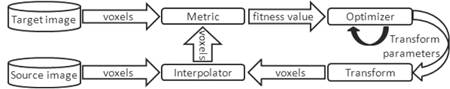
Figure 1
A registration process
Medical image registration can be considered intra patient, inter modality and intra modality3. Routinely, for a patient, different volumes are acquired several times using a single modality or with different medical imaging modalities. The dynamical acquisitions are also frequently performed, generating sequences or time series of images. Inter modality registration allows the combination of complementary information from different image modalities, while intra modality registration allows accurate comparisons between images of the same modality.
Head motion is an important issue involved in any fMRI study and retrospective image registration is, by far, the most commonly used approach on estimating head motion4. Motion correction tools generally assume that head motion can be described using a rigid body transformation, which means that the position of the head can change but that the shape of the head cannot change4.
There are several software packages available for MRI/fMRI image registration; these include SPM, AFNI, AIR, BrainVoyager and FSL4. Another option for image registration is the ITK library which has been widely used in the last decade5, even though its application on fMRI has been scarce. With regard to studies about comparison of MRI/fMRI image registration tools; there are a relatively small number of papers that tackle this subject. The three most common papers in the literature are the studies of 1) Morgan et al.6,7, who compared different versions of the available software packages: SPM, AFNI, AIR, BrainVoyager and FSL using a computer generated phantom, from results they determined that there are few differences among the packages, 2) Ardekany et al.8, who applied the motion correction tools: SPM(99), AIR(3.08), AFNI(98), and TRU to simulated fMRI data and concluded that SPM provides the most accurate motion detection, and 3) Oakes et al.9, who made a more extensive comparison using the packages AFNI, AIR, BrainVoyager, FSL and SPM with real and simulated data, the authors found that AFNI and SPM yield the most accurate motion estimation parameters. At present, we are unaware of any study that has attempted to compare registration tools based on the ITK and SPM.
The paper is organized as follows. First, the registration frameworks are described. Then, the complete process for generating the data sets is detailed. After, the test planning is proposed. Then, experiments carried out to compare the frameworks are presented, and the results obtained are discussed.
Materials and Methods
Data Source
In this stage, the data sets used in the proposed experiments to evaluate the performance of registration frameworks are generated. The anatomical data, which correspond to data describing the structure, shape and relationships of the different parts of an organ, are generated from volumes acquired using brain MRI and the functional data from time series acquired using fMRI scanning. The fMRI data describe the organic activity, considering changes in metabolism, blood flow, and absorption. Two synthetic databases of MRI and fMRI three–dimensional images are constructed from the brain volumes of the database Auditory (available in http://www.fil.ion.ucl.ac.uk/spm/data/auditory/). This brain database is comprised of a MRI anatomical volume and 96 fMRI volumes. The spatial resolution of the MRI volume is 256×256×54 with a voxel size of 1×1×3 mm. Each fMRI volume contains 64 slices. The fMRI slice thickness is of 3 mm. In all fMRI volumes, the slices have an isotropic resolution of 64×64 pixels with a size of 3 mm. The voxel value for all volumes is represented by 12 bits. The Auditory anatomical volume (hereinafter referred to as “MRI target volume”) is used to construct a sequence of 134 brain MRI volumes. In the same context, the average functional Auditory volume (hereinafter referred to as “fMRI target volume”) is used to construct a fMRI synthetic database of 134 functional volumes. The designed synthetic data sets are constructed as follow:
1. Two first’s data sets of MRI and fMRI synthetic volumes, respectively, are constructed in order to simulate the translation component associated with the motion of a rigid body. Each data set consists of 63 volumes generated by applying 63 different translation motions to each target volume. The translation vector {x, y, z} takes values in mm of the set {0, 5, 7, 9}. The motion with components {0, 0, 0} is not considered.
. Two new sets, one for each database are added. The aim is to include into databases a set of 63 volumes which takes into account another component of the rigid motion as rotation. Such volumes are generated by rotating the target volumes certain degrees respect to the axes x, y and z. Values belonging to the set {0, 1, 2, 3} are considered to rotate except of the rotation vector {0, 0, 0}.
3. Finally, a set composed of 8 volumes generated from the combined translation and rotation of each target volume is included in each database. The motions are applied hierarchically. The translation volume is performed by translation computer program and rotation motions are applied to its output using rotation computer program. Values for the translation and rotation motions are chosen according to the results of experiments using the data sets defined above; hence the selection criteria of the translation and rotation values are explained in section 2.3.1.
In summary two databases of brain synthetic data, one of anatomical data and one of functional data, are constructed. These databases are referred to hereinafter as MRI_db for anatomical data and fMRI_db for functional data. Each database contains three data sets, the first is composed only by the translated volumes, the second considers only rotation, and the last set contains volumes with combined translation and rotation. These sets are denoted in Table 1. Therefore, MRI_db = ..MRI È ..MRI È .trMRI and fMRI_db = ..fMRI È ..fMRI È .trfMRI.
| Description | MRI Data set | fMRI Data set |
| Translated volumes | StMRI | StfMRI |
| Rotated volumes | SrMRI | SrfMRI |
| Volumes with combined translation and rotation | StrMRI | StrfMRI |
Registration frameworks
In this section a description of the characteristics for both selected registration frameworks is given.
Selected frameworks
1. SPM is software designed for brain imaging data sequences analysis. The SPM software is selected due to the fact that this computer application is widely accepted by the scientific community. However, SPM has a limited ability to reuse and integrate with other software packages. SPM requires the MatLab. computer program for execution, in this sense, financial investment for use is required10.
2. ITK is a cross–platform object–oriented library that provides developers with a comprehensive set of tools for medical images analysis. Several advantages such as ease of reuse and integration are considered for its selection. ITK is developed under the open source standards and is freely available, therefore, financial investment is not required and, moreover, the computer programs coded using this library do not depend on any other software11. In this sense, for this research a computer program is coded in order to implement the brain image registration process based on this library.
SPM and ITK registration process
The mathematical and computational techniques available for SPM software and ITK library associated with each component of a registration process are presented in Table 1. In order to develop the registration frameworks to compare, for each basic component of the registration process (see Figure 1), the techniques listed in Table 2 have been analyzed and those common, that fulfill the requirements for alignment of brain images, are selected.
· Because the head motion can be described using a rigid body transformation, it is very reasonable to assume that an affine registration scheme (based on affine transformation) can be feasible and sufficient to head motion correction4,12. In this sense and considering that affine transformation is the single transformation codified for SPM, this transformation model based on linear operators is selected. The translation along each axis, rotation around each axis, scaling along each axis (stretching), and shearing along each axis are the linear operators whose combination generates an affine transformation.
· The optimizer corresponds to the Powell13 technique since it is the single optimization technique considered by SPM. ITK proposes seven additional optimizers14.
· Concerning the interpolation technique, β-spline third degree has been selected. This selection is based on suggestions found in the SPM documentation15, in which it is stated that the nearest neighbor interpolation is fast to calculate and implement, however, it is not recommended in the image registration process since it incorporates noise to volumes. Furthermore, trilinear interpolation is reported suitable for positron emission tomography (PET), or realignment and reslicing brain images.
· Regarding the metric, it is necessary to use the mutual information technique because it generates the most acceptable results when coupled to the Powell optimization technique for medical image registration. In ITK there are two implementations of this metric14,16. The Viola and Wells implementation16 is used in both frameworks.
Test planning
This section starts from test planning, then designing test cases, preparing for execution and evaluating. The test planning is prepared considering intra patient and intra modality registration of brain images.
Test Cases
Designed test cases consist of intra modality registration of anatomical data and functional data, hence two cases are considered, MRI–MRI registration and fMRI–fMRI registration.
· Case 1: MRI–MRI Registration. In this case, 134 tests are proposed which correspond to MRI–MRI registration of the MRI target volume respect to each volume in the MRI db. The volumes belonging to datasets ..MRI, ..MRI and .trMRI are considered source volumes.
· Case 2: fMRI–fMRI Registration. These tests consider datasets StfMRI, SrfMRI and StrfMRI. The volumes into these datasets are used individually as source volume in the registration process with the fMRI target volume. A total of 134 tests is proposed in this case.
Table 2. Mathematical and computational techniques for SPM and ITK image registration processes.
| Framework | Transformation | Interpolator | Optimizer | Metric |
| SPM | · Affine transform | · Nearest neighbour · Trilinear · β-spline 2nd to 7th | · Powell | · Mutual information · Normalized mutual information · Entropy correlation coefficient · Normalized cross correlation |
| ITK | · Identity transform · Translation transform · Scale transform · Scale logarithmic transform · Rigid 2–D transform · Rigid 3–D transform · Affine transform · Β-Spline deformable transform · Set of transforms | · Nearest neighbour · Linear · β-spline 2nd to 7th · Windowed sinc | · Nelder–Meade downhill simplex · Gradient descent · Conjugate gradient · Quasi Newton decent · L-BFGS · Evolutionary strategy · Powell · Exhaustive search | · Mean squares · Normalized correlation · Mean reciprocal squared difference · Mutual information · Cardinality match · Gradient difference · Kullback Liebler distance · Kappa statistics · Normalized mutual information · Mean squares histogram · Correlation coefficient histogram |
Test execution and evaluation
At this stage, the tests described in the previous section are executed using the registration module of SPM and the registration computer program developed using the ITK library. Both registration frameworks consider the components described in section 2.2.2.
The evaluation of the proposed cases is performed for each data set. For ..MRI and ..fMRI the transformation matrices obtained by executing the registration frameworks based on SPM and ITK are required in the evaluation. The translation values into the matrices are compared with respect to the translation real values. The absolute error between the theoretical value and the value obtained upon registration is used as error metric.
Meanwhile, for SrMRI and SrfMRI datasets the differences between the target volume and the volumes registered using SPM and the computer program encoded with ITK are used in the evaluation. This difference is quantified using the exclusive OR (XOR), which selects the voxels that are different in both volumes17.
The differences between the target volume and the volumes registered are also used as error metric for the datasets StrMRI and StrfMRI. These data sets contain the volumes with combined translation and rotation data. The translation and rotation values are set according to the following procedure:
1. In sets ..MRI and ..fMRI the volumes that generate, in their corresponding test, the smaller and higher absolute error are identified for each registration framework.
2. The volumes in SrMRI and SrfMRI, for which the error quantified using XOR operator is minimum and maximum, are also identified.
3. Once the volumes, for which the quantified error in the registration process is minimum and maximum, are identified, the translation and rotation values used in the construction of these volumes are recombined in order to construct the data sets StrMRI and StrfMRI according to the procedure described in section 2.1.
Results
In this section the results of applying the test cases over the data sets are presented. Figure 2 shows the boxplots with the absolute error by each axis obtained using both frameworks, for each test case, after registering the volumes in the translation data sets (StMRI and StfMRI). In addition, Figure 3 shows the statistical information concerning the not exclusive OR (XNOR) metric for the two frameworks after registering the rotated volumes (SrMRI and SrfMRI) and the volumes with combined translation and rotation (StrMRI and StrfMRI).
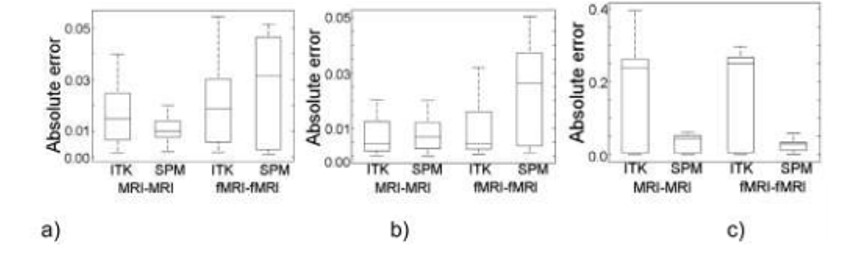
Figure 2.
Registration results of translated volumes. (a) x-axis translation. (b) y-axis translation. (c) z-axis translation.
For MRI–MRI registration and according to Figure 2, the framework that has the best performance is SPM because about 100% of the error values are below the ITK absolute error median. With respect to Figure 2, the ITK has the best error values for fMRI-fMRI registration except for .-axis.
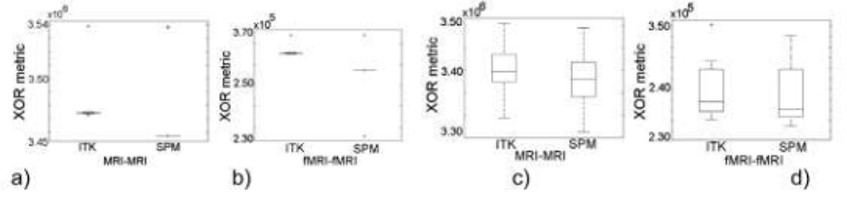
Figure 3.
Registration results. Rotated volumes: (a) MRI-MRI (b) fMRI-fMRI. Volumes with combined translation and rotation: (c) MRI-MRI (d) fMRI-fMRI.
From the analysis of the XNOR metric in Figure 3(a) and 3(b), 100% of the correspondences obtained with ITK framework are above the correspondences reached using the SPM registration framework. In Figures 3(c) and 3(d), the registration framework showing the best performance is also based on ITK library.
Conclusions
The correspondence between the volumes registered and the targets volumes (MRI and fMRI) using the ITK registration framework is greater than that obtained with the SPM framework when a combined translation and rotation motions are considered in both brain images modalities. On the other hand, the results of the test cases (MRI-MRI and fMRI-fMRI) applied to rotation data sets (SrMRI and SrfMRI) prove that ITK based framework generates volumes with greater correspondence (respect to MRI and fMRI references volumes) that the volumes obtained using SPM framework, when only rotation motion is considered. Finally, for the data set StMRI, which only considers translation motion, the SPM based framework shows better performance than the ITK registration framework in the MRI-MRI test case. However, from the analysis of the absolute error for the x- and y- axis, when the data set StfMRI (fMRI translated volumes) is regarded, the ITK framework reaches lower values; while the z-axis absolute error is higher using ITK registration. Since SPM framework has a higher number of registrations with low error comparing the absolute error, this framework is better if the brain is regarded as a rigid object subjected to translation motion only.
As a general conclusion, the framework based on ITK library applied to MRI-MRI and fMRI-fMRI intra modalities registration generates the better spatial transformation useful to align the brain when a combination of translation and rotation motions is used to simulate head motion.
References
1. Pluim, J., Fitzpatrick, J. Image registration. IEEE Trans. Med. Imaging 22(11), 1341-1343, 2003.
2. Zitová, B., Flusser, J. Image registration methods: a survey. Image Vision Comput. 21(11), 977–1000, 2003.
3. Hajnal, J., Hill, D., Hawkes, D. Medical image registration. CRC Press, Boca Raton, 2001.
4. Strother, S. Evaluating fMRI preprocessing pipelines. IEEE Eng Med Biol Mag 25(2), 27-41, 2006.
5. Avants, B., Tustison, N., Stauffer, M., Song, G.,Wu, B., Gee, J. The Insight ToolKit image registration framework. Front Neuroinform; 8(44), 2014.
6. Morgan, V., Pickens, D., Hartmann, S., Price, R. Comparison of functional MRI image realignment tools using a computer–generated phantom. MRM; 46: 510-514, 2001.
7. Morgan, V., Dawant, B., Li, Y., Pickens, D. Comparison of fMRI statistical software packages and trategies for analysis of images containing random and stimulus–correlated motion. Comput Med Imaging Graph; 31: 436–446, 2007.
8. Ardekani, B., Bachman, A., Helpern, J. A quantitative comparison of motion detection algorithms in fMRI. Magn Reson Imaging; 19(7): 959-963, 2001.
9. Oakes, T., Johnstone, T., Walsh, K.O., Greischar, L., Alexander, A., Fox, A., Davidson, R. Comparison of fMRI motion correction software tools. NeuroImage; 28(3), 529-543, 2005.
10. Friston, K., Ashburner, J., Kiebel, S., Nichols, T., Penny, W. Statistical Parametric Mapping: The Analysis of Functional Brain Images. Academic Press, 2007.
11. Johnson, H., McCormick, M., Ibañez, L., Insight Software Consortium. Insight Segmentation and Registration Toolkit (ITK). http://www.itk.org/, 3ed. 2009. Updated for ITK version 4.5.
12. Bankman, I.N. Handbook of Medical Image Processing and Analysis. Academic Press, 2009.
13. Press, W.H., Teukolsky, S.A., Vetterling, W.T., Flannery, B.P. Numerical Recipes in C. The Art of Scientific Computing. Cambridge University Press, New York, 1992.
14. Ibáñez, L., Schroeder, W., Ng, L., Cates, J. The ITK Software Guide. Kitware, 2003.
15. The FIL Methods Group and honorary members, SPM12 Manual. Functional Imaging Laboratory, Institute of Neurology, UCL, 2015, http://www.fil.ion.ucl.ac.uk/spm/doc/manual.pdf.
16. Viola, P. Wells, W. Alignment by maximization of mutual information. Int. J. Comput. Vision 24(2): 137-154, 1997.
17. Russ, J. Russ, J. C. Introduction to image processing and analysis. CRC Press, Boca Raton, 2008.
Notas de autor
a.bravo@unisimonbolivar.edu.co
