Molecular characteristics of vancomycin-susceptible Staphylococcus aureus could help to predict treatment failure due to reduced vancomycin susceptibility
Características moleculares de Staphylococcus aureus suscetível à vancomicina poderia ajudar a prever falhas no tratamento devido à reduzida suscetibilidade à vancomicina
Características moleculares de Staphylococcus aureus susceptible a la vancomicina podría ayudar a predecir fallas en el tratamiento debido a la reducida susceptibilidad a la vancomicina
Molecular characteristics of vancomycin-susceptible Staphylococcus aureus could help to predict treatment failure due to reduced vancomycin susceptibility
Revista de Epidemiologia e Controle de Infecção, vol. 8, no. 4, pp. 422-427, 2018
Universidade de Santa Cruz do Sul

This work is licensed under Creative Commons Attribution 4.0 International.
Received: 28 November 2017
Accepted: 19 September 2018
Resumo: Justificativa e Objetivos:Staphylococcus aureus resistente à meticilina (MRSA) é uma das causas mais frequentes de infecções relacionadas à assistência à saúde e comunitárias, e com seu avanço, a vancomicina tornou-se a principal opção terapêutica. Entretanto, o seu uso indiscriminado favoreceu o surgimento de MRSA com reduzida suscetibilidade à vancomicina, comumente associados com falhas no tratamento, bacteremia persistente, hospitalização prolongada e desfechos clínicos adversos. Este estudo avaliou a ocorrência de MRSA com reduzida suscetibilidade à vancomicina e determinou algumas características moleculares em comparação com MRSA suscetível à vancomicina (VS-MRSA). Métodos: Determinação do perfil de suscetibilidade aos antimicrobianos, a concentração inibitória mínima (CIM) e concentração bactericida mínima (CBM) para vancomicina, tolerância à vancomicina, tipagem do SCCmec e agr foram realizadas em um total de 177 MRSA. Posteriormente, foram triados para hVISA por BHIA-3V e BHIA-6V e confirmados com a Análise do Perfil Populacional - Área Abaixo da Curva (PAP-AUC). Resultados: Os fenótipos VT-MRSA e hVISA foram encontrados em 13,6% e 5,1% dos isolados clínicos de MRSA, respectivamente, e a presença de hVISA foi estatisticamente significativa entre os isolados de VT-MRSA (.<0,05). Em VT-MRSA, SCCmec tipo II foi significativamente mais frequente do que em não-VT-MRSA, assim como a presença do agr grupo II. Conclusões: Características moleculares encontradas em MRSA são importantes para a epidemiologia, bem como para demonstrar um padrão em isolados com reduzida suscetibilidade à vancomicina. Testes não-convencionais para detecção destas características podem ser realizados para evitar a identificação errada de VS-MRSA que, consequentemente, resulta em falhas no tratamento com vancomicina.
Palavras-chave: agr, hVISA, SCCmec, VISA, VT-MRSA.
Abstract: Background and Objectives: Methicillin-resistant Staphylococcus aureus (MRSA) is one of the most frequent causes of healthcare-associated and community-acquired infections and with its advancement, vancomycin became the main therapeutic option. However, its indiscriminate use favored the emergence of MRSA with reduced susceptibility to vancomycin, commonly associated with vancomycin treatment failure, persistent bacteremia, prolonged hospitalization and adverse clinical outcome. This study evaluated the occurrence of MRSA with reduced vancomycin susceptibility and determined some molecular characteristics in comparison with vancomycin-susceptible MRSA (VS-MRSA). Methods:Determination of antimicrobial susceptibility profile, the minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) for vancomycin, vancomycin-tolerance, SCCmec and agr typing were performed in a total of 177 MRSA. Thereafter, they were screened for hVISA by BHIA-3V and BHIA-6V and confirmed with population analysis profile - area under the curve method (PAP-AUC). Results: VT-MRSA and hVISA phenotypes were found in 13.6% and 5.1% of clinical isolates of MRSA, respectively, and the presence of hVISA was statistically significant among VT-MRSA isolates (.<0.05). In VT-MRSA, SCCmec type II was significantly more frequent than in non-VT-MRSA, as well as the presence of agr group II. Conclusion: Molecular characteristics found in MRSA are important for epidemiology, as well as demonstrate a pattern in reduced vancomycin susceptibility isolates. Non-conventional tests for detection of these characteristics might be performed to prevent misidentification of VS-MRSA that, consequently, results in vancomycin treatment failures.
Keywords: agr, hVISA, SCCmec, VISA, VT-MRSA.
Resumen: Justificación y objetivos: Staphylococcus aureus resistente a la meticilina (MRSA) es una de las causas más frecuentes de infecciones relacionadas con la asistencia sanitaria y comunitarias, y con su avance, a la vancomicina se ha convertido en la principal opción terapéutica. Sin embargo, su uso indiscriminado favoreció el surgimiento de MRSA con reducida susceptibilidad a la vancomicina, comúnmente asociados con fallas en el tratamiento, bacteriemia persistente, hospitalización prolongada y resultados clínicos adversos. Este estudio evaluó la ocurrencia de MRSA con reducida susceptibilidad a la vancomicina y determinó algunas características moleculares en comparación con MRSA susceptible a la vancomicina (VS-MRSA). Métodos: Determinación del perfil de susceptibilidad a los antimicrobianos, la concentración inhibitoria mínima (CIM) y la concentración bactericida mínima (CBM) para vancomicina, tolerancia a la vancomicina, tipificación del SCCmec y agr se realizaron en un total de 177 MRSA. Resultados: Los fenotipos VT-MRSA y hVISA se encontraron en el 13,6% y el 5,1% de los aislados clínicos de MRSA, respectivamente, y la presencia de hVISA fue estadísticamente significativa entre los aislados de VT-MRSA (.<0.05). En VT-MRSA, SCCmec tipo II fue significativamente más frecuente que en no-VT-MRSA, así como la presencia del agr grupo II. Conclusiones: Características moleculares encontradas en MRSA son importantes para la epidemiología, así como para demostrar un patrón en aislados con reducida susceptibilidad a la vancomicina. Pruebas no convencionales para la detección de estas características pueden realizarse para evitar la identificación errónea de VS-MRSA que, consecuentemente, resulta en fallas en el tratamiento con vancomicina.
Palabras clave: agr, hVISA, SCCmec, VISA, VT-MRSA.
INTRODUCTION
The pathogenesis of Staphylococcus aureus infections is complex and depends on the host characteristics, expression of virulence factors and ability to develop resistance to antimicrobials.1,2 Methicillin-resistant S. aureus (MRSA) is related to an advancement of healthcare- and community-acquired infections, being vancomycin the primary therapeutic option for the last fifty years, even with the availability of antimicrobials as linezolid, tigecycline, and daptomycin. Infections caused by vancomycin-susceptible MRSA (VS-MRSA) have been associated with vancomycin treatment failures and, also, an increase in mortality.3,4 Hence, its excessive and inappropriate use has led to the emergence of S. aureus with reduced vancomycin susceptibility, as vancomycin-intermediate S. aureus (VISA), heterogeneous VISA (hVISA) and afterwards, vancomycin-tolerant MRSA (VT-MRSA), last two not yet detected routinely.5-8
Infections caused by hVISA and VISA are commonly associated with prolonged hospitalization, persistent bacteremia, and adverse clinical outcome. Although the relevance of hVISA persists unclear, studies speculate that these isolates could be a precursor of VISA phenotype.9-11 MRSA strains also develop vancomycin tolerance, whose infections are even more difficult to treat, especially when they are invasive and present in immunocompromised patients.. Additionally, this reduced vancomycin susceptibility is associated with dysfunction in the accessory gene regulator (agr) locus, a quorum sensing system that controls the expression of genes encoding virulence factors.12,13
In this context, to understand the epidemiology and some molecular characteristics of MRSA with reduced vancomycin susceptibility is required to assist in the antimicrobial therapy and control the spread of these multiresistant microorganism. This study was designed to determine the prevalence of SCCmec and agr types among MRSA isolates, and to evaluate the association of these molecular characteristics with reduced susceptibility to vancomycin.
METHODS
Ethics
The cross-sectional observational study was approved by the institutional Ethics Committee, with permit number 1.212.043. The Central Laboratories from Hospital Mãe de Deus (HMD) and Hospital Nossa Senhora da Conceição (HNSC) provided de MRSA isolates for this study, stored at -20ºC in our sample collection. According to the institutional ethical and type of investigation assessment, informed consent was not required.
Clinical isolates of MRSA
A collection of 177 MRSA clinical isolates from distinct hospitalized patients were analyzed. They were isolated from respiratory tract 72 (40.7%), blood 40 (22.6%), skin and soft tissue 39 (22.0%), bone and connective tissue 10 (5.6%), medical devices 9 (5.1%), urine 5 (2.8%), and others sites 2 (1.1%). S. aureus identification was confirmed in the laboratory using conventional methods, as Gram staining, catalase activity, plasma coagulase production and growth on brain heart infusion agar (BHIA; Oxoid, Basingstoke, England) supplemented with 15% NaCl. Also, methicillin resistance was confirmed by conventional PCR for mecA gene and by the traditional Kirby-Bauer method using cefoxitin (DME, São Paulo, Brazil), according to Clinical and Laboratory Standard Institute (CLSI) guidelines.14,15
Antimicrobial susceptibility
Antimicrobial susceptibility tests were performed by disk diffusion method for clindamycin (2 μg), erythromycin (15 μg), levofloxacin (5 μg), rifampicin (5 μg), teicoplanin (30 μg), tetracycline (30 μg), and trimethoprim/sulfamethoxazole (1.25 μg/ 23.75 μg) (DME, São Paulo, Brazil), and vancomycin susceptibility was determined by microdilution method in Mueller-Hinton broth (MHB; Oxoid, Basingstoke, England), according to the CLSI guidelines.15S. aureus ATCC 25923 and S. aureus ATCC 29213 strain was used as a control, respectively.
Vancomycin tolerance was determined by the minimum bactericidal concentration (MBC)/minimum inhibitory concentration (MIC) ratio of ≥ 32 μg/mL, as described by Cázares-Domínguez et al.. S. aureus ATCC 29213 was used as a control.
Screening for hVISA
All MRSA were submitted to a screening test for hVISA in brain heart infusion agar (BHIA; Oxoid, Basingstoke, England) containing 3 μg/mL vancomycin (BHIA-3V)16 and 6 μg/mL vancomycin (BHIA-6V).15 Briefly, 10 μL of a bacterial suspension (1 x 108 CFU/mL) was spread onto BHIA-3V and incubated for 24h at 37ºC. Cultures that showed growth after 24h w confirmed by Population Analysis Profile - Area Under the Curve (PAP-AUC). hVISA Mu3 (ATCC 700698) was used as a control.
Population Analysis Profile – Area Under the Curve method
PAP/AUC was performed as previously described by Huang et al.11Briefly, a 0.5 McFarland standard suspension from an overnight culture in trypticase soy agar (TSA; Oxoid, Basingstoke, England) was prepared. Serial dilutions (10-1 a 10-5) were prepared in sterile saline, 10 µL from each dilution was spread onto BHI agar plates containing increasing concentrations of vancomycin (0 µg/mL, 1 µg/mL, 2 µg/mL, 3 µg/mL, 4 µg/mL, 5 µg/mL, 6 µg/mL and 8 µg/mL). The BHI agar plates were incubated at 37 °C for 48 h and after that period, the number of Colony Forming Units (CFU) were counted. The value of the dilutions was corrected and the result was graphed as log10 CFU/ml versus the vancomycin concentration using software Excel® (Microsoft, Windows 7, Redmond, WA, EUA). The following strains VSSA (ATCC 29213), hVISA Mu3 (ATCC 700698) and VISA Mu50 (ATCC 700699) were included as controls. Finally, AUC of the tested isolate/AUC of Mu3 ratio was calculated and interpreted as follows: for VSSA, less than 0.9; for hVISA, from 0.9 to 1.3; and for VISA, greater than 1.3.
DNA
Extraction and molecular typing
All MRSA were cultured on blood agar and incubated overnight at 37ºC. DNA was extracted using Chelex.100 (Bio-Rad, Richmond, CA) and Proteinase K (Sigma-Aldrich, Poole, UK). DNA samples were stored at -20ºC until use. Multiplex PCR was used to determine the SCCmec types I-X (including subtypes IVa, IVb, IVc and IVd) of all MRSA, as previously described.14, 17 NCTC 10442 (SCCmec I), N315 (SCCmec II), 85/2082 (SCCmec III), JCSC 4474 (SCCmec IVa), JCSC 2172 (SCCmec IVb), JCSC 4488 (SCCmec IVc), JCSC 4469 (SCCmec IVd), WIS (SCCmec V), HDE 288 (SCCmec VI), JCSC 6082 (SCCmec VII), JCSC 6943 (SCCmec IX) and JCSC 6945 (SCCmec X) were included as controls. The agr typing was performed by multiplex PCR according to the method published by Wu et al.18S. aureus COL (agr I), S. aureus N315 (agr II), S. aureus ATCC 25923 (agr III) and S. aureus A920210 (agr IV) were included as controls.
Statistical analysis
Statistical analyzes were determined using SPSS version 23.0 software (SPSS, Chicago, IL, USA). Chi-square test or Fisher’s exact test for categorical variables were used to calculate . values. A . value of < 0.05 was considered as statistically significant.
RESULTS
Antimicrobial susceptibility
Antimicrobial resistance profile showed lower levels of tetracycline resistance (22.0%), followed by trimethoprim/sulfamethoxazole (27.1%) and rifampicin (31.1%), among all 177 MRSA. On the other hand, higher resistance levels were observed for levofloxacin (49.7%), clindamycin (64.4%) and erythromycin (77.4%). Eighty-eight (49.7%) MRSA presented multidrug resistance (MDR, resistant to three or more antimicrobial classes). Teicoplanin resistance was not found among MRSA isolates.
All isolates were susceptible to vancomycin, with MIC values of 1 µg/mL (n=142; 80.2%) and 2 µg/mL (n=35; 19.8%). Screening for hVISA reveals that 67 (37.8%) isolates showed growth in BHIA-3V and, of these, only four were capable of growing in BHIA-6V. Population analysis profile confirmed that 9 (5.1%) were hVISA, and VISA phenotype was not detected by PAP-AUC (Figure 1). The phenotype VT-MRSA was found in 24 (13.6%) of MRSA isolates and all hVISA isolates were also classified as VT-MRSA (.<0.001) (Table 1).
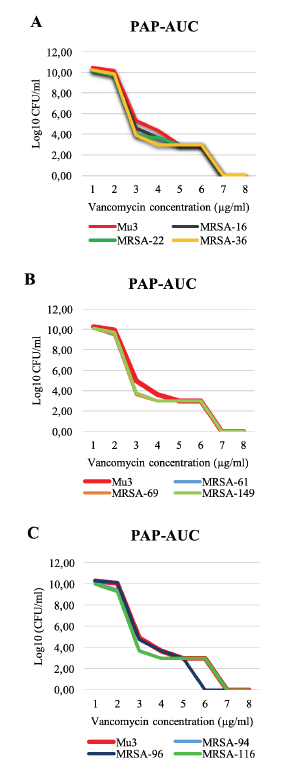
Figure 1
Growth curves of the nine isolates characterized as hVISA compared to the standard strain Mu3, by PAP-AUC method. All isolates demonstrated a ratio greater than 0.9. Graphs A, B, and C illustrate all hVISA isolates; the reference strain Mu3 is shown in all graphs.
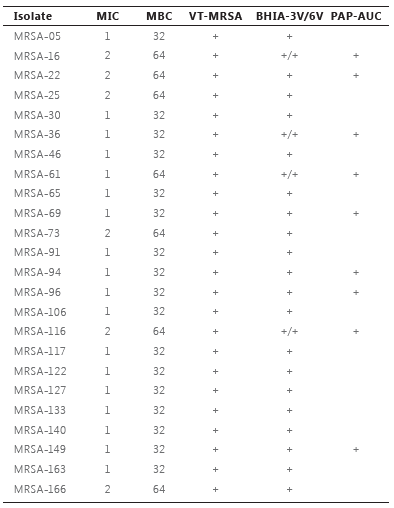
MIC = minimum inhibitory concentration; MBC = minimum bactericidal concentration; VT-MRSA = vancomycin tolerant methicillin-resistant Staphylococcus aureus; BHIA-3V/6V = BHIA with 3 or 6 µg/mL vancomycin; PAP-AUC = population analysis profile – area under the curve.
Significant differences in antimicrobial susceptibility pattern were observed between MRSA and VT-MRSA: all antimicrobial tested, except teicoplanin, demonstrated higher resistance rates in VT-MRSA when compared to MRSA. Also, hVISA demonstrated significantly higher resistance levels to rifampicin, trimethoprim/sulfamethoxazole and tetracycline than non-hVISA (Table 2). One of the possible reasons we could not find a significant difference in hVISA resistance levels for some antimicrobials is the low prevalence of these isolates in the study.
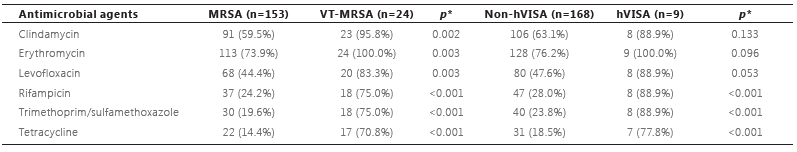
MRSA = methicillin-resistant Staphylococcus aureus; VT-MRSA = vancomycin tolerant methicillin-resistant Staphylococcus aureus; hVISA = heterogeneous vancomycin-intermediate Staphylococcus aureus. Number of isolates (percentage). *Chi-square test or Fisher’s exact test.
Molecular characteristics
SCCmec and agr typing are presented in Table 3. None of the isolates showed SCCmec types VII and X, and the prevalence of SCCmec IV subtypes was: 21 IVa, 2 IVb, 28 IVc and 6 IVd. Among VT-MRSA, the SCCmec type II was the most frequent followed by SCCmec type III. The presence of SCCmec type II in VT-MRSA was significantly higher than in non-VT-MRSA (.=0.025). Among hVISA isolates, the SCCmec type III was the most frequent followed by SCCmec II. There was no significant difference in the presence of SCCmec between hVISA and non-hVISA.
The majority of MRSA belonged to agr group II, followed by agr group I and III. agr group IV was not detected. Twenty-one isolates were untypeable by our assay. The presence of agr group II was significant in VT-MRSA (.=0.032) and hVISA (.=0.046) (Table 3).
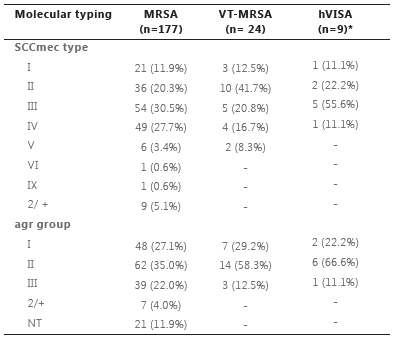
MRSA = methicillin-resistant Staphylococcus aureus; VT-MRSA = vancomycin tolerant methicillin-resistant Staphylococcus aureus; hVISA = heterogeneous vancomycin-intermediate Staphylococcus aureus; SCCmec = staphylococcal cassette chromosome mec; agr = accessory gene regulator. Number of isolates (percentage). *All hVISA isolates were also classified as VT-MRSA.
DISCUSSION
Although vancomycin remains the main therapeutic option for the treatment of MRSA infections, this therapeutic management might be reconsidered regarding epidemiological studies worldwide. In our results, almost 15% of the MRSA isolates presented tolerance to vancomycin, which is in accordance with findings from other studies.7, 19 The selection of VT-MRSA strains occurs likely due to exposure to suboptimal concentrations of vancomycin, which explains the development of hVISA and VISA strains and the association between VT-MRSA and hVISA.3,7,19
In this study, all MRSA were susceptible to vancomycin, with a MIC range of 1-2 μg/mL. Other studies have reported that infections caused by VS-MRSA have been associated with the increase in vancomycin treatment failures and mortality, particularly those isolates with MICs of 1.5 or 2 µg/mL.3,4Around 5% of MRSA isolates presented the hVISA phenotype. In previous reports, the frequency of the hVISA has ranged from 0 to 22.1% of MRSA.11,20,21
Their prevalence among MRSA varies significantly with the geographic area, the source of clinical isolates, and detection methods.11,21 SCCmec III was the most prevalent type among the MRSA strains, as observed in other studies, and types III and II were more prevalent among isolates with reduced susceptibility to vancomycin (hVISA and VT-MRSA), as also observed in previous studies.7,11,20,21,22 Regarding agr typing, group II was more frequent among MRSA isolates, as already seen in other analysis.8,13 Its was also high among VT-MRSA and hVISA, being associated with reduced vancomycin susceptibility.12, 23, 24 Many VISA are highly enriched for the agr group II. In this study, VISA was not detected, but hVISA and VT-MRSA demonstrated the same molecular characteristic previously described for VISA7. The agr group II has been strongly associated with vancomycin treatment failures in MRSA bacteremia, as well as the thickened cell wall in VT-MRSA that harbors SCCmec type II and agr group II.7,25 It suggests that the vancomycin resistance is adaptative and could be inducible by molecular characteristics.
In summary, our study detected the prevalence of SCCmec types II and III and agr group II in MRSA with reduced susceptibility to vancomycin and demonstrated that in fact, these isolates showed higher resistance levels to antimicrobials than non-VT-MRSA and non-hVISA isolates. Conventional tests to detect vancomycin susceptibility do not comprehend the PAP-AUC method, a time-consuming and labor-intensive technique, and this is the scenario that treatment failures by vancomycin may occur since isolates are underestimated due to routinely characterization as susceptible to vancomycin.
Acknowledgments
We are grateful for the fellowship by Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) for Adriana M. Rossato. We also offer thanks to Universidade Federal de Ciências da Saúde de Porto Alegre (UFCSPA) that provided technical support for the development of this study.
REFERENCES
1. Yu F, Liu Y, Lv J, et al. Antimicrobial susceptibility, virulence determinant carriage and molecular characteristics of Staphylococcus aureus isolates associated with skin and soft tissue infections. Braz J Infect Dis 2015; 19 (6): 614-622. http://dx.doi.org/10.1016/j.bjid.2015.08.006
2. Kong C, Neoh H, Nathan S. Targeting Staphylococcus aureus toxins: a potential form of anti-virulence therapy. Toxins 2016; 8 (3): 72. https://doi.org/10.3390/toxins8030072
3. Van Hal SJ, Lodise TP, Paterson DL. The clinical significance of vancomycin minimum inhibitory concentration in Staphylococcus aureus infections: a systematic review and meta-analysis. Clin Infect Dis 2012; 54 (6): 755-771. https://doi.org/10.1093/cid/cir935
4. Jacob JT, Diazgranados CA. High vancomycin minimum inhibitory concentration and clinical outcomes in adults with methicillin-resistant Staphylococcus aureus infections: a meta-analysis. Int J Infect Dis 2013; 17 (2): e93-e100. doi: 10.1016/j.ijid.2012.08.005
5. Casapao AM, Leonard SN, Davis SL, et al. Clinical outcomes in patients with heterogeneous vancomycin intermediate Staphylococcus aureus bloodstream infection. Antimicrob Agents Chemother 2013; 57 (9): 4252-4259. doi: 10.1128/AAC.00380-13
6. Ito T, Kuwahara-Arai K, Katayama Y, et al. Staphylococcal cassette chromosome mec (SCCmec) analysis of MRSA. Methicillin-resistant Staphylococcus aureus (MRSA) protocols. Methods Mol Biol 2014; 1085: 131-148. doi: 10.1007/978-1-62703-664-1_8.
7. Cázares-Domínguez V, Cruz-Córdova A, Ochoa AS, et al. Vancomycin tolerant, methicillin-resitant Staphylococcus aureus reveals the effects of vancomycin on cell wall thickening. PLoS One 2015a; 10 (3): e0118791. https://doi.org/10.1371/journal.pone.0118791
8. Kang CK, Cho JE, Choi YJ, et al. agr dysfunction affects staphylococcal cassette chromosome mec type dependent clinical outcomes in methicillin-resistant Staphylococcus aureus bacteremia. Antimicrob Agents Chemother 2015; 59 (6): 3125-3132. doi: 10.1128/AAC.04962-14
9. Di Gregorio S, Perazzi B, Ordonez AM, et al. Clinical, microbiological, and genetic characteristics of heteroresistant vancomycin-intermediate Staphylococcus aureus bacteremia in a teaching hospital. Microb Drug Resist 2015; 21 (1): 25-34. https://doi.org/10.1089/mdr.2014.0190
10. Hu HC, Kao KC, Chiu LC, et al. Clinical outcomes and molecular typing of heterogenous vancomycin-intermediate Staphylococcus aureus bacteremia in patients in intensive care units. BMC Infect Dis 2015; 15: 444. https://doi.org/10.1186/s12879-015-1215-2
11. Huang SH, Chen YC, Chuang YC, et al. Prevalence of vancomycin-intermediate Staphylococcus aureus (VISA) and heterogeneous VISA among methicillin-resistant S. aureus with high vancomycin minimal inhibitory concentrations in Taiwan: A multicenter surveillance study, 2012 – 2013. J Microbiol Immunol Infect 2015; 49 (5): 701-707. https://doi.org/10.1016/j.jmii.2015.07.003
12. Bibalan MH, Shakeri F, Javid N, et al. Accessory gene regulator types of Staphylococcus aureus isolated in Gorgan, North of Iran. J Clin Diagn Res 2014; 8 (4): DC07- DC09. doi: 10.7860/JCDR/2014/6971.4219
13. Cázares-Domínguez V, Ochoa1 SA, Cruz-Córdova A, et al. Vancomycin modifies the expression of the agr system in multidrug-resistant Staphylococcus aureusclinical isolates. Front Microbiol 2015b; 6: 369. https://doi.org/10.3389/fmicb.2015.00369
14. Lawung R, Chuong LV, Cherdtrakulkiat R, et al. Revelation of staphylococcal cassette chromosome mec types in methicillin-resistant Staphylococcus aureus isolates from Thailand and Vietnam. J Microbiol Methods 2014; 107 (1): 8-12. https://doi.org/10.1016/j.mimet.2014.08.024
15. Clinical and Laboratory Standards Institute. (CLSI). Performance standards for antimicrobial susceptibility testing: twenty-fourth informational supplement. Document M100-S24. Wayne, PA: CLSI; 2015. p.124-139
16. Riederer K, Shemes S, Chase P, et al. Detection of intermediately vancomycin-susceptible and heterogeneous Staphylococcus aureus isolates: comparison of etest and agar screening methods. J Clin Microbiol 2011; 49(6): 2147-2150. doi:10.1128/JCM.01435-10
17. Kondo Y, Ito T, Ma XX, et al. Combination of multiplex PCRs for staphylococcal cassette chromosome mec type assignment: rapid identification system for mec, ccr, and major differences in junkyard regions. Antimicrob Agents Chemother 2007; 51 (1): 264-274. doi: 10.1128/AAC.00165-06
18. Wu D, Li X, Yang Y, et al. Superantigen gene profiles and presence of exfoliative toxin genes in community-acquired meticillin-resistant Staphylococcus aureus isolated from Chinese children. J Med Microbiol 2011; 60 (1): 35-45. doi: 10.1099/jmm.0.023465-0
19. Rose WE, Fallon M, Moran JJM, et al. Vancomycin tolerance in methicillin-resistant Staphylococcus aureus: influence of vancomycin, daptomycin, and telavancin on differential resistance gene expression. Antimicrob Agents Chemother 2012; 56 (8): 4422-4427. doi:10.1128/AAC.00676-12
20. Liu C, Chen ZJ, Sun Z, et al. Molecular characteristics and virulence factors in methicillin-susceptible, resistant, and heterogeneous vancomycin-intermediate Staphylococcus aureus from central-southern China. J Microbiol Immunol Infect 2015; 48 (5): 490-496. doi: 10.1016/j.jmii.2014.03.003
21. Zhang S, Sun X, Chang W, et al. Systematic review and meta-analysis of the epidemiology of vancomycin-intermediate and heterogeneous vancomycin-intermediate Staphylococcus aureus isolates. PLoS ONE 2015; 10 (8): e0136082. https://doi.org/10.1371/journal.pone.0136082
22. Cechinel A, Machado DP, Turra E, et al. Association between accessory gene regulator polymorphism and mortality among critically Ill patients receiving vancomycin for nosocomial MRSA bacteremia: a cohort study. Can J Infect Dis Med Microbiol 2016; 2016: 1-5. http://dx.doi.org/10.1155/2016/8163456
23. Hu J, Ma XX, Tian Y, et al. Reduced vancomycin susceptibility found in methicillin-resistant and methicillin-sensitive Staphylococcus aureus clinical isolates in Northeast China. PLoS ONE 2013; 8 (9): e73300. https://doi.org/10.1371/journal.pone.0073300
24. Azimian A, Havaei SA, Fazeli H, et al. Genetic characterization of a vancomycin-resistant Staphylococcus aureus isolate from the respiratory tract of a patient in a university hospital in Northeastern Iran. J Clin Microbiol 2012; 50 (11): 3581-3585. doi: 10.1128/JCM.01727-12
25. Sakoulas G, Eliopoulos GM, Moellering RC, et al. Accessory gene regulator (agr) locus in geographically diverse Staphylococcus aureus isolates with reduced susceptibility to vancomycin. Antimicrob Agents Chemother 2002; 46 (5): 1492-1502. doi: 10.1128/AAC.46.5.1492–1502.2002