Artículos de investigación
Geastrum chamelense (Geastraceae, Agaricomycetes), a new species with setose endoperidium from the tropical dry forest in Jalisco, Mexico
Geastrum chamelense (Geastraceae, Agaricomycetes), una nueva especie con endoperidio setoso del bosque tropical caducifolio en Jalisco, México
Geastrum chamelense (Geastraceae, Agaricomycetes), a new species with setose endoperidium from the tropical dry forest in Jalisco, Mexico
Acta botánica mexicana, no. 128, e1905, 2021
Instituto de Ecología A.C., Centro Regional del Bajío
Received: 04 June 2021
Revised document received: 02 September 2021
Accepted: 01 December 2021
Published: 07 December 2021
Funding
Funding source: Instituto Politécnico Nacional (IPN)
Contract number: SIP-20210135
Contract number: SIP-20210661
Funding statement: PAPIIT IN-218008 and IN-207311. TR and RV thank the Instituto Politécnico Nacional (IPN) for financial support for their research in the projects SIP-20210135 and SIP-20210661. RV and TR are grateful to the Sistema Nacional de Investigadores (CONACYT)
Abstract
Background and Aims: Geastrum is characterized by stelliform basidiomata, exoperidium with three layers, sessile or stalked endoperidium, and sulcate, plicate, folded or fibrillose peristome, distinctly or indistinctly delimited, sometimes with mycosclereids. The objective of this study is to describe and illustrate Geastrum chamelense with morphological, ecological and molecular data as a new species from the Chamela Biological Station, Jalisco, Mexico.
Methods: Basidiomata collections with different degrees of maturity gathered in 2010 and 2011 from tropical dry forest at the Chamela Biological Station in Jalisco state are described macro- and microscopically. The type material is deposited in the fungal collections of the herbaria ENCB and MEXU. The extraction of DNA, as well as the phylogenetic analyses of ITS, LSU, atp6 and rpb1 sequences, are based on the holotype.
Key results: Geastrum chamelense is distinguished by its greyish brown basidiomata, pseudofornicate, fleshy exoperidium, not hygroscopic, sessile endoperidium, subglobose to depressed with peristome plicate, not delimited, and setae present. The latter character is shared with Geastrum setiferum from Brazil, but that species has shorter and wider setae (95-215 × 20-47 µm) than G. chamelense (102-330 × 10.2-15.3 µm). From a phylogenetic perspective, G. chamelense is sister to G. hieronymi and G. cf. calceum, while G. setiferum is not related, as it appears in a separate clade.
Conclusions: Geastrum chamelense is recognized as a new species based on morphological, ecological and molecular data.
Key words: earthstars fungi+ Geastrales+ phylogeny+ taxonomy.
Resumen
Antecedentes y Objetivos: Geastrum se caracteriza por presentar basidiomas esteliformes, exoperidio con tres capas, endoperidio sésil o estipitado, peristoma sulcado, plegado, doblado o fibriloso, distintiva o indistintivamente delimitado, algunas veces con micoesclereidas. El objetivo de este estudio es describir e ilustrar a Geastrum chamelense con datos morfológicos, ecológicos y moleculares como una especie nueva de la Estación de Biología Chamela, Jalisco, México.
Métodos: Las colecciones de basidiomas con diferentes grados de madurez recolectados en 2010 y 2011 del bosque tropical caducifolio de la Estación de Biología Chamela en el estado de Jalisco se describen macro y micromorfológicamente. El material tipo está depositado en las colecciones micológicas de los herbarios ENCB y MEXU. La extracción de ADN, así como los análisis filogeneticos de secuencias de ITS, LSU, atp6 y rpb1, se basan en el holotipo.
Resultados clave: Geastrum chamelense se distingue por sus basidiomas pardo grisáceos, exoperidio pseudofornicado, carnoso, no higroscópico y endoperidio sésil, subgloboso a deprimido con peristoma plegado, no delimitado, y presencia de setas. Este último carácter se comparte con Geastrum setiferum de Brasil, pero ésta tiene setas más cortas y anchas (95-215 × 20-47 µm) que G. chamelense (102-330 × 10.2-15.3 µm). Desde una perspectiva filogenética, G. chamelense tiene como grupo hermano a G. hieronymi y G. cf. calceum, mientras que G. setiferum no está relacionado porque aparece en un clado separado.
Conclusiones: Geastrum chamelense es reconocida como una especie nueva basada en datos morfológicos, ecológicos y moleculares.
Palabra clave: estrellas de tierra, Geastrales, filogenia, taxonomía.
Introduction
The Chamela Biological Station of the Universidad Nacional Autónoma de México is located within the Chamela-Cuixmala Biosphere Reserve in the municipality La Huerta, Jalisco state, Mexico (Fig. 1). The station has a predominance of tropical dry forest, which safeguards one of the best conserved tropical communities, and this has great biological and ecological importance (Noguera et al., 2002). Although Mexico has extensive areas with tropical dry forest, 160 000 km2 approximately (Challenger, 1998), knowledge of mycobiota in this kind of vegetation and particularly in the Chamela Biological Station is scarce, compared to the great diversity of fungal species in the tropical dry forest (Hawksworth, 1993) and the station. There are several studies that include species from the tropical dry forest such as those from Esqueda et al. (1999), Raymundo et al. (2009, 2014, 2017), Salinas-Salgado et al. (2012), Valenzuela et al. (2012), Álvarez et al. (2016), Contreras-Pacheco et al. (2018), and Reyes et al. (2020). However, few studies are known from the Chamela Biological Station: Ramírez-López et al. (2012), Bautista-Hernández et al. (2015), and Raymundo (2021).
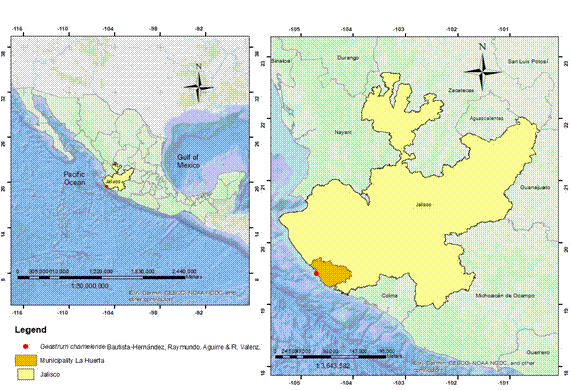
Figure 1:
Type locality of Geastrum chamelense Bautista-Hernández, Raymundo, Aguirre & R. Valenz. in the Chamela Biological Station, Chamela, Jalisco, Mexico.
Seventeen species of Geastrum Pers. have been registered in this type of ecosystem (Esqueda et al., 1996, 1999, 2000, 2003, 2009; Pérez-Silva et al., 1999; Guzmán, 2003; Calonge and Mata, 2004; Calonge et al., 2004; Herrera et al., 2005; Bautista-Hernández et al., 2015). This genus belongs to the family Geastraceae, order Geastrales, subclass Phallomycetidae, class Agaricomycetes, subphylum Agaricomycotina of the phylum Basidiomycota (Index Fungorum, 2021). Three species, Geastrum fimbriatum Fr., G. saccatum Fr. and G. violaceum Rick, have been cited from the Chamela Biological Station (Pérez-Silva et al., 1999; Bautista-Hernández et al., 2015). This genus is characterized by stelliform basidiomata, exoperidium with three layers, sessile or stalked endoperidium, sulcate, plicate, folded or fibrillose peristome, distinctly or indistinctly delimited. Microscopically, it can present ornamented basidiospores and mycosclereids or protruding hyphae (Sunhede, 1989), and setae (Baseia and Milanez, 2002). During explorations carried out in the station about a decade ago, a great diversity of Geastrum species was observed, and several specimens with particular macro- and micromorphological characteristics uncommon in the genus called our attention. The objective of this study is to describe and illustrate Geastrum chamelense as a new species for the Chamela Biological Station, based on morphological, ecological and molecular data.
Materials and Methods
Field work
Material of the undescribed species was collected in September 2010 and 2011 in the Chamela Biological Station, Jalisco, Mexico, ca. 19°27'2.1"N, 105°01'33"W, 250 m a.s.l. (Fig. 1, coordinates and elevation were obtained with a Garmin Etrex 10 GPS, Kansas City, USA). The local ecosystem belongs to the semideciduous tropical forest, according to Rzedowski (2006), with Brosimum alicastrum Sw. and Celtis monoica Hemsl. as the dominant tree species. The holotype was deposited in the fungal collection in the Herbarium of the Escuela Nacional de Ciencias Biológicas of the Instituto Politécnico Nacional (ENCB) and the isotype in the fungal collection of the National Herbarium of Mexico (MEXU) of the Instituto de Biología of the Universidad Nacional Autónoma de México (UNAM).
Morphological analyses
This taxonomic study was based on collections of basidiomata with different degrees of maturity. Morphological examinations were conducted using protocols outlined by Sunhede (1989). The colour of the sporomata was coded according to Kornerup and Wanscher (1978), which is indicated in parentheses in the description. For the microscopic study, temporary preparations were made in 70% alcohol and 5% potassium hydroxide (KOH) to elaborate descriptions of colour, size, shape of basidiospores, setae, and hyphae. The length and width of thirty basidiospores and setae were measured with a micrometric scale. Using scanning microscopy, gleba and endoperidium preparations were made, observing the detail of the spore and capillitium ornamentation, as well as the surface of the endoperidium. For the morphometric study, an optical microscope (MO; Primo Star, Carl Zeiss, Göttingen, Germany) and a scanning electron microscope (SEM; Hitachi Su 1510, Hitachi, Japan) were used. To prepare the taxonomic key, the abovementioned species known to occur in the study area were incorporated.
DNA extraction, amplification, and sequencing
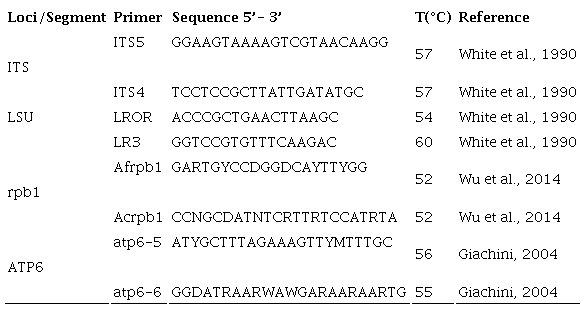
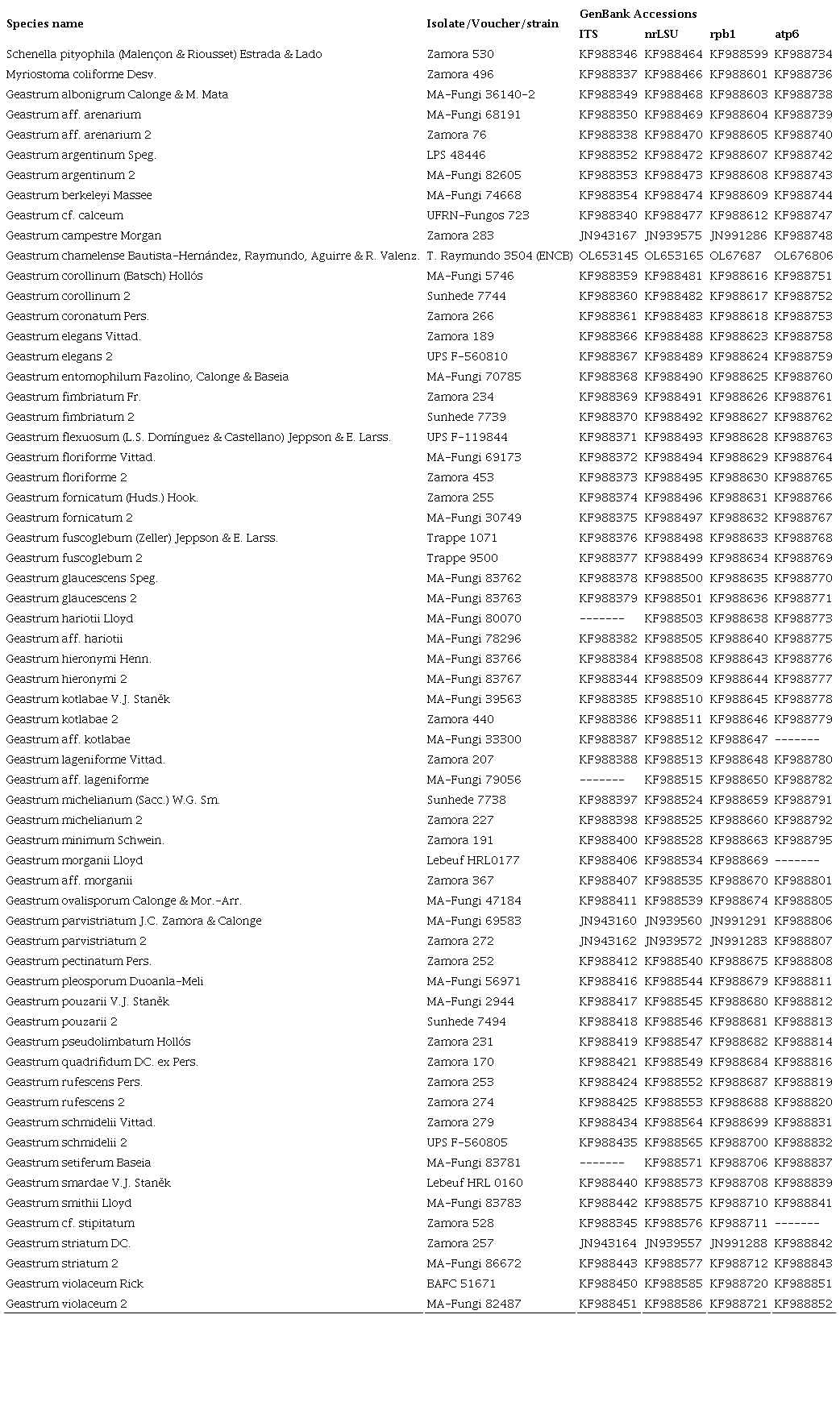
The DNA was obtained from herbarium specimens (Table 1). The CTAB protocol of Martínez-González et al. (2017) was used to extract genomic DNA. The DNA was quantified with a Nanodrop 2000c (Thermo ScientificTM, Wilmington, USA). We prepared dilutions from each sample at 20 ng/µl to amplify the next four regions (Table 2): mitochondrial ATPase subunit 6 (atp6), nuclear large subunit ribosomal DNA (LSU), Internal Transcribed Spacer (ITS) and the largest subunit of RNA polymerase II gene (rpb1). The reaction mixture for PCRs was performed on a final volume of 15 µl containing 1× buffer, 0.8 mM dNTPs mix, 20 pmol of each primer, 2 units of GoTaq DNA (Promega, USA) and 100 ng of template DNA. The PCR products were verified by agarose gel electrophoresis. The gels were run for 1 h at 95 V cm⁻³ in 1.5% agarose and 1× TAE buffer (Tris Acetate-EDTA). The gel was stained with GelRed (Biotium, USA) and the bands were visualized in an Infinity 3000 transilluminator (Vilber Lourmat, Eberhardzell, Germany). The amplified products were purified with the ExoSAP Purification kit (Affymetrix, USA), following the manufacturer’s instructions. They were quantified and prepared for the sequence reaction using a BigDye Terminator v. 3.1 (Applied Biosystems, USA). These products were sequenced in both directions with an Applied Biosystem model 3730XL (Applied BioSystems, Foster City, USA), at the Instituto de Biología, UNAM. The sequences obtained were compared with the original chromatograms to detect and correct possible reading errors. The sequences of both strands of each of the genes were analyzed, edited and assembled using the BioEdit v. 7.0.5 (Hall, 1999) to generate a consensus sequence which were compared with those deposited in GenBank (2020), using the tool BLASTN v. 2.2.19 (Zhang et al., 2000).

Phylogenetic analyses
To explore the phylogenetic relationships of the new species, an alignment was made based on the taxonomic sampling employed by Zamora et al. (2014). Each gene region was independently aligned using the online version of MAFFT v. 7 (Katoh et al., 2002, 2017; Katoh and Standley, 2013). Alignments were reviewed in PhyDE v. 10.0 (Müller et al., 2005), followed by minor manual adjustments to ensure character homology between taxa. The matrices were formed for ITS by 64 taxa (667 characters), for LSU by 63 taxa (875 characters), atp6 by 64 taxa (451 characters), while that of rpb1 consisted of 64 taxa (684 characters). The aligned matrices were concatenated into a single matrix (64 taxa, 2677 characters). Eight partitioning schemes were established: one for the ITS, one for the LSU, three to represent the codon positions of the gene region atp6 and three for the rpb1 gene region, which were established using the option to minimize the stop codon with Mesquite v. 3.70 (Maddison and Maddison, 2021). The best evolutionary model for alignment was sought using PartitionFinder v. 2 (Lanfear et al., 2014, 2016; Frandsen et al., 2015). Phylogeny was performed with Bayesian inference using MrBayes v. 3.2.6 x64 (Huelsenbeck and Ronquist, 2001). The information block for the matrix included two simultaneous runs, four Montecarlo chains, temperature set to 0.2 and sampling 10 million generations (standard deviation ≤0.1) with trees sampled every 1000 generations. The convergence of the chains was displayed in Tracer v. 1 (Suchard et al., 2018). The highest credibility phylogram of the clades recovered with TreeAnnotator v. 1.8 (Bouckaert et al., 2014) was chosen with a 25% burn-in. Trees were visualized and optimized in FigTree v. 1.4.4 (Suchard et al., 2018).
Results
Molecular analysis
The ITS, LSU, atp6 and rpb1 sequences obtained from Geastrum chamelense were deposited in GenBank (Table 2). In the Bayesian analysis, the standard deviation between the chains stabilized at 0.002 after 10 million generations, indicating MC3 reached a stationary phase. To confirm that the sample size was sufficient, the parameter file was examined in Tracer v. 1.6 (Suchard et al., 2018): all parameters had an estimated sample size of over 1500. The posterior probabilities (PP) obtained were estimated by generating a strict consensus tree in MrBayes. Bayesian inference analysis recovered a well-supported clade (PP=1) of the new species (Fig. 2).
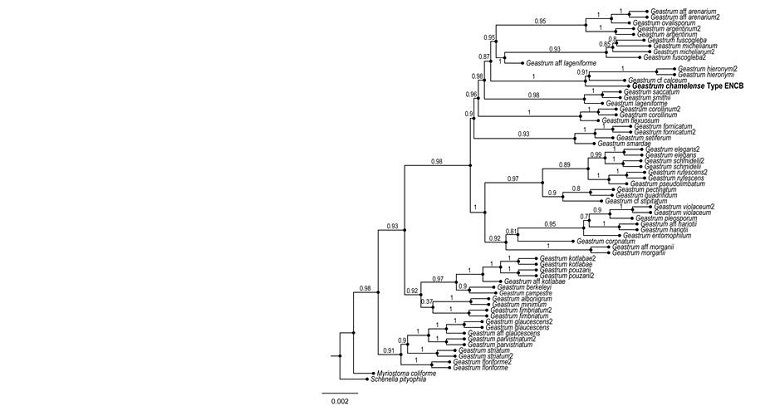
Figure 2:
Bayesian inference phylogram of ITS, LSU, rpb1, atp6 sequences. The new species Geastrum chamelense Bautista-Hernández, Raymundo, Aguirre & R. Valenz. is shown in bold.
The topology of the phylogenetic tree is similar to that reported by Zamora et al. (2014). The new species is phylogenetically distant from G. setiferum Baseia, the taxon with which it bears the greatest morphological similarity, sharing the presence of setae in the endoperidium. It forms a well-supported clade with G. hieronymi Henn. and G. cf. calceum.
Key to species of Geastrum found in the Chamela Biological Station, Jalisco, Mexico
-
1a. Basidiomata with setae on the endoperidial surface .... Geastrum chamelense Bautista-Hernández, Raymundo, Aguirre & R. Valenz.
-
1b. Basidiomata without setae on the endoperidial surface ........................................ 2
-
2a. Exoperidium pink to purplish ….………………… Geastrum violaceum Rick
-
2b. Exoperidium brown grey, brown orange, beige …………………….………….. 3
-
3a. Basidiomata growing on rotten wood or dead leaves, with white subiculum ………………………………… G. schweinitzii (Berk. & M.A. Curtis) Zeller.
-
3b. Basidiomata growing on soil, without white subiculum .................................. 4
-
4a. Basidiomata separating the mycelial layer from the exoperidium easily at maturity ................................................. G. javanicum Lév.
-
4b. Basidiomata with arachnoid aspect, exoperidium with longitudinal ridges, the mycelial layer not easily separating from the exoperidium ........... G. lageniforme Vittad.
Taxonomy
Phylum Basidiomycota
Subphylum Agaricomycotina
Class Agaricomycetes
Subclass Phallomycetidae
Order Geastrales
Family Geastraceae
Geastrum chamelense Bautista-Hernández, Raymundo, Aguirre & R. Valenz., sp. nov. Figs. 3, 4, 5.
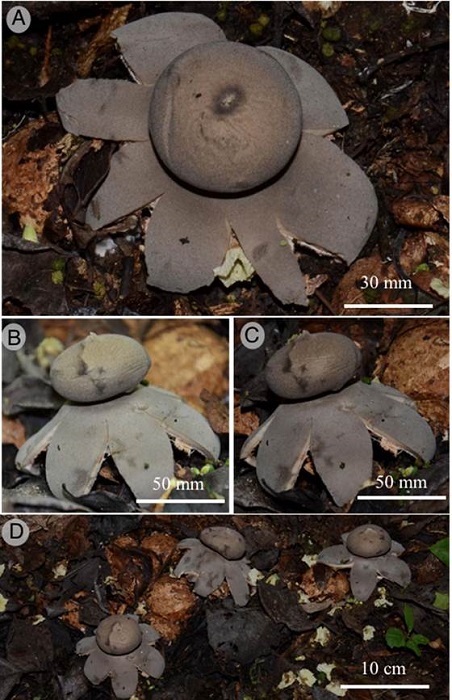

Figure 3:
Geastrum chamelense Bautista-Hernández, Raymundo, Aguirre & R. Valenz. A. endoperidium showing the peristome; B-C. basidiomata showing the expanded exoperidium; D. basidiomata in the field.
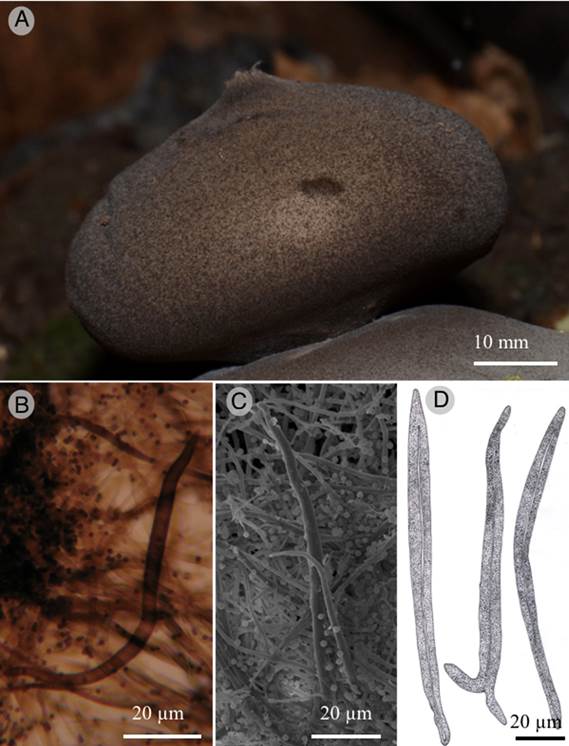

Figure 4:
Geastrum chamelense Bautista-Hernández, Raymundo, Aguirre & R. Valenz. A. endoperidial body, the blackish appearance is due to the presence of setae; B. setae (MO); C. setae (SEM); D. setae (drawing).
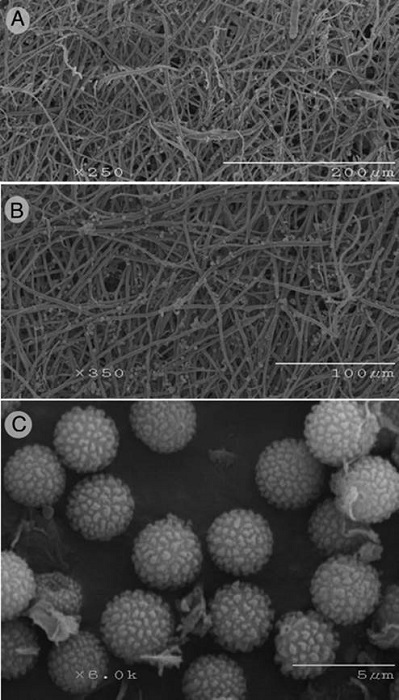

Figure 5:
Geastrum chamelense Bautista-Hernández, Raymundo, Aguirre & R. Valenz. A. endoperidium surface showing the setae (SEM); B. capillitium (SEM); C. basidiospores (SEM).
TYPE: MEXICO. Jalisco, municipality La Huerta, Reserva de la Biosfera Chamela - Cuixmala, Chamela Biology Station, km 60 Barra de Navidad-Puerto Vallarta highway, Eje Central, 250 m, 19°27'2.1"N, 105°01'33"W, 28.IX.2010. T. Raymundo 3504 (holotype: ENCB!, isotype: MEXU!); Mycobank: MB839090.
Geastrum chamelense is distinguished from other species of the genus Geastrum by its depressed, globose, semifornicate basidiomata, exoperidium 60 mm diameter, splitting into 4-7 rays, setose endoperidium, setae 102-330 × 10.2-15.3 μm, plicate peristome, not delimited; basidiospores 4.2-5 μm, globose, densely warty, dark brown.
Basidioma unexpanded, semihypogeous, depressed globose, light brown (5D6), 30 mm diameter; exoperidium 60 mm diameter, splitting into 4-7 rays, with the tips recurved, semifornicate horizontally, brown grayish (10E3), consistency carnose, in dry specimens not hygroscopic; mycelial layer attached to the litter, dimitic, 2.1-2.8 μm wide, with skeletal hyphae, thick-walled, aseptate, not branched, greenish yellow, light brown (5D6); fibrous layer 3.2-4 μm wide, thick-walled, light yellow; pseudoparenchymatous layer 14.4 -34.8 × 12-24 μm, globose to subglobose, brown yellow; without mycelial cords; endoperidium setose, sessile, subglobose to depressed, 15 × 20-30 mm, concolorous with exoperidium, constituted by interwoven hyphae, thick-walled, greenish yellow, 5.1-6.8 μm wide; setae 102-330 × 10.2-15.3 μm, thick-walled, lumen narrow; peristome plicate, not delimited; mycosclereids absent; gleba chocolate brown to blackish; basidiospores 4.2-5 μm diameter, globose, densely warty, dark brown in KOH 5%; basidia not observed; capillitial hyphae 5.6-7.2 μm diameter, aseptate, brown in KOH 5%, thick-walled, lumen narrow, not branched, surface with small warts and litter.
Habit and habitat: growing gregarious on soil in tropical dry forest.
Distribution: only known from the type locality.
Etymology: the specific epithet chamelense refers to the Chamela Biological Station, where this species has been collected.
Additional material examined: MEXICO. Jalisco, municipality La Huerta, Chamela-Cuixmala Biosphere Reserve, Chamela Biology Station, km 60 Barra de Navidad - Puerto Vallarta highway, Eje Central, 50 m, 19°27'2.1"N, 105°01'33"W, 18.IX.2011, T. Raymundo 4064 (ENCB); loc. cit., R. Valenzuela 14534 (ENCB); loc. cit., 28.IX.2010, E. Aguirre y S. Bautista-Hernández (MEXU 27044).
Geastrum javanicum, Búho, 23.IX.2012, E. Aguirre y S. Bautista-Hernández (MEXU 28929).
Geastrum lageniforme, Eje Central, 27.IX.2013, E. Aguirre y S. Bautista-Hernández (MEXU 28996).
Geastrum schweinitzii, Eje Central, 18.IX.2011, E. Aguirre-Acosta (MEXU 28883). Geastrum violaceum, Tejón, 21.X.2009, E. Aguirre y S. Bautista-Hernández (MEXU 25836).
Discussion
Geastrum chamelense was macroscopically characterized because the basidiomata, when mature, have a grey to greyish brown colour, a globose depressed endoperidium, folded and non-delimited peristoma, and a non-hygroscopic and arcuate exoperidium. Microscopically, it presented densely ornamented spores and setiform hyphae on the surface of the endoperidium, which, under a stereomicroscope, were observed as small erect blackish brown spines.
A similar species is G. setiferum, described from Brazil (Baseia and Milanez, 2002; Trierveiler-Pereira et al., 2011) and Argentina (Castiglia et al., 2013), which presents setae in the endoperidium. However, macro- and micromorphological differences delimit both taxa, such as the peristome, colour of the basidiome, and size of the setae and spores, indicating that this is a new species. Baseia and Milanez (2002) described G. setiferum with a fibrillose to almost sulcate and defined peristoma, while Trierveiler-Pereira et al. (2011) mentioned that it was fibrillose to slightly plicated. However, the specimen from Argentina was reported as conical to mammiform, with the apex truncated, finely plicated, and not delimited (Castiglia et al., 2013), showing a heterogeneity with this characteristic in both Brazilian and Argentinian specimens, in addition to presenting a pseudo-stipe and apophysis (Trierveiler-Pereira et al., 2011; Castiglia et al., 2013). In the Mexican specimens, few folds were observed, and it was sessile, not delimited, and without apophysis. Furthermore, the colour of the endoperidium differed because in the specimens from Brazil and Argentina, the tones ranged from greyish orange to light brown, while in the Mexican ones, it was greyish brown. Although macroscopically they are similar, we considered that the differences between the microscopic characteristics were preponderant for the separation of both species.
Geastrum setiferum has setae of 95-215 × 20-47 µm (Baseia and Milanez, 2002; Trierveiler-Pereira et al., 2011; Castiglia et al., 2013), differing notably in the proposed taxon which has setae that are longer and slender, measuring up to 330 × 12-16 µm. Additionally, the setae were observed with dichotomous terminations, a character that was not addressed by the aforementioned authors. Regarding the size of the spores, those of G. setiferum measure 2.5-4 µm diameter (Baseia and Milanez, 2002; Trierveiler-Pereira et al., 2011), while in G. chamelense, they are 4.2-4.9 µm diameter, including ornamentation in both species. Therefore, our new species has larger spores.
Molecular analysis showed that G. hieronymi is found in the same clade as G. chamelense, sharing an arched exoperidium, fibrillose peristome, endoperidial surface with spines, which are formed by bundles of hyphae (Zamora et al., 2014) or strongly asperate with acute or subpyramidal spicules (Ponce de León, 1968). Although these authors refer to G. hieronymi as having a stalked endoperidial body and prominent apophysis, this character was not observed in the studied specimens. Regarding the microscopic characteristics, the spores of G. hieronymi are much larger, up to 6 (m in diameter and warty (Ponce de León, 1968). Therefore, the differences are notable between both species.
Conclusions
Geastrum chamelense is recognized as a new species based on morphological, ecological and molecular data. Although this species is close to G. setiferum because they share the setiferous elements in the endoperidium, the macro- and micromorphological characters and its position in the phylogenetic hypothesis based on ITS, LSU, rpb1 and atp6 markers were decisive to separate them as different species. Worldwide, Geastrum has 109 valid species (Index Fungorum, 2021); of those, 29 species (26.6%) have been reported in Mexico, including G. chamelense. It is important to continue with taxonomic studies of this genus to contribute with new records and new species for the Mexican mycobiota.
Literature cited
Álvarez, V. I., T. Raymundo and R. Valenzuela. 2016. Hongos histerioides (Dothideomycetes, Ascomycota) del bosque tropical caducifolio en el Parque Nacional Lagunas de Chacahua, Oaxaca, México. Acta Botanica Mexicana 116: 49-64. DOI: http://doi.org/10.21829/abm116.2016.1118
Baseia, I. G. and A. I. Milanez. 2002. Geastrum setiferum (Gasteromycetes): a new species with a setose endoperidium. Mycotaxon 84: 135-139.
Bautista-Hernández, S., E. Aguirre-Acosta, T. Raymundo , T. Herrera, J. Cifuentes, S. Cappello-García and R. Valenzuela. 2015. Geastrum violaceum, registro nuevo para México. Revista Mexicana de Biodiversidad 86(3): 569-572. DOI: http://dx.doi.org/10.1016/j.rmb.2015.05.005
Bouckaert, R., J. Heled, D. Kühnert, T. Vaughan, C. H. Wu, D. Xie, M. A. Suchard, A. Rambaut and A. J. Drummond. 2014. BEAST 2: A software platform for Bayesian Evolutionary analysis. PLOS Computational Biology 10(4): e1003537. DOI: https://doi.org/10.1371/journal.pcbi.1003537
Calonge, F. D. and M. Mata. 2004. A new species of Geastrum from Costa Rica and Mexico. Boletín de la Sociedad Micológica de Madrid 28: 331-335.
Calonge, F. D. , G. Guzmán and F. Ramírez-Guillén. 2004. Observaciones sobre los Gasteromycetes de México depositados en los Herbarios XAL y XALU. Boletín de la Sociedad Micológica de Madrid 28: 337-371.
Castiglia, V., I. Cinto and L. Papinutti. 2013. Geastrum trichiferum and Geastrum setiferum (Geastrales, Phallomycetidae) new records for Argentina. Boletín de la Sociedad Argentina de Botánica 48(3-4): 377-380.
Challenger, A. 1998. Utilización y Conservación de los Ecosistemas Terrestres de México. Pasado, presente y futuro. Comisión Nacional para el Conocimiento y Uso de la Biodiversidad (CONABIO), Instituto de Biología, UNAM, Agrupación Sierra Madre, S.C. México. 847 pp.
Contreras-Pacheco, M., R. Valenzuela, T. Raymundo and L. Pacheco. 2018. Hymenochaete cifuentesii, H. potosina and H. raduloides spp nov. from the tropical dry forest of Mexico. Mycotaxon 133(3): 499-512. DOI: https://doi.org/10.5248/133.499
Esqueda, M., E. Pérez-Silva, T. Herrera and R. E. Villegas. 1996. Los Gasteromycetes de Sonora. CESUES 1(2): 3-16.
Esqueda, M. , E. Pérez-Silva , F. San Martín and R. Santos-Guzmán. 1999. Macromicetos de selva baja caducifolia, I. Álamos, Sonora, México. Revista Mexicana de Micología 15: 73-78.
Esqueda, M. , E. Pérez-Silva , T. Herrera , M. L. Coronado and A. Estrada-Torres. 2000. Composición de gasteromicetos en un gradiente de vegetación en Sonora, México. Anales del Instituto de Biología serie Botánica UNAM 71(2): 39-62.
Esqueda, M. , T. Herrera , E. Pérez-Silva and A. Sánchez. 2003. Distribution of Geastrum species from some priority regions for conservation of biodiversity of Sonora, Mexico. Mycotaxon 87: 445-456.
Esqueda, M. A. Sánchez, M. Rivera, M. L. Coronado , M. Lizárraga andR. Valenzuela . 2009. Primeros registros de hongos gasteroides en la Reserva Forestal Nacional y Refugio de Fauna Silvestre Ajos-Bavispe, Sonora, México. Revista Mexicana de Micología 30: 19-29.
Frandsen, P. B., B. Calcott, C. Mayer and R. Lanfear. 2015. Automatic selection of partitioning schemes for phylogenetic analyses using iterative k-means clustering of site rates. BMC Evolutionary Biology 15(1): 13. DOI: https://doi.org/10.1186/s12862-015-0283-7
GenBank. 2020. National Center for Biotechnology Information http://www.ncbi.nlm.nih.gov/genbak/ (consulted October, 2021)
Giachini, A. J. 2004. Systematics, phylogeny, and ecology of Gomphus sensu lato. Doctoral Dissertation. Oregon State University. Corvallis, USA.
Guzmán, G. 2003. Los Hongos de El Edén, Quintana Roo, Introducción a la micobiota tropical de México. Comisión Nacional para el Conocimiento y Uso de la Biodiversidad (CONABIO), Instituto de Ecología, A.C. (INECOL). Xalapa, México. 316 pp.
Hall, T. A. 1999. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series 41: 95-98.
Hawksworth, D. L. 1993. The tropical fungal biota. Census, pertinence, prophylaxis, and prognosis. In: Isaac, S., J. C. Frankland, R. Watling and A. J. S. Walley (eds.). Aspects of tropical mycology. British Mycological Society & Cambridge University Press. Cambridge, UK. Pp. 265-293.
Herrera, T., E. Pérez-Silva, M. Esqueda and V. H. Valenzuela. 2005. Algunos gasteromicetos de Calakmul, Campeche, México. Revista Mexicana de Micología 21: 23-27.
Huelsenbeck, J. P. and F. Ronquist. 2001. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17(8): 754-755. DOI: https://doi.org/10.1093/bioinformatics/17.8.754
Index Fungorum. 2021. An international project to index all formal names in the Fungi Kingdom. http://www.indexfungorum.org/names/Names.asp (consulted November, 2021).
Katoh, K., K. Misawa, K. Kuma and T. Miyata. 2002. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Research 30(14): 3059-3066. DOI: https://doi.org/10.1093/nar/gkf436
Katoh, K. and D. M. Standley. 2013. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Molecular Biology and Evolution 30(4): 772-780. DOI: https://doi.org/10.1093/molbev/mst010
Katoh, K., J. Rozewicki and K. D. Yamada. 2017. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Briefings in Bioinformatics 20(4): 1160-1166. DOI: https://doi.org/10.1093/bib/bbx108
Kornerup, A. and J. H. Wanscher. 1978. Methuen handbook of colour. 3a ed. Eyre Methuen. London, UK. 252 pp.
Lanfear, R., B. Calcott , D. Kainer, C. Mayer and A. Stamatakis. 2014. Selecting optimal partitioning schemes for phylogenomic datasets. BMC Evolutionary Biology 14: 82. DOI: https://doi.org/10.1186/1471-2148-14-82
Lanfear, R., P. B. Frandsen, A. M. Wright, T. Senfeld and B. Calcott . 2016. Partition Finder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Molecular Biology and Evolution 34(3): 772-773. DOI: https://doi.org/10.1093/molbev/msw260
Maddison, W. P. and D. R. Maddison. 2021. Mesquite: a modular system for evolutionary analysis.Version 3.70. http://mesquiteproject.org (consulted October, 2021)
Martínez-González, C. R., R. Ramírez-Mendoza, J. Jiménez-Ramírez, C. Gallegos-Vázquez and I. Luna-Vega. 2017. Improved method for genomic DNA extraction for Opuntia Mill. (Cactaceae). Plant Methods 13: 1-10. DOI: https://doi.org/10.1186/s13007-017-0234-y
Müller, K., D. Quandt, J. Müller and C. Neinhuis. 2005. PhyDE®-Phylogenetic data editor. Program distributed by the authors, version 10.0.https://www.phyde.de (consulted October, 2021)
Noguera, F. A., J. H. Vega-Rivera, A. N. García-Aldrete and M. Quesada-Avendaño (eds.). 2002. Historia Natural de Chamela. Instituto de Biología, Universidad Nacional Autónoma de México. Cd. Mx., México. 568 pp.
Pérez-Silva, E., T. Herrera andM. Esqueda. 1999. Species of Geastrum (Basidiomycotina: Geastraceae) from Mexico. Revista Mexicana de Micología 15: 89-104.
Ponce de León, P. 1968. A revision of the family Geastraceae. Fieldiana: Botany 31(14): 303-349.
Ramírez-López, I., M. Villegas-Ríos y Z. Cano-Santana. 2012. Diversidad de Agaricomycetes clavarioides en la Estación de Biología de Chamela, Jalisco, México. Revista Mexicana de Biodiversidad 83(4): 1084-1095. DOI: http://doi.org/10.7550/rmb.27700
Raymundo, T. 2021. Fuscoporia valenzuelae (Hymenochaetaceae, Basidiomycota), a new species from the tropical dry forest in Mexico. Acta Botanica Mexicana 128: e1844. https://doi.org/10.21829/abm128.2021.1844
Raymundo, T., M. L. Coronado , A. Gutiérrez, M. Esqueda andR. Valenzuela. 2017. New records of Ascomycota from tropical dry forest in Sonora, Mexico. Mycotaxon 132(2): 421-432. DOI: https://doi.org/10.5248/132.421
Raymundo, T. , E. Escudero-Leyva, I. Ortega-López, D. Castro-Bustos, H. León-Avendaño and R. Valenzuela . 2014. Ascomicetos del bosque tropical caducifolio en el Parque nacional Lagunas de Chacahua, Oaxaca, México. Boletín de la Sociedad Micológica de Madrid 38: 9-21.
Raymundo, T., R. Valenzuela and M. Esqueda . 2009. The family Hymenochaetaceae from México 4: New records from Sierra de Álamos-Río Cuchujaqui biosphere reserve. Mycotaxon 110: 387-398.
Reyes, P. E., R. Valenzuela andT. Raymundo. 2020. El género Hypoxylon (Xylariales, Ascomycota) en el bosque tropical caducifolio de cuatro Áreas Naturales Protegidas de México. Acta Botanica Mexicana 127: e1680. DOI: https://10.21829/abm127.2020.1680
Rzedowski, J. 2006. Vegetación de México. 1ra. Edición digital, Comisión Nacional para el Conocimiento y Uso de la Biodiversidad (CONABIO). Cd. Mx., México. 504 pp.
Salinas-Salgado, E., R. Valenzuela , T. Raymundo , M. Cipriano-Salazar, B. Cruz-Lagunas and E. Hernández-Castro. 2012. Macromicetos xilófagos del bosque tropical caducifolio en el municipio de Cocula, Guerrero, México. Polibótanica 34: 137-155.
Suchard, M. A., P. Lemey, G. Baele, D. L. Ayres, A. J. Drummond and A. Rambaut . 2018. Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10 Virus Evolution 4(1): vey016. DOI: http://doi.org/10.1093/ve/vey016
Sunhede, S. 1989. Geastraceae (Basidiomycotina). Morphology, ecology, and systematics with special emphasis on the north european species. Synopsis Fungorum 1: 534.
Trierveiler-Pereira, L., F. D. Calonge and I. G. Baseia. 2011. New distributional data on Geastrum (Geastraceae, Basidiomycota) from Brazil. Acta Botanica Brasilica 25(3): 577-585. DOI: http://doi.org/10.1590/S0102-33062011000300010
Valenzuela, R., T. Raymundo , C. Decock andM. Esqueda . 2012. Aphyllophoroid fungi from Sonora, México 2: New records from Sierra of Álamos-Río Cuchujaqui Biosphere Reserve. Mycotaxon 122: 51-59. DOI: https://doi.org/10.5248/122.51
White, T. J., T. D. Bruns, S. B. Lee and J. W. Taylor. 1990. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis, M. A., D. H. Gelfand, J. J. Sninsky and T. J. White (eds.). PCR Protocols: a guide to methods and applications. Academic Press, New York. 135-322. http://dx.doi.org/10.1016/B978-0-12-372180-8.50042-1
Wu G., B. Feng, J. Xu., X. T. Zhu, Y. C. Li, N. K. Zeng, M. I. Hosen and Z. L. Yang. 2014. Molecular phylogenetic analyses redefine seven major clades and reveal 25 new generic lineages in the fungal family Boletaceae. Fungal Diversity 69: 93-115.
Zamora, J. C., K. Hosaka and M. P. Martín. 2014. Systematics of the genus Geastrum (Fungi: Basidiomycota) revisited. Taxon 63(3): 477-497. DOI: https://doi.org/10.12705/633.36
Zhang, Z., S. Schwartz, L. Wagner and W. Miller. 2000. A greedy algorithm for aligning DNA sequences. Journal of Computational Biology 7(1-2): 203-214. DOI: https://doi.org/10.1089/10665270050081478
Author contributions
Author notes
4Author for correspondence: rvalenzg@ipn.mx
