Artículos de investigación
Systematics of Urostephanus: resurrection of a Mesoamerican taxon of Gonolobinae (Apocynaceae, Asclepiadoideae), with eight new combinations
Sistemática de Urostephanus: resurrección de un taxón Mesoamericano de Gonolobinae (Apocynaceae, Asclepiadoideae), con ocho nuevas combinaciones
Systematics of Urostephanus: resurrection of a Mesoamerican taxon of Gonolobinae (Apocynaceae, Asclepiadoideae), with eight new combinations
Acta botánica mexicana, no. 131, e2302, 2024
Instituto de Ecología A.C., Centro Regional del Bajío
Received: 05 January 2024
Revised document received: 08 March 2024
Accepted: 03 April 2024
Published: 28 May 2024
Abstract
Background and Aims: Recent studies have focused on resolving the systematics of the tribe Gonolobinae of the family Apocynaceae, as well as the controversial genus Matelea s.l. However, many species within Matelea s.l. have not been evaluated phylogenetically. This is the case of M. gonoloboides, previously recognized as the type species of Urostephanus, and other taxa with similar floral morphology. The aim of this study is to test the monophyly of Urostephanus, including some species with morphology similar to M. gonoloboides. Furthermore, we evaluate the taxonomic position of the Urostephanus clade within the subtribe Gonolobinae and compare morphology among related clades.
Methods: To assess phylogenetic relationships within Gonolobinae, we downloaded sequences from four loci for 94 species of the subtribe from GenBank. We extracted DNA and sequenced the trnL-F intergenic spacer and rps16 intron from four species of Dictyanthus and six species of Matelea, which are considered to belong to Urostephanus. We reconstructed the phylogeny with Bayesian inference using the maximum clade credibility tree. Based on the phylogenetic pattern, we performed morphological comparisons between the clades Dictyanthus, Polystemma, and Urostephanus.
Key results: Phylogenetic analysis recovered Urostephanus as monophyletic and sister to Dictyanthus. Based on this phylogenetic pattern and floral morphology, we resurrect the genus Urostephanus. We propose eight new combinations for the species included in the phylogeny, as well as for those that share floral morphological similarity. We designate two lectotypes. Finally, we discuss aspects of morphology between Urostephanus and closely related groups.
Conclusions: The current circumscription of Matelea s.l. includes species with high morphological diversity. With the phylogenetic and morphological evidence among related taxa, we support the resurrection of Urostephanus, more than 120 years after its description. These results contribute to the resolution of the systematics of Gonolobinae in Mesoamerica.
Key words: Diversity+ flower morphology+ gynostegial corona+ Mexico+ molecular phylogenetics+ Sanger sequencing.
Resumen
Antecedentes y Objetivos: Estudios recientes se han centrado en resolver la sistemática de la tribu Gonolobinae, así como del controversial género Matelea s.l. Sin embargo, varias especies dentro de Matelea s.l. no han sido evaluadas. Tal es el caso de Matelea gonoloboides, previamente reconocida como especie tipo de Urostephanus, y otras especies con una morfología floral similar no han sido evaluadas filogenéticamente. Los objetivos de este estudio son probar la monofilia del grupo Urostephanus, incluyendo algunos taxones con morfología similar a M. gonoloboides. Además, evaluamos la posición taxonómica del grupo Urostephanus dentro de la subtribu Gonolobinae y comparamos la morfología entre clados afines.
Métodos: Para evaluar las relaciones filogenéticas dentro de Gonolobinae, descargamos de GenBank las secuencias de cuatro loci para 94 especies de la subtribu. Extrajimos ADN y secuenciamos trnL-F y rps16 de cuatro especies de Dictyanthus y seis especies de Matelea, que se consideran en el grupo Urostephanus. Reconstruimos la filogenia con inferencia Bayesiana, empleando el árbol de máxima credibilidad de los clados. Con base en el patrón filogenético, realizamos una comparación morfológica entre los clados Dictyanthus, Polystemma y Urostephanus.
Resultados clave: El análisis filogenético recupera las especies del grupo Urostephanus como monofiléticas y relacionadas con Dictyanthus. Con base en este patrón filogenético y la morfología floral, resucitamos al género Urostephanus. Proponemos ocho nuevas combinaciones para las especies incluidas en la filogenia, así como para aquellas que comparten similitud morfológica floral. Designamos dos lectotipos. Además, discutimos aspectos sobre la morfología entre Urostephanus y grupos cercanamente relacionados.
Conclusiones: La circunscripción actual de Matelea incluye especies con una gran diversidad morfológica. Con la evidencia filogenética y morfológica entre taxones relacionados apoyamos la resurrección de Urostephanus, más de 120 años después de su descripción. Estos resultados contribuyen a resolver la sistemática de Gonolobinae en Mesoamérica.
Palabras clave: Corona ginostegial, diversidad, filogenia molecular, México, morfología floral, secuenciación Sanger.
Introduction
Gonolobinae is a monophyletic group of Apocynaceae (Asclepiadoideae, Asclepiadeae), dominated by climbing plants with ca. 45 genera and over 540 species with a primarily amphitropical distribution (Endress et al., 2018). However, the generic limits within the subtribe have not been fully established (Krings et al., 2008; McDonnell et al., 2018; Endress et al., 2018). Several studies tried to test the generic circumscriptions and to infer the synapomorphies supporting them (Rapini et al., 2003, 2006; Liede-Schumann et al., 2005; Krings et al., 2008; Morillo, 2015; McDonnell et al., 2018). However, there is still considerable uncertainty regarding the circumscription of several genera, particularly within Matelea Aubl. in the broad sense of Woodson (1941), henceforth Matelea s.l.
Matelea s.l. stands out as the genus with the most remarkable morphological heterogeneity within Gonolobinae (Krings et al., 2008; Endress et al., 2018).Woodson (1941) subordinated 27 genera of Asclepiadoideae to 16 subgenera of Matelea, comprising over 200 species with very diverse and complex floral characters, and retained Gonolobus Michx. and Fischeria DC. as distinct genera (Stevens and Morales, 2009). Phylogenetic studies have recovered several type species, of potentially different genera, nested within Matelea s.l. (Krings et al., 2008). Recent studies recognize Matelea as a paraphyletic group with several different genera within Matelea s.l. (McDonnell et al., 2018). Some works have resurrected genera such as Ibatia Decne., Lachnostoma Kunth (sensu Morillo, 2012), Polystemma Decne. (sensu Stevens, 2001; Stevens and Morales, 2009), Ptycanthera Decne. (Morillo, 2023b), and Trichosacme Zucc. (Alvarado-Cárdenas and Cortez, 2022), or established new genera (Morillo, 2012, 2013, 2015, 2016, 2023a, b; Morillo et al., 2013, 2016; Mangelsdorff et al., 2016; Morillo and Keller, 2016; Keller and Liede-Schumann, 2020; Alvarado-Cárdenas, 2021). It has been suggested that Matelea s.s. (sensuKrings and Morillo, 2015; Morillo, 2015; Morillo et al., 2017, 2023) only contains around 20 species, morphologically consistent with the type species M. palustris Aubl. (Stevens, 2001; Krings and Morillo, 2015). However, generic circumscriptions are far from settled as several recent resurrections provided limited, if any, molecular evidence.
In recent coalescence analyses obtained with multilocus chloroplast and nuclear data with ASTRAL-II, several monophyletic groups within Gonolobinae were recovered (McDonnell et al., 2018), morphologically contrasted to Matelea s.s., such as the clade M. gonoloboides (B.L. Rob. & Greenm.) Woodson sister to D. pavonii Decne. The same results were obtained using more species and only two cpDNA loci (rpl16 and rpoB-trnC), which include the clade M. gonoloboides-M. patalensis (Donn. Sm.) Woodson, sister to four species of Dictyanthus Decne. (McDonnell et al., 2018). These results are consistent with those previously obtained by Krings et al. (2008) with two cpDNA loci (rps16 and trnL-F). As a further contribution to resolving taxonomic issues in the subtribe, our work focuses on M. gonoloboides, initially described as the type for Urostephanus B.L. Rob. & Greenm., and morphologically similar species (henceforth Urostephanus) (Stevens and Morales, 2009; González-Martínez et al., 2023).
Urostephanus was established more than 120 years ago, based on Urostephanus gonoloboides B.L. Rob. & Greenm. (Robinson and Greenman, 1895) from Mexico. The type species was described as having a rotate corolla, arachnoid trichomes on the adaxial corolla, a cyathiform corona adnate to the base of the column, the distal edge of the corona with five interstaminal bifid and filiform lobes (horn-like), and with five oblong-spatulate staminal corona lobes resting on the anthers. Woodson (1941) included it within Matelea subgenus Ibatia (Decne.) Woodson, among 18 other morphologically diverse species. However, Krings et al. (2008) recognized Matelea subgenus Ibatia as an artificial group. Stevens and Morales (2009) pointed out that M. gonoloboides is related to Matelea araneosa (Donn. Sm.) Woodson, M. medusae Woodson, M. patalensis, M. porphyrantha (Standl.) Woodson, and M. warscewiczii (H. Karst.) Woodson. Only M. patalensis, M. porphyrantha, and M. medusae share a cyathiform corona with staminal and interstaminal corona elements; the latter has five bifid and erect lobes, whereas the interstaminal corona of M. araneosa and M. patalensis is made up only of a basal projection, which resembles a stinger denticulated at the base. Four new species were recently described from the specimens previously identified as M. gonoloboides (González-Martínez et al., 2023). This broadens the specific and morphological diversity and the number of species that can potentially be included in the Urostephanus group due to shared characters of taxonomic importance, such as the fusion of the lobes of the interstaminal corona and appendages on the membrane.
The objectives of the present study are to test the monophyly of the Urostephanus group, by including some species that share the floral attributes of the type species U. gonoloboides, to know the phylogenetic relationship of the Urostephanus group with respect to the other taxa of the subtribe Gonolobinae, and to compare the main morphological attributes among the related clades or taxa.
Material and Methods
Taxon sampling for the phylogenetic study
The coalescent analyses obtained by McDonnell et al. (2018) are consistent in recovering Matelea gonoloboides as sister to a Dictyanthus clade (clade A, according to McDonnell et al., 2018). In addition, Polystemma (clade D) is the sister group of clade A. Hereafter we will name them Dictyanthus clade, Polystemma clade, and Urostephanus clade (the species related to M. gonoloboides). To evaluate the phylogenetic relationships of the species in the Urostephanus group and potentially related taxa, we downloaded the sequences available in GenBank (2022) of four plastid DNA regions: rpl16 intron, rps16 intron, trnC-rpoB intergenic spacer, and trnL-F intergenic spacer from Gonolobinae. The sequences were generated by Krings et al. (2008) and McDonnell et al. (2018), also including those of recently proposed genera supported by molecular data such as Anemotrochus Mangelsdorff, Meve & Liede and Tylodontia Griseb. (Mangelsdorff et al., 2016) (Supplementary data 1).
We did not include the sequences of Dictyanthus species, nor previously published Matelea gonoloboides or M. patalensis. When we included these data with those we also sampled, our study produced inconsistencies in phylogenetic relationships (data not shown). We selected Orthosia scoparia (Nutt.) Liede & Meve, Metastelma sp. and Metastelma northropiae Schltr. as the outgroups, as representatives of the subtribes Orthosiinae and Metastelmatinae, respectively, successive sister clades of Gonolobinae, according to Fishbein et al. (2018).
Extraction and amplification of DNA, alignment, partition scheme, and phylogenetic inference
We extracted genomic DNA from dry leaves (20-25 mg) from herbarium specimens of Matelea balrog Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado, M. cornuta Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado, M. gonoloboides, M. jaimesiae Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado, M. lokii Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado, and M. porphyrantha. Furthermore, to broaden the sampling of the putative sister group of Urostephanus, we included four species of Dictyanthus: D. macvaughianus (W.D. Stevens) W.D. Stevens, D. parviflorus Hemsl., D. pavonii Decne., and D. reticulatus (Turcz.) Benth. & Hook. F. ex Hemsl. (Supplementary data 1).
DNA extraction was performed with DNeasy Plant Mini Kits (QIAGEN, Germany), following the manufacturer’s protocol. To be consistent with previous works (Krings et al., 2008; McDonnell et al., 2018), we amplified two plastid fragments using GoTaq DNA Polymerase (Promega, USA): trnL-F (Taberlet et al., 1991) and rps16 (Lee and Downie, 2000), and rpl16 only for M. gonoloboides. These regions have been shown to provide phylogenetic information at a generic and species level (Liede-Schumann et al., 2005; Krings et al., 2008).
We made four matrices for each of the molecular regions: rpl16, rps16, trnC-rpoB, trnL-F (Table 1). The matrices include the type species of putative genera: Anemotrochus, Chthamalia Decne., Dictyanthus, Fischeria, Ibatia, Matelea s. s., Meveampelos Morillo, Polystemma, Prosthecidiscus Donn. Sm., Ptycanthera, and Suberogerens Morillo. We updated the nomenclature of the species included in the phylogeny, according to recent publications, to represent the currently accepted taxa (Supplementary Data 1).
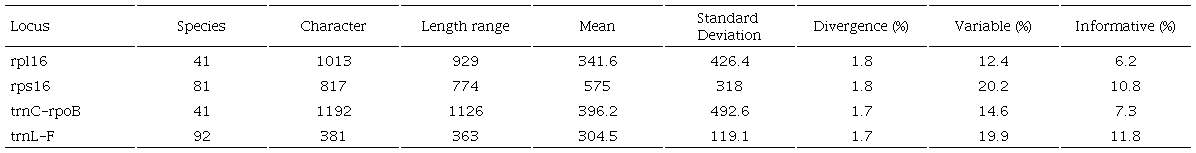
We aligned each locus separately using an online version of MAFFT v. 7 (Katoh et al., 2002, 2017; Katoh and Standley, 2013). After that, we examined the alignments, and manually edited them using AliView v. 1.26 software (Larsson, 2014) (Supplementary data 2). The four matrices were concatenated in a single matrix with 104 taxa and 3403 characters (3529 characters including indels) using Mesquite v. 3.2 software (Maddison and Maddison, 2017). We selected a partition scheme for each of the four molecular regions. We evaluated nucleotide substitution models for each of the molecular partitions using the Bayesian Information Criterion (BIC) in PartitionFinder v. 2.1.1 on XSEDE (Lanfear et al., 2017), implemented on the CIPRES webserver (Miller et al., 2010). The TVM+I was the best model for each molecular region. The TVM model is not currently available in MrBayes. We implemented the next best-fitting parameter, the GTR with a proportion of invariant sites (I) and gamma-distributed (G) (nst=6 with I+G).
We inferred the phylogeny with Bayesian phylogenetic inference using MrBayes on XSEDE v. 3.2.6 (Huelsenbeck and Ronquist, 2001) on the CIPRES webserver (Miller et al., 2010). We used four Markov chains Monte Carlo (three heated chains and one cold chain), with default temperature values with two independent runs to assess convergence on a stationary distribution. We sampled a total of 50 million generations, sampling one tree every 1000 trees (mencionado en el texto 3). The phylogenetic inference included calculating the Bayesian posterior probabilities support values for each node (PP). We assessed the convergence and stationarity of chains with effective size sample >200 using Tracer v. 1.6.0 software (Rambaut et al., 2014). We obtained the phylogram of maximum clade credibility tree discarding 2500 trees as a burnin (25%, according to trace plot) with mean heights and recovered with TreeAnotator v. 1.8.4 software (Bouckaert et al., 2014).
Additionally, as the highest percentage of species with complete molecular data are found in the rps16 and trnL-F, we performed an exploratory analysis considering only 77 species from the Gonolobinae subtribe and one from Metastelmatinae, to assess consistency in topology and changes in posterior probability values. We used the same parameters as described above for alignment, evolutionary models, and phylogenetic analyses (see alignment in Supplementary Data 4).
Morphology comparison
We recorded the states for vegetative, floral (inflorescence type, corolla, staminal and interstaminal corona, gynostegium), and fruit characters consulting the taxonomic literature. We performed a morphological comparison of the groups related to Urostephanus (Robinson and Greenman, 1895; González-Martínez et al., 2023), including Dictyanthus (Stevens, 1988; Endress et al., 2018; González-Martínez et al., 2019), and Polystemma (Stevens, 2001; Lozada Pérez, 2010; McDonnell and Fishbein, 2016; McDonnell et al., 2018; Endress et al., 2018). We complemented the morphological comparison with observations made directly in the field and of floral samples under the stereoscopic microscope (Leica, EZ4 HD, Solms, Germany) to clarify the organization of the corona in the different genera. We followed the classification of Liede and Kunze (1993), and Kunze (1995) for the description of the gynostegial corona, where Ci=interstaminal corona and Cs=staminal corona.
In this work, we regard taxa (genus and species) as explanatory hypotheses that attempt to describe and explain biological diversity. These hypotheses are constructed based on abductive inference, which requires a background knowledge (causal theories and hypotheses), together with our observations and knowledge of causal conditions (Fitzhugh, 2005; 2009), along with assumptions about evolutionary history (Griffiths, 1996). Based on the concept of genus proposed by Stuessy (1990) and phylogenetic theory, we suggest that a genus is a species, or a group of species (explanatory hypotheses) characterized by attributes (morphological and genetic), originated by some unspecified mechanism(s) during tokogeny, and fixed in the descendants of individuals with those characters, that make it different and separated by some unspecified mechanism(s) from any other species or group of species (monophyletic groups). Our species hypothesis follows the cohesion species concept (Templeton, 1989); taxa are suggested based on their phenotypic, ecological, and genetic cohesion. The molecular data of the individuals used in the phylogenetic analyses are associated with the species hypotheses generated in previous studies (González-Martínez et al., 2023).
Geographic distribution
We made distribution maps for the clades to represent the distribution patterns. Geographic distribution records were obtained from a previous review of M. balrog, M. cornutus, M. jaimesiae, and M. lokii specimens (González-Martínez et al., 2023), except for M. araneosa, M. medusae, and M. patalensis, which were obtained from public databases available at GBIF (2023). For Dictyanthus species (17/19 species), Polystemma (5/8 species, excluding the Matelea species nested within the clade), geographical records were obtained from GBIF (2023). We filtered the records of GBIF to include only data with preserved specimen coordinates (download links in Supplementary data 5). We excluded duplicate records. Maps were produced in QGIS software v. 3.32.3 (QGIS Development Team, 2019).
Taxonomic treatment
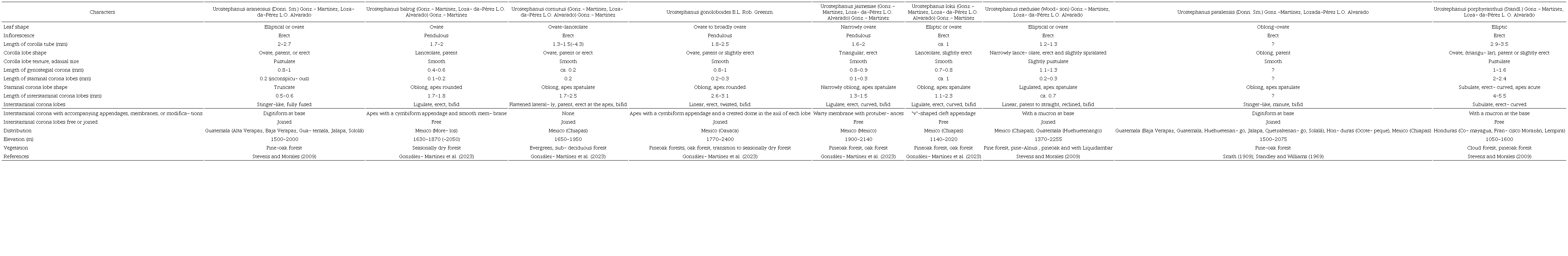
We update the generic description of Urostephanus (Robinson and Greenman, 1895) to reflect the morphological diversity of the species Matelea araneosa (Smith, 1909; Standley and Williams, 1969; Stevens and Morales, 2009), M. balrog, M. cornutus, M. jaimesiae, M. lokii (González-Martínez et al., 2023), M. medusae (Standley and Steyermark, 1944; Stevens and Morales, 2009), M. patalensis (Smith, 1909; Standley and Williams, 1969), and M. porphyrantha (Standley, 1938). We consulted the collections of the following herbaria: ENCB, IBUG, IEB, MEXU, and UAMIZ (Thiers, 2022-updated continuously), as well as the digital collections of the herbaria MO, NY, and US (Thiers, 2022-updated continuously) to update the description. Although not all species were included in the phylogeny, the morphological similarity in gynostegial corona attributes allows us to recognize them as members of the same genus.
Additionally, we generated a comparative table between the species we include within Urostephanus, using morphological, geographical, and ecological attributes of taxonomic importance. For detailed information on the organization of floral morphology, we suggest consulting the illustrations of González-Martínez et al. (2023).
Results
Phylogenetic relationships of Gonolobinae
Our phylogenetic pattern is, in general terms, consistent with previous phylogenetic analyses (Krings et al., 2008; Mangelsdorff et al., 2016; McDonnell et al., 2018; Keller and Liede-Schumann, 2020). However, the posterior probability of the nodes is lower at deep levels in the phylogeny and at tips level resolution due to the incomplete nature of the sequences or the phylogenetic informativeness at those levels of the loci employed (Fig. 1). Similar values are obtained when using an analysis with a smaller species sampling and with only two complete loci for all species (Supplementary data 6). We recovered in clades the species now recognized as genera distinct from Matelea, which is indirectly supported by morphological attributes or synapomorphies. The maximum clade credibility tree from Bayesian inference recovers two main clades. In the first clade (PP:0.27), we recovered the following taxa as monophyletic: Chthamalia (PP:1), Dictyanthus (PP:1), Microstelma and Prosthecidiscus (PP:0.94), Pherotrichis (PP:1), Suberogerens and Talayotea (PP:0.99), Urostephanus (PP:1); and Polystemma as paraphyletic (PP:0.94). The second clade (PP:0.23) comprised Chloropetalum (PP:0.35), Gonolobus (PP:0.86), Ibatia (PP:0.86), Ptycanthera (PP:0.12), Schubertia (PP:0.98), and Tylodontia (PP:0.97) clades as monophyletic, and Fischeria as paraphyletic. We recovered the genus Anemotrochus in another clade (PP:0.96), with their sister clade Meveampelos (PP:0.86). We recovered Matelea s.l. as a paraphyletic genus; the type species Matelea palustris is sister to M. pedalis (E. Fourn.) Fontella & E.A. Schwarz in the second clade (PP:0.82) (Fig. 1). There are some species of Matelea s.l. that form monophyletic groups that can potentially represent different genera due to their morphological similarities, as occurs with the Matelea reticulata clade.
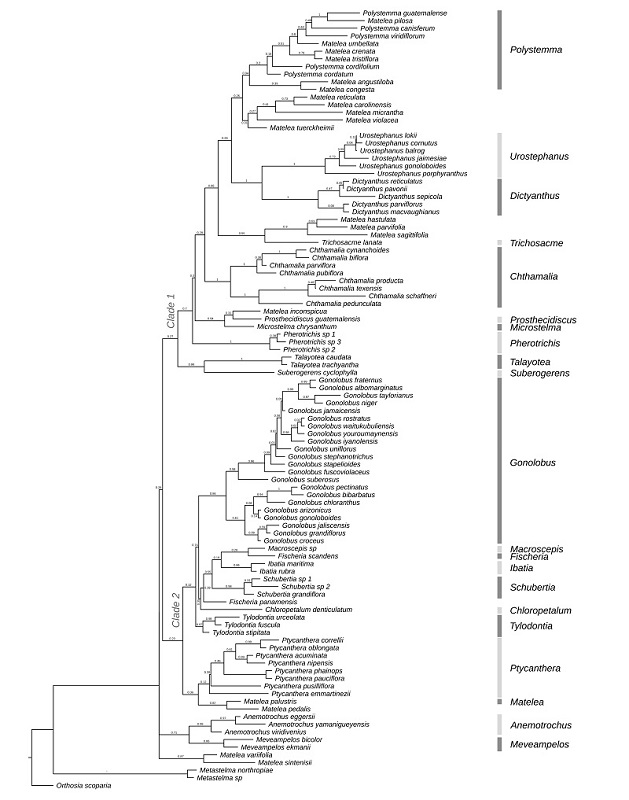
Figure 1:
Maximum clade credibility phylogeny showing the phylogenetic relationship of Urostephanus B.L. Rob. & Greenm. within Gonolobinae indicating the position of the genus in the phylogeny. The numbers of the clades indicate the main clades recovered. The numbers indicate the support of branches, measured by Bayesian Posterior Probability.
Phylogenetic relationships of Urostephanus and related clades
Based on our alignment and phylogenetic pattern, the Urostephanus group is recovered in a clade that has several putative molecular synapomorphies, in both the trnL-F intergenic spacer and rps16 intron regions of the cpDNA. In rps16, character 733 is thymine in Urostephanus species instead of guanine; in character 744, adenine vs. cytosine; in character 800, guanine vs. thymine. In the trnL-F intergenic spacer data set, the first character is thymine vs. cytosine; in character 163, thymine vs. cytosine; in character 209, a cytosine vs. adenine; and the character 257, a thymine vs. cytosine (see alignments in Supplementary data 2).
We recovered Urostephanus with Dictyanthus as sister group (PP:1), and the clade of Polystemma s.l. and the Matelea reticulata clade (PP:0.26) as the sister group of the Urostephanus-Dictyanthus clade (PP:0.28) (Fig. 1). Each of these groups was recovered as monophyletic, supported by morphological attributes or potential synapomorphies and restricted to geographic areas.
The Urostephanus clade (PP=1) included the type species Urostephanus gonoloboides (Matelea gonoloboides sensuWoodson, 1941), as well as U. balrog (Matelea balrog), U. cornutus (M. cornuta), U. jaimesiae (M. jaimesiae), U. lokii (M. lokii), and U. porphyranthus (Matelea porphyrantha sensuWoodson, 1941).
The combinations in Urostephanus are presented as new below. The members of the Urostephanus group share a cyathiform corona with bifid interstaminal corona (McDonnell et al., 2018). The species mentioned are restricted to the Mesoamerican region in Mexico, Guatemala, and Honduras, with high diversity in southern Mexico (Fig. 2A). Morphological and molecular similarities suggest the nomenclatural change of Matelea species included in the clade to Urostephanus.
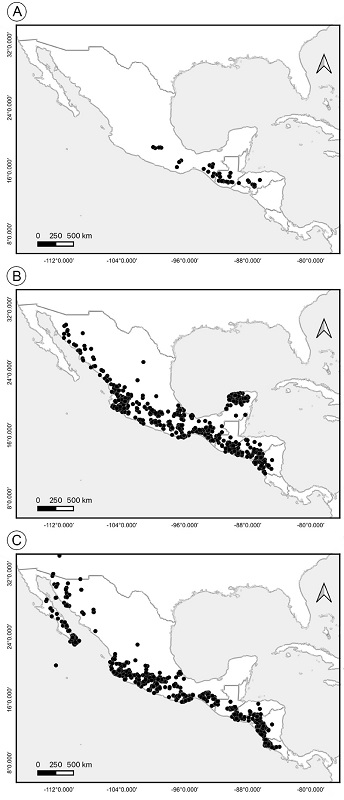
Figure 2:
Geographic distribution pattern of species of the clades related to Urostephanus B.L. Rob. & Greenm.; A. distribution of Urostephanus; B. distribution of Dictyanthus Decne.; C. distribution of Polystemma Decne.
Dictyanthus was recovered as monophyletic (PP=1), represented by D. pavonii (type species), D. macvaughianus, D. parviflorus, D. reticulatus, and D. sepicola. Uncinate trichomes and urceolate to campanulate sacciform corollas with confluent coronas are shared by these species and could be putative synapomorphies (McDonnell et al., 2018). The members of Dictyanthus are distributed from northern Mexico to Nicaragua, with the highest diversity and endemism in Mexico (Stevens, 2000; González-Martínez et al., 2019; Alvarado-Cárdenas et al., 2020, 2022) (Fig. 2B).
Also, we recovered the Polystemma clade (PP=0.94) as paraphyletic, represented by P. viridiflorum Decne. (type species), P. canisferum McDonnell & Fishbein, P. cordifolium (A. Gray) McDonnell & Fishbein, P. guatemalense (Schltr.) W.D. Stevens, and several species still included in Matelea (sensuWoodson, 1941). Under the current circumscription, the genus is recognized by having trichomes with crystalline inclusions, corona with multiple teeth and mottled fusiform fruits (Stevens and Morales, 2009; McDonnell et al., 2018). They are distributed from the southwest USA to Nicaragua. (Fig. 2C).
Finally, the Matelea reticulata clade, with M. carolinensis (Jacq.) Woodson, M. micrantha L.O. Williams, M. reticulata (Engelm. ex. A. Gray) Woodson, M. tuerckheimii (Donn. Sm.) Woodson, and M. violacea Woodson. We could not determine synapomorphies for this clade but noted that species share rotate and reticulate flowers, and muricate fruits, except M. carolinensis.
Morphology of Urostephanus and related clades
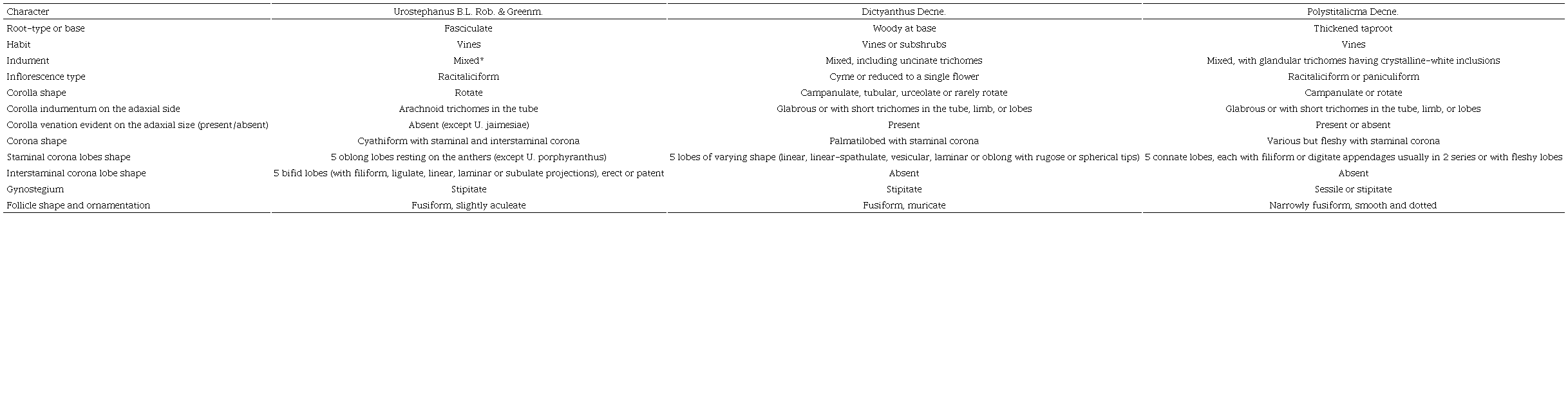
According to our phylogenetic pattern and correlated with the morphology, Urostephanus is easily distinguished from Dictyanthus and Polystemma by numerous floral and follicle attributes (Table 2). In the Urostephanus group, the flowers have rotate corollas with arachnoid trichomes in the corolla tube without evident venation (except M. jaimesiae). These features contrast with Dictyanthus, which generally has campanulate corollas that are glabrous (or with short trichomes in the tube) and evident venation. In the case of Polystemma, the flowers are rotate to campanulate, without arachnoid trichomes, and several of its species have reticulate corollas. The gynostegial corona is one of the main attributes to diagnose genera and species. In Urostephanus, the corona is cyathiform, made up of the staminal corona (Cs) and interstaminal corona (Ci), while Dictyanthus and Polystemma only have Cs. Furthermore, in the fruits, we also find morphological differences, mainly in shape, the presence of projections, and their color. The fruits of the Urostephanus group are slightly aculeate and uniform in color; in Dictyanthus, they are muricate and with dark mottling, while in Polystemma they are smooth and dotted or with irregular stripes (Fig. 3; Table 2).

Figure 3:
Floral and fruit morphology of representative species of Urostephanus B.L. Rob. & Greenm., Dictyanthus Decne., and Polystemma Decne. A. Urostephanus gonoloboides B.L. Rob. & Greenm., flower, bar=1 cm; B. U. gonoloboides, gynostegium and corona, bar=0.1 cm; C. Dictyanthus pavonii Decne., flower, bar=1.5 cm; D. D. pavonii, gynostegium and corona, bar=0.1 cm; E. Polystemma viridiflorum Decne., flower, bar=1.5 cm; F. Polystemma mirandae Lozada-Pérez, gynostegium and corona, bar=0.5 cm; G. from left to right, fruit of U. porphyranthus (Standl.) Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado (image of Katya Romero Soler), D. asper (Mill.) W.D. Stevens, and P. guatemalense (Schltr.) W.D. Stevens, bar=1 cm. Cg=cyathiform gynostegial corona; Ci=interstaminal corona; Cs=staminal corona; G=gynostegium.
Urostephanus spans a conspicuous range of floral morphological variation. The corolla is rotate, yellow-greenish, brown-purple or reddish (color may be variable within species). The tube has arachnoid pubescence or is nearly glabrous. With regard to shape, the lobes are triangular, ovate or lanceolate, patent or erect (Fig. 3). The gynostegium is pentagonal with notches between the anthers. The lobes of the Cs are subulate or oblong, resting on the anthers (subulate, erect-curved in M. porphyrantha). The lobes of the Ci are free or joined at the base, erect or patent, and bifid, filiform, ligulate, linear, laminar, subulate or stinger-like; the bases of the lobes have membranes that can be corrugated, with warty protuberances, or smooth (see Appendix). Based on the phylogenetic pattern and morphological and molecular characters, which differ from the other taxa, and according to our explanatory hypothesis, we propose the recognition of Urostephanus as an independent genus (we include here a description) and provide the corresponding combinations of the species names.
Taxonomic treatment
Urostephanus B. L. Rob. & Greenm., Amer. J. Sci., ser. 3, 50: 159. 1895.
TYPE: Urostephanus gonoloboides B.L. Rob. & Greenm.
≡ Matelea gonoloboides (B.L. Rob. & Greenm.) Woodson
Vines herbaceous or woody with suberous base; roots fleshy and fasciculate; stems with mixed indumentum of short glandular trichomes, black, and long straight trichomes, white or yellowish; leaves opposite, petiolate, blades ovate to elliptic, base cordate or lobate with colleters, apex attenuate to acuminate; inflorescences congested-racemiform, extra-axillary, one per node; bracts lanceolate; calyx with colleters inside the lobes; corolla rotate, yellow-greenish, brown-purple or reddish; tube with arachnoid pubescence or nearly glabrous; lobes triangular, ovate or lanceolate, aestivation contort dextrorse; gynostegium stipitate, staminal appendices terminal and adpressed at the margin of the apex of the style; pollinia horizontal, distally sterile and excavated at the base; style apex pentagonal, convex or flattened, slightly emarginate between the anthers; gynostegial corona cyathiform, adnate basally to gynostegium, orange or purple to black; staminal corona (Cs) with 5 lobes, subulate or oblong, apex truncate or rounded, resting on the anthers (or 5 lobules, subulate, erect-curved in Urostephanus porphyranthus), emerging internal to the middle part of the gynostegial corona; interstaminal corona (Ci) with denticulate, cymbiform, or emarginate appendages at the apex, 5 lobes bifid, filiform, ligulate, linear, laminar, subulate or stinger-like, erect or patent, free or joined at the base, emerging external to the middle part of the gynostegial corona; the lobes are accompanied at the base by corrugated, warty, or smooth membranes; follicle fusiform, slightly aculeate; seeds not seen.
Distribution and ecology: species of Urostephanus are distributed from central Mexico to northern Honduras, in mountainous regions. The species of this genus grow in cloud forest, pine-oak forest, oak forest, and rarely in seasonally dry tropical forest.
Taxonomic notes: Urostephanus is an available name for this genus based on the type of U. gonoloboides (Robinson and Greenman, 1895). We apply the principle of priority to restore the generic name (see article 11 of the ICN, Turland et al., 2018). Urostephanus comes from Latin “uro-” which refers to the elongated or tail-like appendage, and “stephanos”, which is the Greek word for describing a surrounding structure like a corona or garland (Stearn, 2006). The etymology refers to the shape of the gynostegial corona and its different lobes and appendices.
1. Urostephanus araneosus (Donn. Sm.) Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado, comb. nov.
TYPE: GUATEMALA. Alta Verapaz, montibus inter Tactic & Cobán, 1650 m, VII.1908, H. von Türckheim II 2332 (lectotype: US00111104! (designated here); isolectotype: US00111105!).
≡ Gonolobus araneosus Donn. Sm., Bot. Gaz. 47: 257. 1909.
≡ Matelea araneosa (Donn. Sm.) Woodson, Ann. Missouri Bot. Gard. 28: 223. 1941.
Taxonomic notes: in the original description (Smith, 1909) the specimen Hans von Türckheim II 2332 was assigned as type. However, there are two duplicates in the US herbarium. We designate specimen US00111104 as lectotype because it contains an original label from the herbarium of John Donnell Smith. It is also of higher taxonomic value because it contains more flowers.
2. Urostephanus balrog (Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado) Gonz.-Martínez, comb. nov.
TYPE: MEXICO. Morelos, municipio Cuernavaca, Buenavista del Monte a 2.5 km de la entrada a los viveros de Otongo, 1871 m, 7.X.2009, A. Flores Morales 151 (holotype: MEXU1482690!).
≡ Matelea balrog Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado, Kew Bull. 78: 522. 2023.
3. Urostephanus cornutus (Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado) Gonz.-Martínez, comb. nov.
TYPE: MEXICO. Chiapas, municipio Siltepec, Cascada, 5.VIII.1937, E. Matuda 1613 (holotype: MEXU88547!).
≡ Matelea cornuta Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado, Kew Bull. 78: 523. 2023.
4. Urostephanus gonoloboides B.L. Rob. & Greenm. Amer. J. Sci., ser. 3, 50: 159. 1895.
TYPE: MEXICO. Oaxaca, hills above Oaxaca, 6.VIII.1894, C. G. Pringle 4753 (lectotype: GH00076876!, designated by González-Martínez et al., 2023).
≡ Matelea gonoloboides (B.L. Rob. & Greenm.) Woodson, Ann. Missouri Bot. Gard. 28: 222. 1941.
Taxonomic notes: several specimens have been identified as Matelea gonoloboides (= Urostephanus gonoloboides) because of their apparent similarity in floral morphology with the type and the lack of adequate delimitation of the species (Standley and Williams, 1969; Stevens and Morales, 2009). The taxonomic descriptions made so far do not include more precise descriptions of the attributes of the corona, such as the fusion of the Ci lobes, as well as the appendages or membranes that may be present, which are consistent in the populations, as well as in the specimens studied and therefore of taxonomic importance.
Urostephanus gonoloboides differs from the rest of the species in having Ci with a cymbiform appendage and a crested dome in the axil of each lobe, as well as bifid, linear, erect, twisted lobes, joined at the base. Urostephanus gonoloboides is similar to U. jaimesiae. However, it differs from it by having Cs with oblong lobes with rounded apices (vs. narrowly oblong lobes with spatulate apices), Ci with a crested dome in the axil of each lobe (vs. warty membranes with protuberances), and twisted linear lobes (vs. curved ligulate lobes) that are joined at the base (vs. free at the base).
Matelea gonoloboides was described in the Flora of Guatemala (Standley and Williams, 1969), and also included in Flora Mesoamericana (Stevens and Morales, 2009). However, the taxonomic description of Standley and Williams (1969) mentions, among other characteristics, that the corolla lobes are linear to narrowly deltoid-lanceolate, the corona has five internal, hornlike lobes (corresponding to Cs) and five external lobes with two filiform flexuous tails (corresponding to Ci), and a depressed style apex. These characters differ markedly from U. gonoloboides, mainly because of the staminal corona shape that resembles the corona morphology of the U. porphyranthus. More evaluation of Guatemalan specimens could reveal that it is a distinct species. A detailed reassessment and study of the populations previously recognized as M. gonoloboides are necessary for proper identification.
5. Urostephanus jaimesiae (Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado) Gonz.-Martínez comb. nov.
TYPE: MEXICO. Estado de México, municipio San Simón de Guerrero, cerca de Ranchería Cucha alrededor del panteón, 1900 m, 20.VIII.2005, V. Juárez J. et al. 762 (holotype: MEXU1179513!).
≡ Matelea jaimesiae Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado, Kew Bull. 78: 525. 2023.
6. Urostephanus lokii (Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado) Gonz.-Martínez, comb. nov.
TYPE: MEXICO. Chiapas, municipio Villacorzo, ejido Sierra Morena al O del poblado, 1140 m, 6.VIII.2002, L. Alvarado C. et al. 341 (holotype: MEXU1043130!).
≡ Matelea lokii Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado, Kew Bull. 78: 526. 2023.
7. Urostephanus medusae (Woodson) Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado, comb. nov.
TYPE: GUATEMALA, Huehuetenango, rocky slopes above La Libertad, on Cerro Pueblo Viejo, 20.VIII.1942, J. A. Steyermark 50987 (holotype: MO078255!; isotype: F0048964F!).
≡ Matelea medusae Woodson, Publ. Field Mus. Nat. Hist., Bot. Ser. 23: 78. 1944.
8. Urostephanus patalensis (Donn. Sm.) Gonz.-Martínez, Lozada-Pérez & L.O. Alvarado, comb. nov.
TYPE: GUATEMALA. Baja Verapaz, Patal, 1600 m, VII.1908, H. von Türckheim II 2371 (lectotype: US00111124! (designated here); isolectotype: US00147749!).
≡ Gonolobus patalensis Donn. Sm., Bot. Gaz. 47(4): 256-257. 1909.
≡ Matelea patalensis (Donn. Sm.) Woodson, Ann. Missouri Bot. Gard. 28: 223. 1941.
Taxonomic notes: from the original description (Smith, 1909) until now, there is no formal lectotype designation in any of the treatments where the species has been addressed (e.g. Standley and Williams, 1969; Stevens, 2001). From the two type specimens (Hans von Türckheim II 2371) stored in the US herbarium, we designate as lectotype the specimen bearing the original herbarium label of John Donnell Smith. This designation is in accordance with the information given on the Tropicos website (Tropicos, 2024) for the basionym.
9. Urostephanus porphyranthus (Standl.) Gonz.-Martínez, Lozada-Pérez & L. O. Alvarado, comb. nov.
TYPE: HONDURAS. Comayagua, Siguatepeque, near river, 5.VII.1936, T. G. Yuncker et al. 5676 (holotype: F0049063F!; isotype: MO344389!, NY00318839!, S1213183!).
≡ Vincetoxicum porphyranthum Standl., Publ. Field Mus. Nat. Hist., Bot. Ser. 17: 387. 1938.
≡ Matelea porphyrantha (Standl.) Woodson, Ann. Missouri Bot. Gard. 28: 223. 1941.
Discussion
Phylogenetic analyses
The Urostephanus group was recovered as a monophyletic and well-supported clade. Its members (here transferred) share genetic and morphological traits of the flower and gynostegial corona, distinct from other genera of Gonolobinae. Recognizing Urostephanus as a genus is consistent with our explanatory hypothesis, with each clade being composed of a set of hypothesized species that share attributes that are distinct from other clades with different attributes, separated by some unspecified mechanism.
Our results are similar to previous phylogenetic analyses, recovering Dictyanthus (sensuStevens, 2000) as the sister group of the species recognized here in Urostephanus; furthermore, the Polystemma clade (sensuStevens, 2001) is recognized as closely related to this clade (Krings et al., 2008; McDonnell et al., 2018). Although previous phylogenetic contributions have not aimed at elucidating the phylogenetic relationships of these groups, the phylogenetic hypotheses obtained from a large sample of Gonolobinae have consistently recovered these clades. It seems likely that a broader sampling of species would have recovered the same topology with regard to these clades, given the similarity of our results with previous analyses, even when including new sequences together with published sequences (e.g. Mangelsdorff et al., 2016).
Despite obtaining a topology partially consistent with previous studies (Mangelsdorff et al., 2016; McDonnell et al., 2018), in our analyses, the support or posterior probability values at nodes were relatively low at deep levels of the phylogeny, as well as close to the tips. This is likely due to incomplete sampling of loci for all species included in the phylogeny, as well as the phylogenetic informativeness and incongruence provided by the loci to resolve at different phylogenetic levels. Poor resolution and incongruence at deep levels of the phylogeny have been mentioned to reflect rapid evolution of lineages along with incomplete lineage sorting (McDonnell et al., 2018), during their divergence in the middle Miocene (Rapini et al., 2007). Although most genera included in the analysis are recovered as monophyletic and with reliable posterior probability, we consider taking with caution the resolution at deep levels of the phylogeny of our analyses.
Considerations in Urostephanus
Urostephanus is a monophyletic group according to our molecular analysis, a finding congruent with morphological attributes shared among U. araneosus, U. balrog, U. cornutus, U. gonoloboides, U. jaimesiae, U. lokii, U. medusae, U. patalensis, and U. porphyranthus. The main diagnostic of Urosteophanus is the cyathiform gynostegial corona with Cs and Ci lobes, and the Ci with five bifid lobes. In U. balrog, U. gonoloboides, U. jaimesiae, and U. lokii, the Ci is bifid, linear or ligulate, and erect, but these species can be recognized by differences in the length of the lobes of Ci, the degree of fusion, the ornamentations, and appendages at the bases of the Ci lobes (González-Martínez et al., 2023). In the case of U. araneosus, U. cornutus, U. medusae, U. patalensis, and U. porphyranthus, they have markedly different Ci morphology from each other. Urostephanus araneosus has Ci lobes stinger-like and completely fused with denticulated bases (Smith, 1909), whereas these are bifid, laminar, and laterally flattened in U. cornutus (González-Martínez et al., 2023). Urostephanus medusae showed Ci lobes bifid, linear, patent with a mucron at the base (González-Martínez et al., 2023); in U. patalensis, they are stinger-like and diminutively bifid with denticulated bases (Smith, 1909); and in U. porphyranthus, the Ci lobes are bifid and subulate (Stevens and Morales, 2009).
Other morphological differences are found in the shape and dimensions of the corolla. Intraspecific variation is relatively low and (mainly in corolla color), with species having relatively marked morphological cohesion, based on our review of herbarium specimens. The difficulties for the delimitation of the species in the group have arisen mainly because there are few specimens in herbaria and few or no vegetative details to distinguish them, in addition to the challenges of working with pressed flowers. Therefore, detailed observations of flower structures are necessary. Additionally, the few specimens collected each of the species we examined and their low representativeness in herbaria made it difficult to obtain samples for phylogenetic analysis, so not all the species studied could be included in the analyses.
Before our analysis, Stevens and Morales (2009) pointed out that several species, which we include in the genus, form a complex of species that require attention for their delimitation. On the one hand, he recognized that Matelea araneosa (here U. araneosus), M. medusae (here U. medusae), and M. patalensis (here U. patalensis) (henceforth referred to as group 1) can only be distinguished by the shape and size of the corolla and could possibly be a widely variable species. On the other hand, he further added that these species, together with M. gonoloboides (here U. gonoloboides), M. warscewiczii and M. porphyranta (here U. porphyranthus) (henceforth referred to as group 2), form a species complex (Stevens and Morales, 2009). Our view is that group 1 cannot be recognized as a single widely variable species because it has readily diagnostic corolla and corona features that distinguish it from all other putative relatives. Although U. araneosus, U. medusae, and U. patalensis share arachnoid trichomes in the corolla tube and cyathiform gynostegial coronas, other aspects of the morphology of the gynostegial corona differ notably. Urostephanus araneosa has ovate corolla lobes (4.8 cm long on average; Stevens and Morales, 2009), with inconspicuous Cs, and the Ci with lobes with digitate bases and fully fused projections short and stinger-like curved upwards (Smith, 1909; reinterpreted from Field Museum, 2017), while U. patalensis has lanceolate corolla lobes (4.5 cm long on average, Stevens and Morales, 2009), Cs with oblong lobes resting on the anthers, and Ci has lobes with digitiform bases and a short, upward-curved projection (Smith, 1909; reinterpreted from detailed flowers observations of specimen Stevens 1249, MEXU). In contrast, U. medusae differs markedly from the previous species in having narrowly lanceolate lobes (19.5 mm long on average, Stevens and Morales, 2009), four times longer than wide; the Cs has oblong lobes resting on the anthers, and the Ci has bifid lobes that are linear and patent. In group 2, U. gonoloboides and U. porphyrantha share the diagnostic features of the genus, but differ in shape of the Ci lobes, which are linear and subulate, respectively. However, M. warscewiczii differs significantly, not only from these species but also from group 1. The corolla is subrotate-campanulate, and the Cs and Ci are very complex with multiple ascending projections (for more details, consult the original illustrated description in Karsten, 1869). Based on this, we decided not to include it within Urostephanus.
It is pertinent to point out that Stevens and Morales (2009) cited the specimen Stevens & Martínez 25699 (MEXU, MO) as M. patalensis, which was sampled in our analyses and those of McDonnell et al. (2018); however, the specimen differs markedly from the original description (Smith, 1909). In the specimen Stevens & Martínez 25699, the morphology of the corona is akin to Urostephanus, and it is treated as an isotype of Matelea cornuta (González-Martínez et al., 2023). We corroborated the original description of M. patalensis (Smith, 1909) with the observation of the digital image of the type specimens (Tuerckheim 2371, US00111124!), and with the detailed observation of the flower of the specimens Stevens 1249 (MEXU) which was also included by Stevens and Morales (2009). Urostephanus cornutus and U. patalensis resemble each other by having cyathiform gynostegial coronas with lobes of the Cs oblong resting on the anthers; however, they differ in the morphology of the Ci. The lobes of the Ci are laminar, patent and distally bifid with ascendant projections in U. cornutus vs. lobes of the Ci with digitate bases and a short, upward-curved projection in M. patalensis (according to Smith, 1909).
The Gonolobinae are diverse and difficulties in the circumscription of the genera have been mainly due to the complexity of the structures that make up the gynostegial corona (e.g., Woodson, 1941; Krings et al., 2008). Our results highlight the need for studies to understand the morphology of the flowers, gynostegial corona, and fruits; likewise, the understanding of the morphology of our species hypothesis will improve our phylogenetic analyses to resolve the relationships of the taxa, as well as provide causal information about the process involved in the evolution of the subtribe.
Acknowledgements
To the two anonymous reviewers for their valuable contributions and comments to improve the manuscript. To Ericka Belén Cortez for providing samples of flowers fixed in ethanol of one of the species. To César Ramiro Martínez González for his support, assistance and suggestions during the molecular work for this study. To María Eugenia Muñiz Díaz de León for her support and for allowing the use of the facilities at the Taller de Biología de Plantas I y II of the Facultad de Ciencias, UNAM, where the molecular work was carried out and stereomicroscope photographs were taken. We are grateful to the curators and technicians of the herbarium MEXU for their great help in obtaining samples of the species studied. We thank Laura Márquez and Nelly López, who performed the DNA sequencing in the Laboratorio Nacional de la Biodiversidad. We thank Mark Fishbein for his substantial comments and recommendations that significantly improved the quality of the article.
Literature cited
Alvarado-Cárdenas, L. O. 2021. Talayotea (Apocynaceae: Asclepiadoideae, Gonolobinae), un género nuevo y endémico de México. Phytoneuron 62: 1-18.
Alvarado-Cárdenas, L. O. and E. B. Cortez. 2022. Pasado y presente de Trichosacme lanata (Apocynaceae: Asclepiadoideae): Revisión de un taxón endémico y poco conocido en México. Phytoneuron 33: 1-10.
Alvarado-Cárdenas, L. O. , A. López Hernández and R. Sánchez Crisóstomo. 2022. A remarkable new species of Dictyanthus (Apocynaceae, Asclepiadoideae, Asclepiadeae, Gonolobinae) restricted to Jalisco, México. Phytotaxa 558(2): 178-184. DOI: https://doi.org/10.11646/phytotaxa.558.2.2
Alvarado-Cárdenas, L. O. , L. Lozada-Pérez, C. S. Islas-Hernández, E. B. Cortez, K. G. Maya-Mandujano and M. G. Chávez-Hernández. 2020. Apocináceas de ayer y hoy. Conocimiento histórico y reevaluación de la diversidad y distribución de Apocynaceae en México. Botanical Sciences 98(2): 393-416. DOI: https://doi.org/10.17129/botsci.2525
Bouckaert, R., J. Heled, D. Kühnert, T. Vaughan, C. H. Wu, D. Xie, M. A. Suchard, A. Rambaut and A. J. Drummond. 2014. BEAST 2: A software platform for Bayesian Evolutionary Analysis. PloS Computational Biology 10(4): e1003537. DOI: https://doi.org/10.1371/journal.pcbi.1003537
Endress, M. E., U. Meve, D. J Middleton and S. Liede-Schumann. 2018. Apiales, Gentianales (except Rubiaceae). In: Kadereit, J. W. and V. Bittrich (eds.). Flowering plants: Eudicots. The Families and Genera of Vascular Plants, vol. XV. Springer. Cham, Switzerland. DOI: https://doi.org/10.1007/978-3-319-93605-5
Field Museum. 2017. Matelea araneosa. Neotropical Live Plant Photos. https://plantidtools.fieldmuseum.org/en/nlp/catalogue/3684652 (consulted, November 2018).
Fishbein, M., T. Livshultz, S. C. K. Straub, A. O. Simões, J. Boutte, A. McDonnell and A. Foote. 2018. Evolution on the backbone: Apocynaceae phylogenomics and new perspectives on growth forms, flowers, and fruits. American Journal of Botany 105(3): 495-513. DOI: https://doi.org/10.1002/ajb2.1067
Fitzhugh, K. 2005. The inferential basis of species hypotheses: the solution to defining the term ‘species’. Marine Ecology 26(3-4): 155-165. DOI: https://doi.org/10.1111/j.1439-0485.2005.00058.x
Fitzhugh, K. 2009. Species as explanatory hypotheses: refinements and implications. Acta Biotheoretica 57: 201-248. DOI: https://doi.org/10.1007/s10441-009-9071-3
GBIF. 2023. Global Biodiversity Information Facility. https://www.gbif.org (consulted, September 2023).
GenBank. 2022. National Center for Biotechnology Information National Center for Biotechnology Information https://www.ncbi.nlm.nih.gov/genbank/ (consulted September, 2022).
González-Martínez, C. A., L. Lozada-Pérez and L. O. Alvarado-Cárdenas. 2019. Dictyanthus stevensii (Apocynaceae; Asclepiadoideae; Gonolobinae), a new species from Chimalapas region, Oaxaca, Mexico. Phytotaxa 394(1): 79-88. DOI: https://doi.org/10.11646/phytotaxa.394.1.5
González-Martínez, C. A., L. Lozada-Pérez and L. O. Alvarado-Cárdenas. 2023. Four new species of Matelea (Apocynaceae: Asclepiadoideae) endemic to Mexico. Kew Bulletin 78(4): 519-531. DOI: https://doi.org/10.1007/s12225-023-10121-z
Griffiths, P. E. 1996. The historical turn in the study of adaptation. British Journal for the Philosophy of Science 47(4): 511-532. DOI: https://doi.org/10.1093/bjps/47.4.511
Huelsenbeck, J. P. and F. Ronquist. 2001. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17(8): 754-755. DOI: https://doi.org/10.1093/bioinformatics/17.8.754
Karsten, H. 1869. Callaeolepium warscewiczii Krst. Florae Columbiae terraumque adiacentium specimina selecta in peregrinatione duodecim annorum observata. Apud Ferdinand Duemmler Successores. Berlin, Alemania. Pp. 123-124. DOI: https://doi.org/10.5962/bhl.title.400
Katoh, K. and D. M. Standley. 2013. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Molecular Biology and Evolution 30(4): 772-780. DOI: https://doi.org/10.1093/molbev/mst010
Katoh, K., K. Misawa, K. Kuma and T. Miyata. 2002. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Research 30(14): 3059-3066. DOI: https://doi.org/10.1093/nar/gkf436
Katoh, K., J. Rozewicki and K. D. Yamada. 2017. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Briefings in Bioinformatics 20(4): 1160-1166. DOI: https://doi.org/10.1093/bib/bbx108
Keller, H. A. and S. Liede-Schumann. 2020. Caa, a new South American genus of Gonolobinae (Apocynaceae: Asclepiadeae: Asclepiadoideae). Lilloa 57(1): 81-88. DOI: https://doi.org/10.30550/j.lil/2020.57.1/6
Krings, A. and G. Morillo. 2015. A new species in the Matelea palustris complex (Apocynaceae, Asclepiadeae) and a synopsis of the complex of the Guianas and Northern Brazil. Systematic Botany 40(1): 214-219. DOI: https://doi.org/10.1600/036364415X686521
Krings, A., D. T. Thomas and Q. Xiang. 2008. On the generic circumscription of Gonolobus (Apocynaceae, Asclepiadoideae): evidence from molecules and morphology. Systematic Botany 33(2): 403-415. DOI: https://doi.org/10.1600/036364408784571527
Kunze, H. 1995. Floral morphology of some Gonolobeae (Asclepiadaceae). Botanische Jahrbücher fur Systematik, Pflanzengeschichte und Pflanzengeographie 117(1): 211-238.
Lanfear, R., P. B. Frandsen, A. M. Wright, T. Senfeld and B. Calcott. 2017. PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Molecular Biology and Evolution 34(3): 772-773. DOI: https://doi.org/10.1093/molbev/msw260
Larsson, A. 2014. AliView: a fast and lightweight alignment viewer and editor for large datasets. Bioinformatics 30(22): 3276-3278. DOI: https://doi.org/10.1093/bioinformatics/btu531
Lee, B. Y. and S. R. Downie. 2000. Phylogenetic analysis of cpDNA restriction sites and rps16 intron sequences reveals relationships among Apiaceae tribes Caucalideae, Scandiceae and related taxa. Plant Systematics and Evolution 221: 35-60. DOI: https://doi.org/10.1007/BF01086379
Liede, S. and H. Kunze. 1993. A descriptive system for corona analysis in Asclepiadaceae and Periplocaceae. Plant Systematics and Evolution 185: 275-284. DOI: https://doi.org/10.1007/BF00937663
Liede-Schumann, S., A. Rapini, D. J. Goyder and M. W. Chase. 2005. Phylogenetics of the New World subtribes of Asclepiadeae (Apocynaceae-Asclepiadoideae): Metastelmatinae, Oxypetalinae and Gonolobinae. Systematic Botany 30(1): 184-195. DOI: https://doi.org/10.1600/0363644053661832
Lozada Pérez, L. 2010. Polystemma mirandae (Apocynaceae, Asclepiadoideae), una nueva especie de México. Novon 20(4): 429-431. DOI: https://doi.org/10.3417/2009004
Maddison, W. P. and D. R. Maddison. 2017. Mesquite: a modular system for evolutionary analysis. Ver. 3.31. http://mesquiteproject.org (consulted, January 2018).
Mangelsdorff, R. D., U. Meve and S. Liede-Schumann. 2016. Phylogeny and circumscription of Antillean Anemotrochus, gen. nov., and Tylodontia (Apocynaceae: Asclepiadoideae: Gonolobinae). Willdenowia 46(3): 443-474. DOI: https://doi.org/10.3372/wi.46.46311
McDonnell, A. and M. Fishbein. 2016. Polystemma canisferum (Apocynaceae, Asclepiadoideae): a distinctive new gonoloboid milkweed vine from Sonora, Mexico. Phytotaxa 246(1): 78-84. DOI: https://doi.org/10.11646/phytotaxa.246.1.6
McDonnell, A., M. Parks and M. Fishbein. 2018. Multilocus phylogenetics of New World milkweed vines (Apocynaceae, Asclepiadoideae, Gonolobinae). Systematic Botany 43(1): 77-96. DOI: https://doi.org/10.1600/036364418X697021
Miller, M. A., W. Pfeiffer and T. Schwartz. 2010. Creating the CIPRES Science Gateway for Inference of Large Phylogenetic Trees. Proceedings of the Gateway Computing Environments Workshop (GCE): 1-8. DOI: https://doi.org/10.1109/GCE.2010.5676129
Morillo, G. 2012. Aportes al conocimiento de las Gonolobinae (Apocynaceae-Asclepiadoideae). Pittieria 36: 13-57.
Morillo, G. 2013. Aportes al conocimiento de las Gonolobinae II (Apocynaceae, Asclepiadoideae). Pittieria 37: 115-154.
Morillo, G. 2015. Aportes al conocimiento de las Gonolobinae parte III (Apocynaceae, Asclepiadoideae). Pittieria 39: 191-258.
Morillo, G. 2016. Nuevas especies y nuevas combinaciones en las Gonolobinae (Apocynaceae, Asclepiadoideae, Asclepiadeae) de Centro y Sudamérica. Lilloa 53(1): 89-106. http://www.scielo.org.ar/scielo.php?script=sci_arttext&pid=S2346-96412016000100006 (consulted, November 2019).
Morillo, G. 2023a. Aportes al conocimiento de las Gonolobinae (Apocynaceae-Asclepiadoideae) parte IV. Memoria de la Fundación La Salle de Ciencias Naturales 81(190): 45-90.
Morillo, G. 2023b. Aportes al conocimiento de las Gonolobinae (Apocynaceae, Asclepiadoideae) Parte V. Chthamalia, Meveampelos gen. nov., Phaeostemma, Polystemma, Ptycanthera. Memoria de la Fundación La Salle de Ciencias Naturales 81(191): 37-90.
Morillo, G. and H. A. Keller. 2016. Un nuevo género y dos nuevas combinaciones en las Gonolobinae (Apocynaceae: Asclepiadoideae). Bonplandia 25(2): 129-143. https://www.jstor.org/stable/26413083 (consulted, November 2019).
Morillo, G., J. Fontella Pereira and M. V. DóriaBarbosa. 2013. Austrochthamalia (Apocynaceae, Asclepiadoideae, Gonolobinae), un género nuevo segregado de Chthamalia Decne. Revista de Biología Neotropical 10(1): 1-8. DOI: https://doi.org/10.5216/rbn.v10i1.27827
Morillo, G. , S. A. Cáceres Moral and H. A. Keller. 2016. Cristobalia, un nuevo género Sudamericano de Gonolobinae (Apocynaceae, Asclepiadoideae, Asclepiadeae). Pittieria 40(1): 122-143.
Morillo, G., I. L. de Morais and M. A. Farinaccio. 2017. Matelea atrolingua, una nueva Apocynaceae cuyos lóbulos corolinos semejan la lengua de un mamífero muerto. Iheringia Série Botânica 72(3): 319-324. DOI: https://doi.org/10.21826/2446-8231201772301
Morillo, G., U. Meve and S. Liede-Schumann. 2023. A new species of Matelea (Apocynaceae, Gonolobinae) from Bolivia and an assessment of Matelea in Bolivia. Novon 31(1): 126-131. DOI: https://doi.org/10.3417/2023815
QGIS Development Team. 2019. QGIS Geographic Information System. Open Source Geospatial Foundation Project. http://qgis.osgeo.org (consulted, May 2019).
Rambaut, A., M. A. Suchard, D. Xie and A. J. Drummond. 2014. Tracer ver. 1.6. http://beast.bio.ed.ac.uk/Tracer (consulted, May 2018).
Rapini, A., M. W. Chase, D. J. Goyder and J. Griffiths. 2003. Asclepiadeae classification: evaluating the phylogenetic relationships of the New World Asclepiadoideae (Apocynaceae). Taxon 52(1): 33-50. DOI: https://doi.org/10.2307/3647436
Rapini, A., M. W. Chase and T. U. P. Konno. 2006. Phylogenetics of South American Asclepiadoideae (Apocynaceae). Taxon 55(1): 119-124. DOI: https://doi.org/10.2307/25065533
Rapini, A. , C. van den Berg and S. Liede-Schumann. 2007. Diversification of Asclepiadoideae (Apocynaceae) in the New World. Annals of the Missouri Botanical Garden 94(2): 407-422. DOI: https://doi.org/10.3417/0026-6493(2007)94[407:DOAAIT]2.0.CO;2
Robinson, B. L. and J. M. Greenman. 1895. New and Noteworthy Plants chiefly from Oaxaca collected by Messrs. C. G. Pringle, L. C. Smith and E. W. Nelson. Contributions from the Gray Herbarium of Harvard University 9: 150-168. https://www.jstor.org/stable/41763753 (consulted, November 2019).
Smith, J. D. 1909. Undescribed plants from Guatemala and other Central American republics. XXXI. Botanical Gazette 47(4): 253-262. https://www.journals.uchicago.edu/doi/pdf/10.1086/329872 (consulted, December 2023).
Standley, P. C. 1938. Vincetoxicum Walt. Field Museum of Natural History 17(4): 387-388.
Standley, P. C. and J. A. Steyermark. 1944. Studies of Central American Plants-IV. Field Museum of Natural History 23(2): 31-109.
Standley, P. C. and L. O. Williams. 1969. Asclepiadaceae. In: Standley, P. C. and L. O. Williams (ed.). Flora of Guatemala, part VIII, number 4. Fieldiana Botany 24: 407-472.
Stearn, W. T. 2006. Latín Botánico. Ediciones Omega. Barcelona, España. Pp. 644.
Stevens, W. D. 1988. A synopsis of Matelea subg. Dictyanthus (Apocynaceae: Asclepiadoideae). Annals of the Missouri Botanical Garden 75(4): 1533-1564. DOI: https://doi.org/10.2307/2399300
Stevens, W. D. 2000. New and interesting milkweeds (Apocynaceae, Asclepiadoideae). Novon 10(3): 242-256. DOI: https://doi.org/10.2307/3393108
Stevens, W. D. 2001. Flora de Nicaragua: Asclepiadaceae R. Br. http://legacy.tropicos.org/Name/42000279?projectid=7 (consulted, November 2019).
Stevens, W. D. and J. F. Morales 2009. Flora Mesoamericana: Apocynaceae Flora Mesoamericana: Apocynaceae http://legacy.tropicos.org/Name/42000278?projectid=3&langid=66 (consulted, November 2019).
Stuessy, T. F. 1990. Plant Taxonomy. The Systematic Evolution of Comparative Data. Columbia University Press. New York, USA. 515 pp.
Taberlet, P., L. Gielly, G. Pautou and J. Bouvet. 1991. Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Molecular Biology 17(5): 1105-1109. DOI: https://doi.org/10.1007/BF00037152
Templeton, A. R. 1989. The meaning of species and speciation: A genetic perspective. In: Otte, D. and J. A. Endler (eds.). Speciation and its consequences. Sinauer Associates. Sunderland, UK. Pp. 3-27.
Thiers, B. 2022-updated continuously. Index Herbariorum, a global directory of public herbaria and associated staff. New York Botanical Garden’s Virtual Herbarium. New York, USA. http://sweetgum.nybg.org/ih (consulted, March 2024).
Tropicos. 2024. Tropicos.org. Missouri Botanical Garden. https://tropicos.org. (consulted, March 2024).
Turland, N. J., J. H Wiersema, F. R. Barrie, W. Greuter, D. L. Hawksworth, P. S. Herendeen, S. Knapp, W.-H. Kusber, D.-Z. Li, K. Marhold, T. W. May, J. McNeill, A. M. Monro, J. Prado, M. J. Price and G. F. Smith (eds.). 2018. International Code of Nomenclature for Algae, Fungi, and Plants (Shenzhen Code) adopted by the Nineteenth International Botanical Congress Shenzhen, China, July 2017. Regnum Vegetabile 159. Koeltz Botanical Books. Glashütten, Germany. 254 pp. DOI: https://doi.org/10.12705/Code.2018
Woodson, R. E. Jr. 1941. The North American Asclepiadaceae I. Perspective of the genera. Annals of the Missouri Botanical Garden 28(2): 193-244. DOI: https://doi.org/10.2307/2394270
Supplementary data
Supplementary data 1: Accession numbers of the sequences of the Gonolobinae used in the analyses (Krings et al., 2008; Mangelsdorff et al., 2016; McDonnell et al., 2018).
Supplementary data 2: Documents with the alignment files for each of the loci in nexus format. They can be visualized with any freely available software such as AliView www.github.com/AliView (Larsson, 2014) or Mesquite http://mesquiteproject.org (Maddison and Maddison, 2017).
Supplementary data 3: Nexus file with the information to perform the Bayesian analysis using all loci in MrBayes software http://brahms.biology.rochester.edu/software.html (Huelsenbeck and Ronquist, 2001). The file is a plain text document and can be viewed with any text editing software.
Supplementary data 4: Nexus file with the information to perform the Bayesian analysis using only two loci (rps16 and trnL-F) in MrBayes software http://brahms.biology.rochester.edu/software.html (Huelsenbeck and Ronquist, 2001). The file is a plain text document and can be viewed with any text editing software.
Supplementary data 5: Databases used for the geographic distribution of Urostephanus B. L. Rob. & Greenm. and related clades. The links come fromthe GBIF database of 19 September 2023.
Supplementary data 6: Results of phylogenetic analysis with Bayesian inference using only two loci. The file contains the phylogenetic tree, and the table with the statistics of the loci.
To cite as:
Author contributions
Funding
Author notes
*Author for correspondence: leonardoac@ciencias.unam.mx
