ECUADORIAN POTATO LANDRACES: TRADITIONAL NAMES AND GENETIC IDENTITY
PAPAS NATIVAS ECUATORIANAS: NOMBRES TRADICIONALES E IDENTIDAD GENÉTICA
ECUADORIAN POTATO LANDRACES: TRADITIONAL NAMES AND GENETIC IDENTITY
Revista Fitotecnia Mexicana, vol. 40, núm. 4, pp. 481-489, 2017
Sociedad Mexicana de Fitogenética, A.C.
Recepción: 22 Noviembre 2016
Aprobación: 21 Mayo 2017
Resumen: Las papas nativas ecuatorianas (Solanum tuberosum L.) son recursos genéticos importantes, pero se han descrito pobremente. Con el fin de estudiar la diversidad genética de estas papas se aplicaron marcadores de Secuencias Simple Repetidas (SSR) en 152 papas nativas colectadas en tres áreas de alta diversidad: provincias de Carchi, Chimborazo y Loja. Estos marcadores SSR fueron previamente aplicados en el genotipeo de más de 800 variedades de papas europeas. El número de alelos y valores del Contenido Informativo del Polimorfismo (PIC) de los marcadores fueron comparables entre las europeas y nuestro estudio; sin embargo, el traslape de alelos fue pequeño. La relación entre nombres locales de las papas nativas y la identidad genética, basada en datos SSR, mostró que había algunas papas nativas con diferentes nombres que tenían perfiles moleculares idénticos. Dicha relación también mostró que papas nativas con nombres idénticos, pero obvias diferencias en la morfología del tubérculo, fueron casi siempre diferentes genéticamente. No hubo un agrupamiento claro del material colectado de acuerdo con las regiones investigadas, lo que sugiere un movimiento extensivo de semillas de papas a lo largo de Ecuador.
Palabras clave: Solanum tuberosum, diversidad genética, nombres, potatoes, variedades nativas.
Abstract: Ecuadorian potato landraces (Solanum tuberosum L.) are an important genetic resource, but they have been poorly described. Simple Sequence Repeats (SSR) markers were applied to 152 landraces to assess the genetic diversity of potatoes collected in three areas of high diversity: the Carchi, Chimborazo and Loja provinces. These SSR markers were previously used in the genotyping of more than 800 European potato varieties. The number of alleles and Polymorphism Information Content (PIC) of the markers found in this study were similar to those in European cultivars; however, the overlap in alleles was small. Based on SSR data, the relationship between local names of landraces and genetic identity showed several landraces with different names but identical molecular profiles. It also showed that landraces with identical names but obvious differences in tuber morphology were almost always genetically different. There was no clear grouping of material collected according to the regions under study that suggests extensive movement of seed potatoes all over Ecuador.
Keywords: Solanum tuberosum, genetic diversity, landraces, names, potato.
INTRODUCTION
Potatoes (Solanum tuberosum L.) are cultivated throughout the Andes, and the greatest diversity ranges from Southern Peru to Northern Bolivia (De Haan and Rodriguez, 2016; Hawkes, 1988). Potato diversity in the Andes includes different ploidy levels. In Peru, for example, farmers grow mixtures of diploid, triploid, tetraploid and pentaploid potatoes in a single field (De Haan et al., 2010; De Haan and Rodriguez, 2016). Ecuador has an important but poorly described resource of potato landraces that also includes multiple ploidy levels (Hawkes, 2004), although the extent to which potatoes with different ploidy levels are grown is still partially unknown. Three cultivated tuber-bearing Solanum tuberosum diploid, triploid and tetraploid Andigenum group are known to be present in Ecuador (Spooner 2013, Spooner et al., 2014).
Andean potatoes have rich nomenclature. Hawkes (1947) described the origin and meaning of South American Indian potato names. Most of the Ecuadorian potatoes had Spanish or native Quechua names, or a combination of both. Understanding variety naming by farmers can be important to understand the genetic diversity present in a crop (Nuijten and Almekinders, 2008). Quiros et al. (1990) and De Haan et al. (2010) studied the consistency between the folk naming system and genetic profiles of potatoes in Peru. Farmer identification and electrophoretic phenotypes were well correlated in the study of Quiros et al. (1990), but De Haan et al. (2010) found poor correlations using SSRs markers. Both studies reported possible under-estimation of genetic variation in farmer fields due to landraces with the same name representing different genetic profiles. The relationship between names and genetic profiles has not been studied previously for the cultivated potatoes of Ecuador.
Compared to other marker systems, microsatellites (SSRs) have proven to be very effective because they are co-dominant, reproducible, cost-effective, simple to use and highly polymorphic (McGregor et al., 2000; Milbourne et al., 1997). Studies on the genetic diversity of local varieties of potatoes using SSRs have been conducted in Argentina (Ispizúa et al., 2007), Tenerife, Spain (Barandalla et al., 2006), UK (Reid and Kerr, 2007), Canada (Fu et al., 2009), Russia (Ryzhova et al., 2010), and Chile (Muñoz et al., 2016). Furthermore, a set of nine SSRs have been used to differentiate more than 800 potato varieties from the European Union Common Catalogue (Reid et al., 2009).
For this research, passport data (including local names) of previous collections guided the new collections in three areas of high diversity in Ecuador: provinces Carchi, Chimborazo and Loja (Database of INIAP´s genebank). These areas differ not only in climatic and edaphic conditions but also in ethnicity. The objective was to use SSR markers to determine the relationship between local names and the genetic identity of the landraces and to describe the extent of the potato diversity found in these Ecuadorian provinces.
MATERIALS AND METHODS
Plant material
A total of 152 Ecuadorian potato landraces were collected, 39 in Carchi, located in northern Ecuador between 0º 27' to 1º 10' N, 65 in Chimborazo in central Ecuador from 1º 33' to 2º 55' S, and 48 in Loja, in the South between latitude 3º 18' and 4º 45' S (Figure 1). The altitudinal range of the potato collection varied in Carchi from 2950 to 3400 meters above sea level (masl), in Chimborazo from 2750 to 3950 masl, and in Loja from 2250 to 2900 masl. Individual farmers provided one or more landraces. Passport data included information about landrace name and origin. Ten well-known and commonly used Dutch varieties were included for comparison.
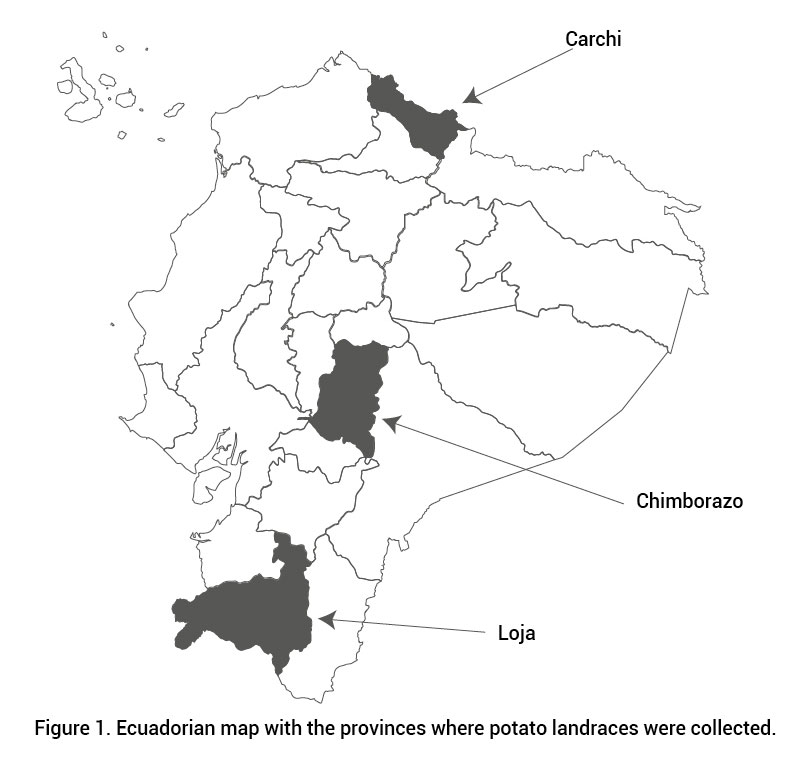
Figure 1
Figure 1. Ecuadorian map with the provinces where potato landraces were collected.
Ploidy level determination
In vitro plants were prepared by using a routine tissue culture protocol. One plant per landrace was used to determine ploidy level by flow cytometry using a Cyflow® Space, Partec, flow cytometer (Guo et al., 2010; INIAP-DENAREF, 2009). Reference varieties were: NK2-162 Yema de huevo CIP 704218 (CIP, 2010) for diploid potato. Ecuadorian varieties Fripapa and Natividad (INIAP, 2017) for tetraploids, and FMFHRA 005 Chihuila negra for triploid potato.
DNA extraction
DNA extraction was performed using the protocol described by Colombo et al. (1998) with the modifications introduced by Morillo and Miño (2011). In short, genomic DNA was isolated from young leaves or tuber sprouts depending on availability of the material (either greenhouse or field, respectively). The tissue was suspended in 200 µL of extraction buffer (50 mM Tris HCl pH 8.0, 1 M NaCl, 20 mM EDTA, 1 % PVP and 1 % CTAB); then, 800 µL of extraction buffer and 12 µL of β-mercapthoethanol were added, and the suspension was thoroughly mixed, incubated at 60 ºC for 2 h and centrifuged at 13000 rpm for 10 min (Eppendorf® centrifuge 5415 D, Hamburg, Germany). The supernatant was recovered and a mixture of 750 µL of chloroform:isoamyl alcohol (24:1) was added, mixed and centrifuged again at 13000 rpm for 5 min. The supernatant was transferred to a new tube and 750 µL of ethanol 100 % was added and incubated at -20 ºC for 10 min. The suspension was centrifuged at 13000 rpm for 3 min followed by a washing step with 70 % ethanol. The tubes were dried at room temperature overnight. If small drops were still observed in the tubes then they were dried again at 37 ºC for 30 min. DNA was re-suspended in 200 µL of TE buffer (10 mM Tris HCl, 1 mM EDTA pH 8.0), incubated at 65 ºC with 2 µL RNase (10 ng µL-1) per 100 µL of DNA solution). DNA was further purified using the PureLink™ 96 Genomic DNA Kit, Invitrogen®, as recommended by the manufacturer. Purified DNA was stored at -20 ºC in TE-buffer.
Microsatellite analysis
To characterize the plant material, nine SSR loci were evaluated (Reid et al., 2009). The nine markers were amplified in three multiplex PCR reactions each containing three markers, as described by Reid et al. (2009), with minor modifications. Instead of 30 cycles described in the protocol, 40 cycles for multiplex 1 and 2, and 35 cycles for multiplex 3 were used. PCR products were analyzed using a DNA sequencer (Applied Biosystems 3130 XL Genetic Analyzer, Foster City, California) with POP-7TM. The peaks present for each microsatellite were visualized using GeneMapper Software v 3.6 (Applied Biosystems) and scored using the protocol described by Reid et al. (2009).
Data analysis
SSR alleles were scored as binary data (present or absent, 1 and 0). A distance matrix was calculated using the Nei and Li coefficient (Nei and Li, 1979) and from this an Unweighted Pair Group Method with Arithmetic Mean (UPGMA) tree was obtained using Treecon® (Van de Peer and De Wachter, 1994). The population genetic structure among the three research areas (Carchi, Chimborazo and Loja) was compared with an Analysis of Molecular Variance (AMOVA) performed by Arlequin 3.11 (Excoffier et al., 2005). The Polymorphism Information Content (PIC) value was based on allelic phenotypes (Becher et al., 2000; Esselink et al., 2003) using the formula PIC = 1-S(pi)², where “pi” is the frequency of the i-th allelic phenotype detected (Nei, 1973).
RESULTS AND DISCUSSION
Naming of landraces
The structure of some landrace names in Chimborazo included Kichwa words, e.g. Kuchi chupa “pig tail”; Uchu rumi “stone to grind chili”; Ashco chaqui “dried dog”; Papa yerac “white potato” and Pudzu uvilla “grey berry” or a combination of Spanish-Kichwa names such as Yana pera “black pear”, Yana tabla “black long and flat tuber” or Cacho blanco, or Cuerno blanco. “Cacho” and “Cuerno” refer to the shape of the tuber like a bull horn and “blanco” means white. Kichwa names in Loja included Guano de kuchi “pig excrement” and Papa de chakra “small field potato” for example. Table 1 lists the collected landraces and includes Spanish (43 %), Spanish-Kichwa (26 %) and Kichwa (12 %) names; 19 % of the names were not classified due to uncertainty.
Early sprouting landraces are named mainly by the generic Chaucha meaning “soft or easy”. This was consistent across the three research areas. This generic name is followed by tuber related characteristics such as color of the tuber, e.g. Blanca “white”, Amarilla “yellow”, Roja “red” or Negra “black”, or animal related names, e.g. Borrega “sheep”, Ratona “mouse” or tuber shape, e.g. Botella “bottle”. In some cases, these two naming components are accompanied by a third component that is the tuber shape, e.g. Chaucha amarilla redonda (“redonda”: round shaped). Exceptions to the naming rule for these early sprouting potatoes are Tulca, Castillo, Wicupa, Mambera and Tabaquera.
Names of potato landraces may also refer to women’s names, such as Manuela and Catalina. Others allude to their apparent origin, such as Cañareja (from Cañar province) or Norteña (from the North) and Leona del Carchi (Lion shape tuber from Carchi). Others refer to animal related features, e.g. Coneja “rabbit ears shape” and Rabo de Gato “cat’s tail”. Other names refer to objects, e.g. Alpargata “children’s shoes”; or gender, as in Tulca hembra (“hembra”: female).
Ploidy levels
The ploidy level of only 134 landraces and their distribution over the three provinces was analyzed. The Ecuadorian landraces consisted of 22 % diploids, 6 % triploids and 72 % tetraploids (Figure 2). No triploids were present in Loja, and pentaploid potatoes were not found in this study. It was observed that farmers usually plant landraces with different ploidy level on the same fields.
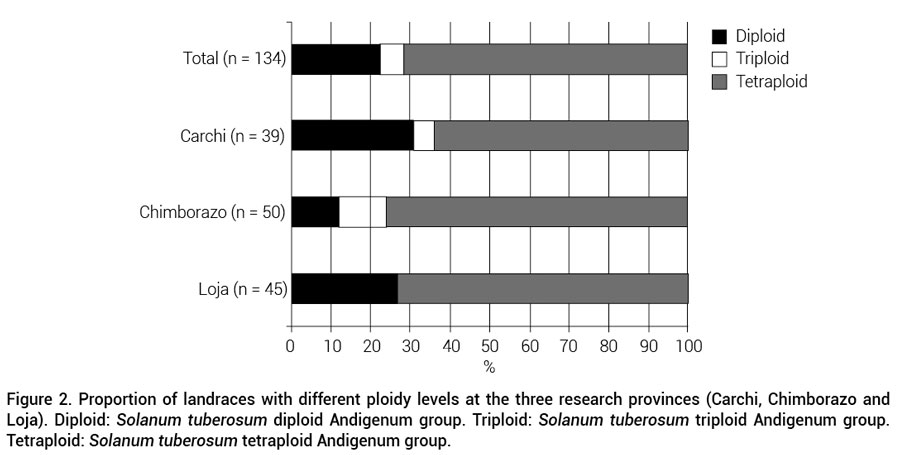
Figure 2
Figure 2. Proportion of landraces with different ploidy levels at the three research provinces (Carchi, Chimborazo and Loja). Diploid: Solanum tuberosum diploid Andigenum group. Triploid: Solanum tuberosum triploid Andigenum group.Tetraploid: Solanum tuberosum tetraploid Andigenum group.
Molecular characterization
Eight of the nine SSRs used in this study produced clear peaks. Marker STM 0019 had a considerable number of missing data in this plant material; therefore, it was not used for this analysis. Overall, the eight polymorphic markers produced 72 alleles in the 152 landraces. Table 1 shows the number of alleles and PIC values for each of the markers in the Ecuadorian landraces (with previous ploidy determination N = 134) and in European varieties. In the Ecuadorian tetraploid landraces 12 alleles were found that are not present in the European varieties, while 24 alleles present in the European collection were not found in the Ecuadorian landraces.
Cluster analysis
The UPGMA tree for the 152 Ecuadorian landraces and 10 Dutch varieties is presented in Figure 3. The dendrogram shows three main branches: 1) a group of two landraces named Uva, collected in Carchi; 2) a group consisting of the Dutch varieties; 3) all Ecuadorian landraces. The Ecuadorian landraces further split into two genetic clusters that are not associated with obvious morphological characteristics, e.g., tuber morphology or collection site. Many Ecuadorian landraces are very similar, or even identical in their SSR patterns as compared to the Dutch varieties that are all unique.
Landraces with similar names did not always cluster together. Chaucha amarilla, for example, was subdivided in three study regions: 1) four landraces collected as Chaucha amarilla; 2) two landraces under the name Chaucha amarilla alargada; and 3) two landraces collected as Chaucha amarilla redonda. The dendrogram (Figure 3) shows that Chaucha amarilla from Carchi and two landraces of Chaucha amarilla redonda from Loja are genetically identical. Two landraces from Loja, Chaucha amarilla and Chaucha amarilla alargada are also identical, but different from another Chaucha amarilla alargada collected in Loja. Finally, two Chaucha amarilla accesions, one from Carchi and the other from Chimborazo, are genetically different from each other, and from the other groups (Figure 3). Even landraces with identical names collected in the same locality did not always cluster together; for example, Colorada Antigua from Loja. On the other hand, landraces were found with different names to be genetically identical; for example, landraces Carriza, Negra or Catalina from Loja; Carriza, Negra (Morasurco) and Huevo de Indio from Carchi; and Norte Roja from Chimborazo. All of these landraces have similar tuber skin: black and white.
Mixed genotypes were also found in landraces collected at a single location. Three such cases were identified when the landrace was multiplied in the field and subsequently analyzed with the SSR markers. These were María Esperanza, Semibolona 2 and Ratona amarilla. The different representatives of these landraces ended up separated from each other in the dendrogram (Figure 3).
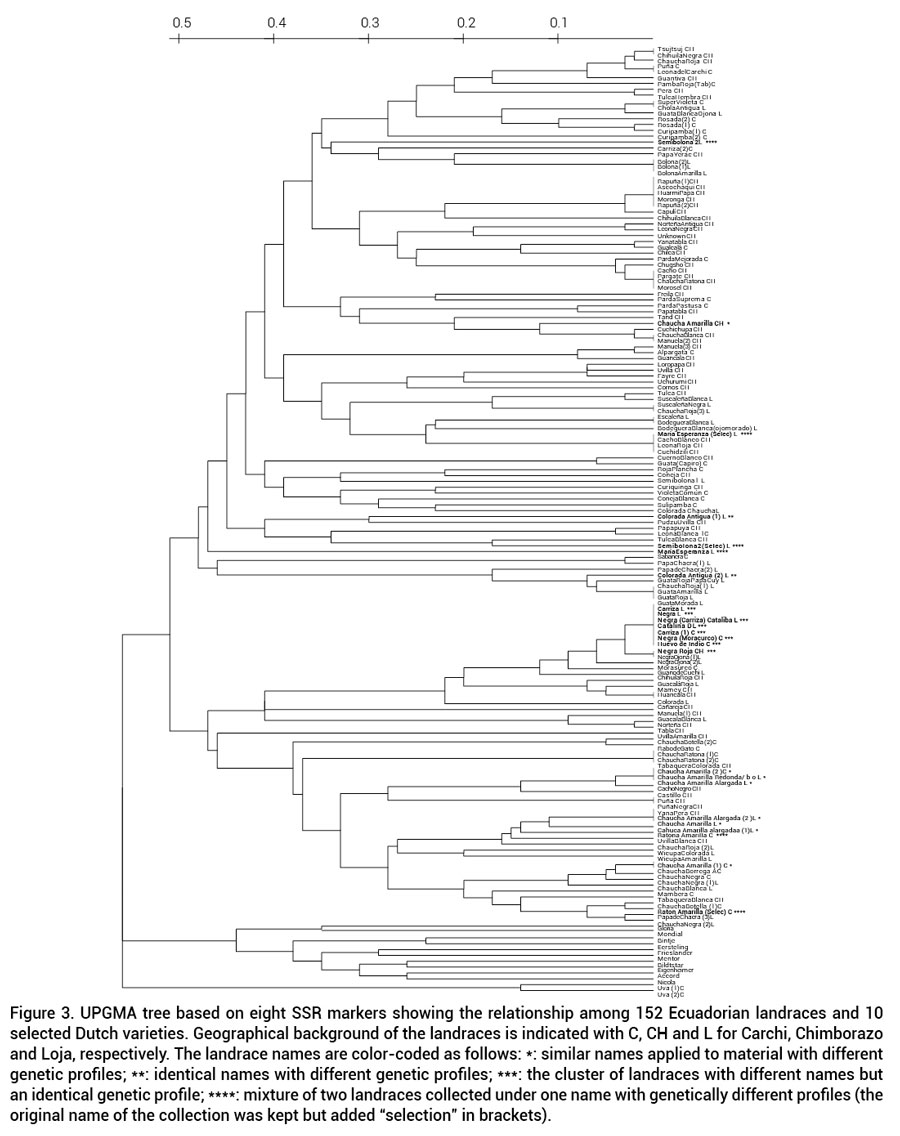
Figure 3
Figure 3. UPGMA tree based on eight SSR markers showing the relationship among 152 Ecuadorian landraces and 10 selected Dutch varieties. Geographical background of the landraces is indicated with C, CH and L for Carchi, Chimborazo and Loja, respectively. The landrace names are color-coded as follows: *: similar names applied to material with different genetic profiles; **: identical names with different genetic profiles; ***: the cluster of landraces with different names butan identical genetic profile; ****: mixture of two landraces collected under one name with genetically different profiles (theoriginal name of the collection was kept but added “selection” in brackets).
Population differentiation
The Fixation Index (Fst) was used to estimate the differences in genetic structure among several variables. First, the distribution of genetic variation among the three ploidy levels was analyzed. Fst among diploid, triploid and tetraploid material was 0.157 (P = 0.000). Due to the highly significant differences between ploidy levels and the differences based on geographical origin, ploidy level per province were analyzed. Overall Fst for diploid materials was 0.081 (P = 0.005), with pairwise Fst values as follows: Carchi-Chimborazo 0.092 (P = 0.054), Carchi-Loja 0.041 (P = 0.153) and Chimborazo-Loja 0.128 (P = 0. 045). Overall Fst for tetraploid potatoes was 0.034 (P = 0.000), with pairwise Fst comparisons: Carchi-Chimborazo 0.045 (P = 0.009), Carchi-Loja 0.021 (P = 0.045) and Chimborazo-Loja 0.035 (P = 0.000). Triploids were not analyzed as they were present in low numbers and not in all study areas.
Relationship between landrace name and genetic profile
The names of the Ecuadorian potato landraces either include tuber characteristics, such as color and shape, or are related to animals, persons, gender or objects, and are consistent with Hawkes (1947) for South America and De Haan et al. (2010) for Peru. In this study, potato names are mostly in Spanish, but with some Spanish-Kichwa and Kichwa names that reflect the mixed ethnic groups holding potato diversity in the studied areas, and is in line with observations by Hawkes (1947); however, in contrast to Hawkes, more Spanish-Kichwa names than pure Kichwa (Hawkes refers to Quechua, while we refer to Kichwa, the official name in Ecuador) were found. The decrease of transmission of Kichwa among generations (King, 1999) resulted in less Kichwa words in common potato names over the last 60 years. It was not possible to determine the etymology of all landrace names studied.
The results on the relationship between landrace name and genetic profile could lead to either under- or overestimation of the genetic diversity present in farmer fields. Some of the potato landraces with identical names were genetically different (Figure 3), which would suggest underestimation. Finding a mixture of two landraces under one name, as is the case for Semibolona 2, María Esperanza and Ratona amarilla, suggests that even more genetic diversity might be present. To really address the extent of possible underestimation of the diversity resulting from identical names attached to genetically different material a much more extensive study should be carried out. On the other hand, landraces collected under different names were found that turned out to be genetically identical (Figure 3). This would indicate that relying on names only would lead to an overestimation of diversity. Sampling on the basis of names combined with morphology, as done, possibly provides the best results.
Genetic structure based on SSR markers
In this study, SSR markers that were originally developed for the identification of European potato varieties (Reid et al., 2009; Reid et al., 2011) were used and were useful for characterizing Ecuadorian landraces (Table 1). In the largest ploidy group, containing 96 Ecuadorian tetraploid landraces, the number of alleles per genotype and PIC values was comparable to that in the European collection of 892 varieties (Reid et al., 2011). This suggests that more alleles may be found with a larger sample size including more Ecuadorian provinces.
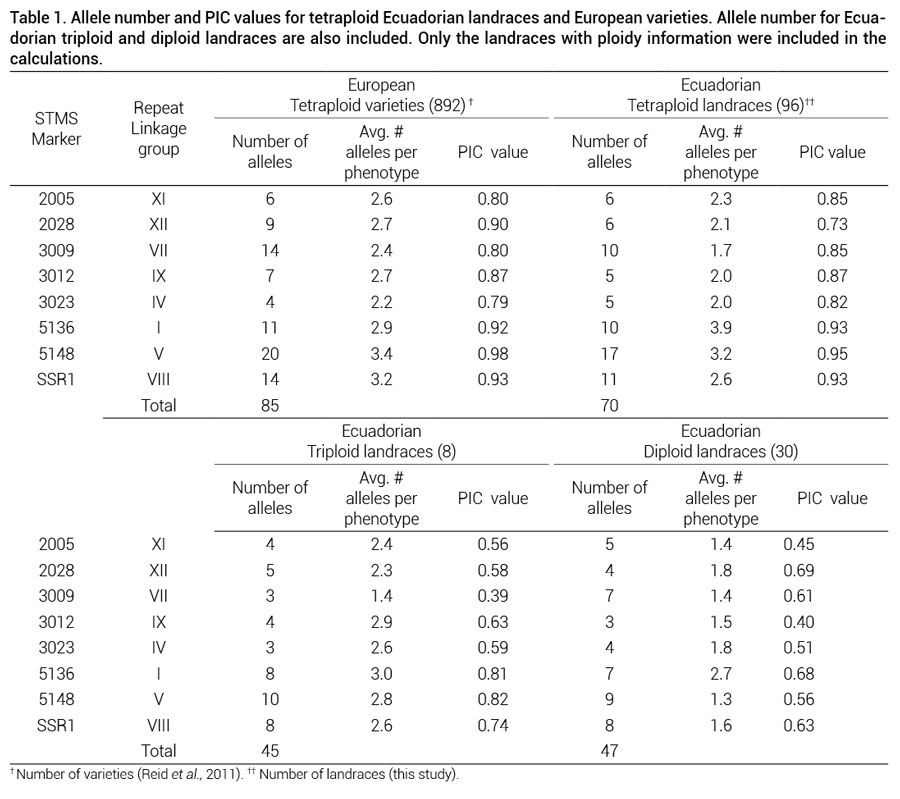
Table 1
Table 1. Allele number and PIC values for tetraploid Ecuadorian landraces and European varieties. Allele number for Ecuadorian triploid and diploid landraces are also included. Only the landraces with ploidy information were included in the calculations.
Similarities among the European and Ecuadorian material may be explained by the fact that European material was derived from Andean and Chilean potatoes (Ames and Spooner, 2008; Spooner et al., 2005; Spooner et al., 2007). The unique alleles present in the European materials might originate from Chilean potatoes or from crosses with wild relatives. The presence of unique alleles in Ecuadorian landraces shows that there is unexploited variability in these potatoes, similar to Quiros et al. (1990) for Peruvian potatoes. The difference in the SSR markers between Ecuadorian and European material is also apparent in the dendrogram of Figure 3, where there is a clear separation between the Ecuadorian landraces and the Dutch varieties.
Branch 3 in the dendrogram consists of a group of two landraces named Uva collected in Carchi. Uva landraces differentiate from other groups probably because they have been selected by farmers for late blight resistance (Cuesta et al., 2015; Pers. Comm.1).
Ploidy levels of Ecuadorian potato landraces
The ploidy level determinations confirm the presence in Ecuador of Solanum tuberosum diploid Andigenum Group (Spooner et al., 2014), formerly S. tuberosum Phureja Group (Huamán and Spooner, 2002; Spooner et al., 1992); S. tuberosum triploid Andigenum Group (Spooner et al., 2014), formerly S. tuberosum Chaucha Group (Huamán and Spooner, 2002); and S. tuberosum tetraploid Andigenum Group (Spooner et al., 2014), formerly S. tuberosum Andigenum Group (Huamán and Spooner, 2002). Ecuadorian farmers usually use the word “Chaucha” to refer to early sprouting potatoes, but it is not related to triploid species S. chaucha (S. tuberosum triploid Andigenum Group). No pentaploid cultivated potatoes were identified among the studied landraces.
The distribution of landraces over the ploidy classes is similar to what Jackson et al. (1980) found for Peru. Tetraploids are more common than diploids and triploids. Tetraploids offer advantages such as the possibility of longer storage for food and late sprouting for the next planting season to avoid diseases or dry periods. With the exception of Loja, where no triploid potatoes were found, all ploidy levels were found at each research site (Figure 2). Data shows that farmers maintain potatoes with different ploidy levels in their fields, which is similar to reports for Peru (De Haan et al., 2010; Zimmerer and Douches, 1991). The absence of triploid potatoes in Loja might be due to under-sampling in that province.
Molecular data helped distinguish the Ecuadorian landraces of potato; however, apparent grouping inconsistencies were observed with respect to ploidy levels within identical materials (Figure 3). For example, in the cluster containing Rapuña 1, Capulí, Huarmi papa, Moronga, Rapuña 2 and Ascochaqui all from Chimborazo, the ploidy level is intermixed with triploids and tetraploids. Another example is the cluster including Yanapera (Chimborazo-4x) and two landraces named Chaucha amarilla from Loja (2x). The exact nature of the apparent inconsistencies is unknown and needs further research.
Do the landraces from the three provinces constitute genetically different gene pools?
The research areas located at the North, Center and South of Ecuador present different climatic and edaphic conditions. When the three research areas are compared, significant differences between them for diploid (Fst = 0.081) as well as tetraploid (Fst = 0.034) landraces are found. On this respect, findings in this study are different from similar ones in Peru, where no differentiation was found between regions in Cusco (Brush et al., 1995) and Huancavelica. When pairwise comparisons were made between materials from different regions, it was observed that Fst for diploid materials is only significant in the Chimborazo-Loja comparison. For tetraploid landraces, the pairwise comparisons between the three provinces showed highly significant Fst values between Carchi-Chimborazo and Chimborazo-Loja.
The dendrogram does not show any genetic grouping matching the regions (Figure 3), which means that alleles are shared among the landraces from the three provinces, suggesting an exchange of landraces among the areas. An indication of such exchange is the groups of genetically identical landraces, either with the same or different names collected in two or three provinces.
Over time, farmers seem to embrace landraces as their own and maintain them for production under their local conditions. For example, landraces Parda pastusa, Parda mejorada and Parda suprema in Carchi may actually be varieties from Colombia. Interestingly, farmers from each province described their landraces mainly as “local”.
Carchi, Chimborazo and Loja hold rich potato diversity. The potato collection in this study aimed, in part, at collecting materials for ex situ conservation. Results suggest that collections are never exhaustive and that under-representation of the genetic diversity is difficult to avoid; therefore, complementary in situ conservation strategies are necessary to prevent the loss of the unique alleles and genotypes present in Ecuador.
CONCLUSIONS
The Ecuadorian landraces are known by farmers, who use common names as a traditional way to identify varieties, which also offer ethnobotanical and ethno-agricultural information. SSR molecular markers were used to effectively discriminate genetically between accessions with the same or different common names. The molecular data provided information of important genetic diversity present in the Ecuadorian potato varieties of Carchi, Chimborazo, and Loja, which could be increased with the integration of more materials sampled from other localities in Ecuador.
ACKNOWLEDGEMENTS
We thank R. van den Berg and B. Vosman of Wageningen University and Research Center, and Charles W. Barnes of INIAP, for their advice and revision to the manuscript. We also thank C. Costa and A. Navarro of INIAP, for technical support during DNA extraction and ploidy identification.
BIBLIOGRAPHY
Ames M. and D. M. Spooner (2008) DNA from herbarium specimens settles a controversy about origins of the European potato. American Journal of Botany 95:252-257.
Barandalla L., J. I. Ruiz de Galarreta, D. Rios and E. Ritter (2006) Molecular analysis of local potato cultivars from Tenerife Island using microsatellite markers. Euphytica 152:283-291.
Becher S. A., K. Steinmetz, K. Weising, S. Boury, D. Peltier, J. P. Renou, G. Kahl and K. Wolff (2000) Microsatellites for cultivar identification in Pelargonium. Theoretical and Applied Genetics 101:643-651.
Brush S., R. Kesseli, R. Ortega, P. Cisneros, K. Zimmerer and C. Quiros (1995) Potato diversity in the Andean center of crop domestication. Conservation Biology 9:1189-1198.
CIP, Centro Internacional de la Papa (2010) Genetic resources characterization and conservation Division: Genebank. Centro Internacional de la Papa. Peru. https://research.cip.cgiar.org/genebankdb/link.php?op=filter&grp=id&cipnr=704218&tmpl=default_prn&frmt=print (September 2016).
Colombo C., G. Second, T. Losada V. and A. Charrier (1998) Genetic diversity characterization of cassava cultivars (Manihot esculenta Crantz). I) RAPD markers. Genetics and Molecular Biology 21:105-113.
De Haan S., J. Núñez, M. Bonierbale and M. Ghislain (2010) Multilevel agrobiodiversity and conservation of Andean potatoes in central Peru. Mountain Research and Development 30:222-231.
De Haan S. and F. Rodriguez (2016) Potato origin and production. In: Advances in Potato Chemistry and Technology. J. Singh and L. Kaur (eds.). Elsevier. Amsterdam, The Netherlands. pp:1-32.
Esselink G. D., M. J. M. Smulders and B. Vosman (2003) Identification of cut rose (Rosa hybrida) and rootstock varieties using robust sequence tagged microsatellite site markers. Theoretical and Applied Genetics 106:277-286.
Excoffier L., G. Laval and S. Schneider (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evolutionary Bioinformatics Online 1:47-50.
Fu Y. B., G. W. Peterson, K. W. Richards, T. R. Tarn and J. E. Percy (2009) Genetic diversity of Canadian and exotic potato germplasm revealed by simple sequence repeat markers. American Journal of Potato Research 86:38-48.
Guo X., C. Xie, X. Cai, B. Song, L. He and J. Liu (2010) Meiotic behavior of pollen mother cells in relation to ploidy level of somatic hybrids between Solanum tuberosum and S. chacoense. Plant Cell Reports 29:1277-1285.
Hawkes J. (1947) On the origin and meaning of South American Indian potato names. Botanical Journal of the Journal of the Linnean Society 53:205-250.
Hawkes J. G. (1988) The evolution of cultivated potatoes and their tuber-bearing wild relatives. Die Kulturpflanze 36:189-208.
Hawkes J. G. (2004) Hunting the Wild Potatoes in South American Andes. Memories of the British Empire Potato Collecting Expedition to South America 1938-1939. Agrico Research BV. Emmeloord, The Netherlands. 224 p.
Huamán Z. and D. M. Spooner (2002) Reclassification of landrace populations of cultivated potatoes (Solanum sect. Petota). American Journal of Botany 89:947-965.
INIAP, Instituto Nacional Autónomo de Investigaciones Agropecuarias (2017) Variedades liberadas por el INIAP. Instituto Nacional Autónomo de Investigaciones Agropecuarias. Ecuador.http://www.iniap.gob.ec/nsite/index.php?option=com_content&view=article&id=346:semillas-y-plantas-variedades-liberadas&catid=9&Itemid=345 (Mayo 2017).
INIAP-DENAREF, Instituto Nacional Autónomo de Investigaciones Agropecuarias. Departamento Nacional de Recursos Fitogenéticos (2009) Informe anual 2008. Estación Experimental Santa Catalina. Instituto Nacional Autónomo de Investigaciones Agropecuarias. Santa Catalina, Ecuador. 200 p.
Ispizúa V. N., I. R. Guma, S. Feingold and A. M. Clausen (2007) Genetic diversity of potato landraces from northwestern Argentina assessed with simple sequence repeats (SSRs). Genetic Resources and Crop Evolution 54:1833-1848.
Jackson M. T., J. G. Hawkes and P. R. Rowe (1980) An ethnobotanical field study of primitive potato varieties in Peru. Euphytica 29:107-113.
King K. A. (1999) Language revitalisation processes and prospects: Quichua in the Ecuadorian Andes. Language and Education 13:17-37.
McGregor C. E., C. A. Lambert, M. M. Greyling, J. H. Louw and L. Warnich (2000) A comparative assessment of DNA fingerprinting techniques (RAPD, ISSR, AFLP and SSR) in tetraploid potato (Solanum tuberosum L.) germplasm. Euphytica 113:135-144.
Milbourne D., R. Meyer, J. E. Bradshaw, E. Baird, N. Bonar, J. Provan, W. Powell and R. Waugh (1997) Comparison of PCR-based marker systems for the analysis of genetic relationships in cultivated potato. Molecular Breeding 3:127-136.
Morillo V. E. y G. Miño C. (2011) Marcadores Moleculares en Biotecnología Agrícola: Manual de Técnicas y Procedimientos en INIAP. Instituto Nacional Autónomo de Investigaciones Agropecuarias. Estación Experimental Santa Catalina. Santa Catalina, Ecuador. 121 p.
Muñoz M., C. Folch, F. Rodriguez, J. Kalazich, S. Orena, J. Santos, R. Vargas, A. Fahrenkrog and A. Puga (2016) Genotype number and allelic diversity overview in the national collection of Chilean potatoes. Potato Research 59:227-240.
Nei M. (1973) Analysis of gene diversity in subdivided populations. Proceedings of the National Academy of Sciences of the United States of America 70:3321-3323.
Nei M. and W. H. Li (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proceedings of the National Academy of Sciences of the United States of America 76:5269-5273.
Quiros C. F., S. B. Brush, D. S. Douches, K. S. Zimmerer and G. Huestis (1990) Biochemical and folk assessment of variability of Andean cultivated potatoes. Economic Botany 44:254-266.
Reid A. and E. M. Kerr (2007) A rapid simple sequence repeat (SSR)-based identification method for potato cultivars. Plant Genetic Resources: Characterization and Utilization 5:7-13.
Reid A., L. Hof, D. Esselink and B. Vosman (2009) Potato cultivar genome analysis. In: Plant Pathology: Techniques and Protocols. Methods in Molecular Biology. Vol. 508. R. Burns (ed.)., Humana Press. New York, USA. pp:295-308.
Reid A., L. Hof, G. Felix, B. Rücker, S. Tams, E. Milczynska, D. Esselink, G. Uenk, B. Vosman and A. Weitz (2011) Construction of an integrated microsatellite and key morphological characteristic database of potato varieties on the EU common catalogue. Euphytica 182:239-249.
Ryzhova N. N., E. V. Martirosyan and E. Z. Kochieva (2010) Analysis of microsatellite locus polymorphism in potato (Solanum tuberosum) cultivars of Russian breeding. Russian Journal of Genetics 46:425-430.
Spooner D. M. (2013)Solanum tuberosum (Potatoes). In: Brenner’s Encyclopedia of Genetics. S. Maloy and K. Hughes (eds.). 2nd Edition. Academic Press. Waltham, MA. pp:481-483.
Spooner D. M., R. Castillo T. and L. López J. (1992) Ecuador, 1991 potato germplasm collecting expedition: taxonomy and new germplasm resources. Euphytica 60:159-169.
Spooner D. M., J. Núñez, F. Rodríguez, P. S. Naik and M. Ghislain (2005) Nuclear and chloroplast DNA reassessment of the origin of Indian potato varieties and its implications for the origin of the early European potato. Theoretical and Applied Genetics 110:1020-1026.
Spooner D. M., J. Núñez, G. Trujillo, M. R. Herrera, F. Guzmán and M. Ghislain (2007) Extensive simple sequence repeat genotyping of potato landraces supports a major reevaluation of their gene pool structure and classification. Proceedings of the National Academy of Sciences of the United States of America 104:19398-19403.
Spooner D. M., M. Ghislain, R. Simon, S. H. Jansky and T. Gavrilenko (2014) Systematics, diversity, genetics, and evolution of wild and cultivated potatoes. The Botanical Review 80:283-383.
Van de Peer Y. and R. De Wachter (1994) TREECON for Windows: a software package for the construction and drawing of evolutionary trees for the Microsoft Windows environment. Computer Applications in the Bioscience 10:569-570.
Zimmerer K. S. and D. S. Douches (1991) Geographical approaches to crop conservation: the partitioning of genetic diversity in Andean potatoes. Economic Botany 45:176-189.
Cuesta X., J. Rivadeneira, C. Monteros, C. Tello y E. Yanez (2015) Mejoramiento para resistencia a plagas en papa. In: Memorias del 1er Simposio Internacional de Manejo Integrado de Plagas en Solanaceas: Conocimiento para Producción Sana de Alimentos. J. Zambrano, C. Iglesias, V. Sánchez y M. Herrera (eds.). Instituto Nacional de Investigaciones Agropecuarias-INIAP. Quito, Ecuador. pp:134-135.↩
Nuijten E. and C. J. M. Almekinders (2008) Mechanisms explaining variety naming by farmers and name consistency of rice varieties in The Gambia. Economic Botany 62:148-160.
Notas de autor
alvaro.monteros@iniap.gob.ec