Editorial
Published: 23 June 2025
INTRODUCTION
In this second act, we continue discussing open science (OS) and its implementation in research in Administration. The first editorial (Limongi & Rogers, 2025) discussed the foundations of OS, starting from the reproducibility crisis to defend the engagement of advisors and graduate programs. The message was direct: OS must be the foundation for reliability and the training of new researchers. To this end, we listed five operational steps: (1) transparent planning, (2) data and code sharing, (3) open access and communication, (4) open collaboration, and (5) ethics and integrity.
It is necessary, however, to go further. The reproducibility crisis, while a catalyst (Crüwell et al., 2019), is only a symptom of deeper structural challenges. OS is a vast topic. Taxonomies such as those of Silveira et al. (2023) and Zarghani et al. (2023) map a complex ecosystem of practices ranging from infrastructure to governance. At first glance, this amplitude intimidates the individual researcher and requires a strategic cut. The solution to the crisis and the construction of a more vigorous science begin with pragmatic changes, implementable at the level of the researcher (Alessandroni & Byers-Heinlein, 2022). Therefore, this editorial focuses on this battlefront: the practices and tools we can incorporate into our daily work.
Our analysis will focus on the micro perspective of OS (Rogers, 2024). The focus is on the scientist’s workflow, who seeks to make their research transparent, reproducible, and replicable. This approach aligns with the ‘publication and sharing’ dimension of Zarghani et al. (2023), which encompasses the elements of individual scientific work: open access, open data, and reproducibility. We deliberately set aside the macro perspective - interactions with funding agencies, journals, and society - to focus on the researcher’s workflow. We believe that the transition to a more open science begins with concrete and manageable changes at our benches, digital or not. It’s a bottom-up approach. The researcher is the protagonist.
This editorial’s purpose is, therefore, informative. We present an arsenal of solutions, tools, and practices so that researchers can, in fact, practice OS and, with that, contribute to a more resilient and transparent scientific ecosystem.
REPRODUCIBLE RESEARCH
In the previous editorial (Limongi & Rogers, 2025), we outlined five steps for OS. They were a script. To equip the researcher with a practical arsenal, we focus our firepower on the first two: (1) transparent planning and (2) data and code sharing.
Why this choice? Because our concern here is with the process, not the product or the complex external interactions. We expect that by mastering planning and sharing, we will lay the foundations for genuine open collaboration (step 4). Precise planning and well-documented data are the cornerstones of any practical cooperation; without them, joint efforts become inefficient, if not impractical. The other stages - (3) open access and communication and (5) ethics and integrity - although vital, require discussions on editorial and institutional policies that are beyond the scope of the individual workflow. Therefore, planning and sharing are not isolated steps. They operate in symbiosis. A plan that anticipates sharing is born more meticulous and transparent. Conversely, preparing data and code for external scrutiny exposes gaps and forces more rigorous planning. It creates a positive feedback loop: effective planning encourages sharing, while the desire to share improves planning quality.
This editorial, therefore, presents some of the tools that enable the practical execution of these two interdependent stages. The focus is on workflow. Adopting tools that support both processes can catalyze this virtuous cycle.
LOCAL WORKFLOW
The transition to open research practices begins on the researcher’s machine (Alessandroni & Byers-Heinlein, 2022). Below we present a local workflow to adopt OS and its details.
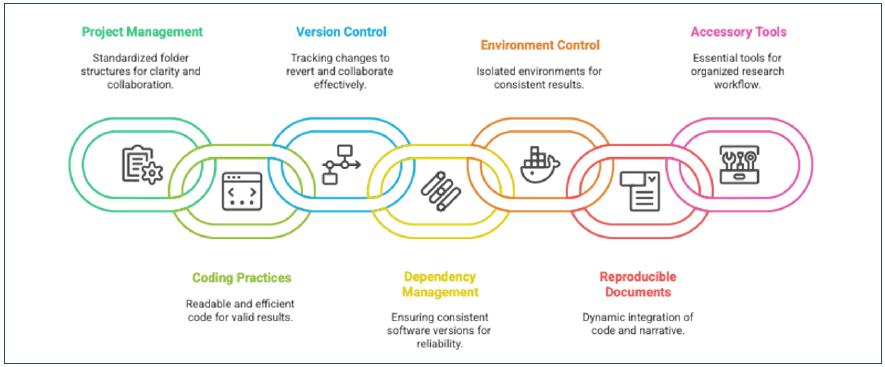
Figure 1
Workflow to Adopt Open Science.
Source: Elaborated by the authors.
Project and folder management
Directory clutter is a primary source of non-reproducibility. A standardized, logical folder structure is not a luxury, but a necessity so that others - and our future selves - can understand, replicate, and expand research (Alston & Rick, 2021; Kathawalla et al., 2021; Zandonella Callegher & Massidda, 2022). For this purpose, we refer to the TIER Protocol (https://www.projecttier.org/). It specifies a detailed directory structure for data, scripts, and documents, establishing a clear project roadmap from raw data to deliverables, simplifying cohesive sharing later (Domingos & Batista, 2021).
Good coding practices
In any quantitative or qualitative-quantitative analysis, the quality of the code determines the validity of the results. Adopting good coding practices means producing code that is not only functional but also readable, efficient, and verifiable (Rodrigues, 2023). We highlight two approaches. Functional programming breaks down complex problems into smaller, manageable functions, improving the maintainability and reusability of code (Zandonella Callegher & Massidda, 2022). For R users, the Tidyverse Style (https://style.tidyverse.org/) is a set of principles that promote a more readable data manipulation code (Rodrigues, 2023). Implementing these practices is a prerequisite for effective sharing and the ability of others to faithfully reproduce analyses.
Version control
Version control is not optional; it is a requirement for high levels of reproducibility. A version control system records all changes made to a set of files over time, allowing reversion to previous versions, comparison of modifications, and facilitation of collaboration (Vuorre & Curley, 2018). The default tool for this task is Git (https://git-scm.com/). Git streamlines the scientific workflow by enabling meticulous change tracking and simultaneous collaborative work (Vuorre & Curley, 2018). It is the basis for constructing reproducible analytical pipelines, ensuring the transparency of the research development process (Rodrigues, 2023).
Dependency management
Code that works today may fail tomorrow. The cause? Software dependencies that change without warning. Dependency management identifies and manages the correct versions of the packages and libraries a project depends on. This ensures the code works simultaneously in different environments and times (Peikert et al., 2021). For R, the renv package (https://rstudio.github.io/renv/) creates an isolated package environment and records the exact versions used (Zandonella Callegher & Massidda, 2022; Rodrigues, 2023). For Python, the conda (https://www.anaconda.com/) fulfills an analogous function. This control minimizes the risk of divergent outcomes due to package updates or incompatibilities.
Environment control
Dependency management mitigates the problem. Containerization solves it. Environment control goes a step further by creating completely isolated computational environments that encapsulate not only the packages but also the operating system and all its configurations (Moreau et al., 2023). The tool for this is Docker (https://www.docker.com/). Docker containers solve the so-called ‘dependency hell’ by eliminating system compatibility issues (Moreau et al., 2023). The Rocker Project (https://rocker-project.org/) provides ready-made and optimized Docker images for R users. By packaging the entire analytics environment, Docker ensures that it can be faithfully recreated by anyone, anywhere, at any time, overcoming the limitations of solutions such as R’s renv package (Rodrigues, 2023; Zandonella Callegher & Massidda, 2022).
Reproducible documents
The dissociation between the analysis code and the final manuscript is a critical point of failure. Reproducible documents dynamically integrate the research narrative, analysis code, and results (figures, tables) (Zandonella Callegher & Massidda, 2022). The advantage is straightforward: the final document can be recompiled at any time, ensuring that the reported results are always in sync with the data and code. This eliminates copy-and-paste errors. The cutting-edge tool for this purpose is Quarto (https://quarto.org/), a scientific publishing system that allows the creation of dynamic documents in R, Python, and Julia, exportable to multiple formats (Rogers, 2024; Rodrigues, 2023).
Accessory tools
There are tools that, although not directly linked to analysis, are part of an organized workflow. They are already in our daily lives but must be integrated intentionally. Zotero (https://www.zotero.org/) is an open-source solution for reference management, superior to many commercial alternatives (Thomas, 2023). Cloud storage services such as OneDrive, Google Drive, and Dropbox are indispensable for the secure backup and sharing of project files, complementing the functionalities of tools such as Zotero (Vuorre & Curley, 2018).
REMOTE WORKFLOW
An organized and structured local workflow is necessary but insufficient. OS requires collaboration and dissemination, which are inherently remote processes. Therefore, online platforms and repositories are the second pillar of our workflow, essential for sharing data, code, and collaboration at scale.
The repository ecosystem is vast, ranging from multidisciplinary platforms such as Zenodo (https://zenodo.org/) and Figshare (https://figshare.com/) to specialized data repositories such as Mendeley Data (https://data.mendeley.com/) and Harvard Dataverse (https://dataverse.harvard.edu/), and preprints such as arXiv (https://arxiv.org/). Within this universe, two platforms stand out for their centrality. The Open Science Framework (OSF) (https://osf.io/) positions itself as a hub that connects and supports the entire research lifecycle, from pre-registration to data and manuscript archiving, integrating with multiple other services (Sullivan et al., 2019). GitHub (https://github.com/), meanwhile, is the default platform for developing, versioning, and sharing code, facilitating technical collaboration through Git (Vuorre & Curley, 2018). The combined use of these platforms, leveraging their integrations, allows for a cohesive remote workflow, serving both search advertising and teamwork.
Workarounds: Cloud computing environments
Local containerization with Docker offers unprecedented environmental control, but its learning curve and management can be prohibitive for some researchers. Here, cloud computing platforms emerge as strategic alternatives. They abstract away the complexity of containerization, offering pre-configured browser-accessible environments that naturally facilitate collaboration and sharing. These platforms democratize access to computational tools, making reproducible research more accessible (Wiebels & Moreau, 2021).
Reproducibility operates on a spectrum: from the basic level (data and code sharing) to the intermediate level (package versioning), and finally to the high level (complete encapsulation of the computing environment). Cloud tools serve different points on this spectrum. The choice depends on the level of reproducibility required by the project.
-
Google Colab (https://colab.research.google.com/): A free, Python-focused cloud Jupyter notebook environment that requires no configuration. It is an agile tool for teaching and prototyping, reaching a basic to intermediate level of reproducibility. In the long run, the stability of the environment can be a challenge if package versions are not actively managed within the notebook.
-
Posit Cloud (https://posit.cloud/): This makes the RStudio IDE available in the browser, eliminating the need for local installation. It allows sharing of entire projects, including the R package environment, which can be controlled with the renv package. It offers an intermediate level of reproducibility, where the researcher manages the packages, but Posit manages the underlying infrastructure.
-
JupyterHub (https://jupyter.org/hub) and Binder (https://mybinder.org/): JupyterHub allows you to serve Jupyter notebooks to multiple users, with a level of reproducibility that depends on your configuration. Binder goes further: it turns a Git repository into a collection of interactive, executable notebooks in the cloud, automatically building Docker images from configuration files. Binder requires an explicit definition of the environment, promoting a high level of reproducibility and making analytics interactive for anyone, without local installation.
-
Nextjournal (https://nextjournal.com/) and Code Ocean (https://codeocean.com/) are platforms designed for the highest level of reproducibility. Nextjournal focuses on creating completely reproducible ‘computational narratives’ with automatic versioning of code and environment. Code Ocean uses ‘compute capsules’ that encapsulate code, data, environment, and results, automating the generation of Dockerfiles and versioning with Git (Clyburne-Sherin et al., 2019). Both platforms are solutions for researchers requiring long-term preservation of their computational work and exact reproducibility.
Choosing the right tool is a strategic decision. It is not a single solution, but a portfolio of options that allows each researcher to find the most appropriate path to make their science verifiable, mitigating inequalities in access to computational resources.
INTEGRATED WORKFLOW
For researchers in the R ecosystem, the RStudio IDE consolidates itself as a hub that integrates the solutions discussed in a single interface. It natively combines Quarto for reproducible documents, a graphical interface for Git with GitHub integration (Vuorre & Curley, 2018; Zandonella Callegher & Massidda, 2022), and the management of dependencies via renv (Rodrigues, 2023; Zandonella Callegher & Massidda, 2022).
This integrated environment can be enhanced. The organization of the project with the TIER Protocol, the backup in cloud services, and the sharing of the products on OSF or GitHub complement the flow. For the maximum level of control, RStudio itself can be run inside a Docker container through Project Rocker, encapsulating the entire development environment (Rodrigues, 2023; Zandonella Callegher & Massidda, 2022). RStudio thus transcends the IDE function to become a centerpiece in orchestrating an open and reproducible workflow.
For technical deepening, we recommend two works: The Open Science Manual by Zandonella Callegher and Massidda (2022) and Building Reproducible Analytical Pipelines with R by Rodrigues (2023). These readings provide the basis for researchers to stay up to date and consolidate their practices.
Essential tools for the open researcher workflow.

Note. Developed by the authors.
CONCLUSION
The journey towards more transparent science is built on consistent habits, not just on adopting advanced technologies. Reproducible research, as Alston and Rick (2021) point out, “is not just about advanced tools but also about simple work habits.” (p. 12).
The pursuit of perfection should not be an obstacle. The advice of Klein et al. (2018, p. 8) and Kathawalla et al. (2021, p. 4) is pragmatic: “Don’t let the perfect be the enemy of the good. Even if you can’t share all the details of your research, sharing something is better than nothing.” Every incremental step counts. Planning is key. “Share and document what you can. ... Start early,” recommend Klein et al. (2018), as this avoids problems and saves time.
A powerful avenue for cultural change is teaching. Researchers, as educators, can introduce these practices from an early age. Two straightforward strategies proposed by Alessandroni and Byers-Heinlein (2022) are introducing OS practices in courses and adopting open educational resources (OER). In doing so, we model the behaviors we want to see in the next generation of scientists. Our course, “Future-Proof Article: Open Science Journey in Practice” (Rogers, 2024), is an example of this approach, and the reader will find more detailed information there than that presented here (https://phdpablo.github.io/curso-open-science/).
The transition doesn’t have to be abrupt for quantitative researchers accustomed to graphical interfaces. Software such as JASP (https://jasp-stats.org/) and Jamovi (https://www.jamovi.org/) serve as bridges (Baker et al., 2023). Both are free, open-source, and R-based, offering user-friendly interfaces for statistical analysis and, in some cases, allowing export of the underlying code syntax. While they are not a substitute for the robustness of a fully versioned script, they represent a significant step beyond the ‘black boxes.’
Finally, we recognize the existence of ‘ready-made’ integrated solutions, such as WORCS (Van Lissa et al., 2021) and REPRO (Peikert et al., 2021), that offer complete workflows. However, we argue that the researcher must understand the ‘process’ before adopting the ‘product.’ The autonomy to adapt and solve problems emerges from this fundamental understanding. Based on this premise, this series’ third and final editorial will present our own ‘template,’ not as a magic solution, but as a tool to apply the knowledge built so far. The journey to OS is continuous.
REFERENCES
Alessandroni, N., & Byers-Heinlein, K. (2022). Ten strategies to foster open science in Psychology and beyond. Collabra: Psychology, 8(1), 57545. https://doi.org/10.1525/collabra.57545
Alston, J. M., & Rick, J. A. (2021). A Beginner’s Guide to Conducting Reproducible Research. The Bulletin of the Ecological Society of America, 102(2), e01801. https://doi.org/10.1002/bes2.1801
Baker, D. H., Berg, M., Hansford, K., Quinn, B. P. A., Segala, F. G., & English, E. (2023). ReproduceMe: Lessons from a pilot project on computational reproducibility [Preprint]. PsyArXiv. https://doi.org/10.31234/osf.io/k8d4u
Clyburne-Sherin, A., Fei, X., & Green, S. A. (2019). Computational Reproducibility via Containers in Psychology. Meta-Psychology, 3. https://doi.org/10.15626/MP.2018.892
Crüwell, S., Van Doorn, J., Etz, A., Makel, M. C., Moshontz, H., Niebaum, J. C., Orben, A., Parsons, S., & Schulte-Mecklenbeck, M. (2019). Seven Easy Steps to Open Science: An Annotated Reading List. Zeitschrift Für Psychologie, 227(4), 237-248. https://doi.org/10.1027/2151-2604/a000387
Domingos, A., & Batista, I. R. (2021). The TIER Protocol is a map for transparency and replicability in empirical social science. Revista Política Hoje, 40-86. https://doi.org/10.51359/1808-8708.2021.245776
Kathawalla, U.-K., Silverstein, P., & Syed, M. (2021). Easing Into Open Science: A Guide for Graduate Students and Their Advisors. Collabra: Psychology, 7(1), 18684. https://doi.org/10.1525/collabra.18684
Klein, O., Hardwicke, T. E., Aust, F., Breuer, J., Danielsson, H., Mohr, A. H., IJzerman, H., Nilsonne, G., Vanpaemel, W., & Frank, M. C. (2018). A Practical Guide for Transparency in Psychological Science. Collabra: Psychology, 4(1), 20. https://doi.org/10.1525/collabra.158
Limongi, R., & Rogers, P. (2025). Open Science in Three Acts: Foundations, Practice, and Implementation - First Act. BAR - Brazilian Administration Review, 22(1), e250079. https://doi.org/10.1590/1807-7692bar2025250079
Moreau, D., Wiebels, K., & Boettiger, C. (2023). Containers for computational reproducibility. Nature Reviews Methods Primers, 3(1), 50. https://doi.org/10.1038/s43586-023-00236-9
Peikert, A., Van Lissa, C. J., & Brandmaier, A. M. (2021). Reproducible Research in R: A Tutorial on How to Do the Same Thing More Than Once. Psych, 3(4), 836-867. https://doi.org/10.3390/psych3040053
Rodrigues, B. (2023). Building reproducible analytical pipelines with R. https://raps-with-r.dev/
Rogers, P. (2024, June). Artigo à Prova de Futuro: Jornada de Open Science na Prática. Zenodo. https://doi.org/10.5281/zenodo.12593928
Silveira, L. D., Calixto Ribeiro, N., Melero, R., Mora-Campos, A., Piraquive-Piraquive, D. F., Uribe Tirado, A., Machado Borges Sena, P., Polanco Cortés, J., Santillán-Aldana, J., Couto Corrêa da Silva, F., Ferreira Araújo, R., Enciso Betancourt, A. M., & Fachin, J. (2023). Open Science Taxonomy: Revised and expanded. Encontros Bibli: Electronic Journal of Library Science and Information Science, 28. https://doi.org/10.5007/1518-2924.2023.e91712
Sullivan, I., DeHaven, A., & Mellor, D. (2019). Open and Reproducible Research on Open Science Framework. Current Protocols Essential Laboratory Techniques, 18(1), e32. https://doi.org/10.1002/cpet.32
Thomas, P. A. (2023). Using Zotero for citation management: A step-by-step guide to organizing and citing your research. University of Kansas Libraries. https://kuscholarworks.ku.edu/handle/1808/34983
Van Lissa, C. J., Brandmaier, A. M., Brinkman, L., Lamprecht, A.-L., Peikert, A., Struiksma, M. E., & Vreede, B. M. I. (2021). WORCS: A workflow for open, reproducible code in science. Data Science, 4(1), 29-49. https://doi.org/10.3233/DS-210031
Vuorre, M., & Curley, J. P. (2018). Curating research assets: A tutorial on the git version control system. Advances in Methods and Practices in Psychological Science, 1(2), 219-236. https://doi.org/10.1177/2515245918754826
Wiebels, K., & Moreau, D. (2021). Leveraging containers for reproducible psychological research. Advances in Methods and Practices in Psychological Science, 4(2), 251524592110178. https://doi.org/10.1177/25152459211017853
Zandonella Callegher, C., & Massidda, D. (2022). The open science manual: Make your scientific research accessible and reproducible. https://doi.org/10.5281/zenodo.6521850
Zarghani, M., Nemati-Anaraki, L., Sedghi, S., Noroozi Chakoli, A., & Rowhani-Farid, A. (2023). The Application of Open Science Potentials in Research Processes: A Comprehensive Literature Review. Libri, 73(2), 167-186. https://doi.org/10.1515/libri-2022-0007
Notes
Author notes
Corresponding author: Ricardo Limongi, Universidade Federal de Goiás, Faculdade de Contabilidade Economia e Administração, Rua Samambaia, s/n, Chácaras Califórnia, CEP 74001-970, Goiânia, GO, Brazil.
